External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G25780 219 / 1e-72
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
AT4G25790 155 / 3e-47
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
AT5G57625 151 / 1e-45
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
AT4G30320 143 / 5e-43
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
AT1G01310 142 / 1e-41
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
AT4G31470 140 / 2e-41
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
AT3G19690 129 / 2e-37
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
AT4G33720 127 / 1e-36
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
AT5G02730 119 / 3e-33
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
AT5G26130 115 / 3e-32
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G096007
160 / 2e-49
AT4G30320 215 / 1e-72
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
Potri.006G171300
144 / 2e-43
AT4G30320 192 / 2e-63
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
Potri.018G007000
136 / 6e-40
AT4G31470 185 / 6e-60
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
Potri.009G083100
131 / 2e-38
AT2G14610 201 / 4e-67
pathogenesis-related gene 1 (.1)
Potri.009G083300
130 / 7e-38
AT2G14610 203 / 9e-68
pathogenesis-related gene 1 (.1)
Potri.009G083000
128 / 4e-37
AT4G33720 192 / 2e-63
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
Potri.009G083600
116 / 2e-32
AT1G50060 205 / 1e-68
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
Potri.009G082900
113 / 6e-31
AT2G14610 179 / 5e-58
pathogenesis-related gene 1 (.1)
Potri.006G215600
108 / 8e-29
AT3G09590 132 / 1e-38
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10006981
275 / 3e-94
AT4G25780 235 / 2e-79
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
Lus10005557
147 / 2e-44
AT4G31470 199 / 2e-65
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
Lus10006980
145 / 9e-44
AT4G30320 203 / 1e-67
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
Lus10001319
145 / 2e-43
AT4G30320 201 / 6e-67
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
Lus10013693
145 / 3e-43
AT4G31470 196 / 1e-64
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
Lus10019993
145 / 5e-43
AT4G30320 197 / 7e-65
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
Lus10011318
140 / 1e-41
AT4G31470 164 / 1e-51
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
Lus10015522
140 / 1e-41
AT4G30320 199 / 7e-66
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
Lus10003990
131 / 2e-36
AT1G01310 140 / 3e-40
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
Lus10025697
120 / 5e-34
AT1G50060 213 / 1e-71
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0659
CAP
PF00188
CAP
Cysteine-rich secretory protein family
Representative CDS sequence
>Potri.018G096028.1 pacid=42800484 polypeptide=Potri.018G096028.1.p locus=Potri.018G096028 ID=Potri.018G096028.1.v4.1 annot-version=v4.1
ATGAGAGCACAATTGTTCTTCGTCTTTATTCACTTCACCATCCTCACTACCAACATCATTTTAGTCACTTCCCAGTCATCTTCATTAGCAGAAACACCGC
TAAAAAAGGTTGATAATGAGACCATTTACAAGGTCTCCAAGCAACTCTGTTGGGGTTGCTTAGGAGAGTCACTACAATTCTTGTTTGCGCATAATTTAGT
AAGGGCAGCCAAATGGGAGCTACCCTTGATGTGGGACTTTCAGCTTGAAAAATATGCAGGATGGTGGGCTGGCCTAAGAAAAGCAGACTGCAAACTGCAA
CATTCGTTTCCAGAATATGATTTCAAGCTAGGAGAAAATATTTATTGGGGGAGTGGCTCCACATGGACACCAACCGATGCAGTGGGTACCTGGGCTGGTG
AAGAGAAGTACTATAATTATGCACAAAACACATGTCAGGAGGGTCAAATGTGCGGACACTATACACAGATTGTGTGGAAGACTACGAGGAGAATTGGGTG
TGCTCGTGTTGTCTGTGATGATGGGGATGTGTTTATGACTTGTAACTATGATCCTCCTGGAGAGAAGCCTGCGAGTTCCACTCATGCTCTAATTATTGGA
GAATATGAGAAGCGTCTTTGCAGAAAGTCTGGTACAAGGTTTGTCGCACGTCATAAAAAATCAGCGCGGCATCGGCTGGCGATTTTATCTTTTGTTATGC
GTTTGCTTGGTTCGTTGGAGTCGCCACCTAGTATTTAA
AA sequence
>Potri.018G096028.1 pacid=42800484 polypeptide=Potri.018G096028.1.p locus=Potri.018G096028 ID=Potri.018G096028.1.v4.1 annot-version=v4.1
MRAQLFFVFIHFTILTTNIILVTSQSSSLAETPLKKVDNETIYKVSKQLCWGCLGESLQFLFAHNLVRAAKWELPLMWDFQLEKYAGWWAGLRKADCKLQ
HSFPEYDFKLGENIYWGSGSTWTPTDAVGTWAGEEKYYNYAQNTCQEGQMCGHYTQIVWKTTRRIGCARVVCDDGDVFMTCNYDPPGEKPASSTHALIIG
EYEKRLCRKSGTRFVARHKKSARHRLAILSFVMRLLGSLESPPSI
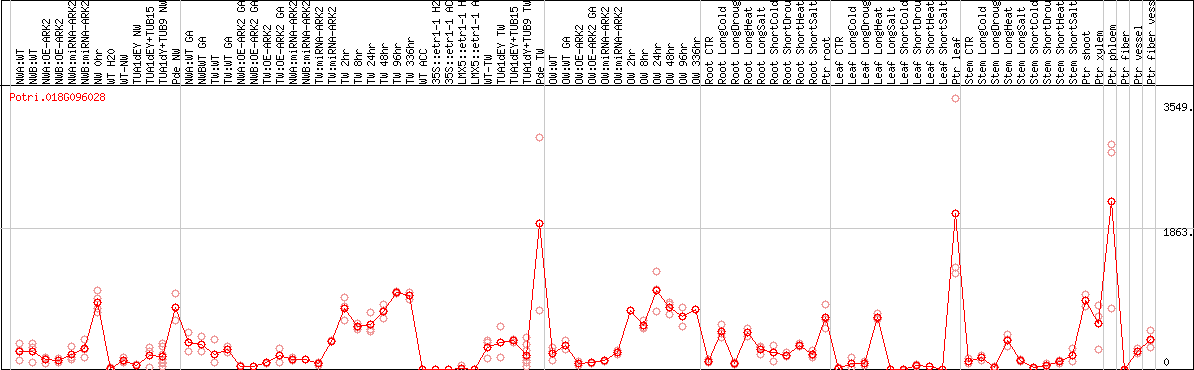
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.018G096028 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.