Potri.018G102500 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.018G102500.1 pacid=42801004 polypeptide=Potri.018G102500.1.p locus=Potri.018G102500 ID=Potri.018G102500.1.v4.1 annot-version=v4.1
ATGGCTCCCAGAACTGGAAAGGCCAAGCCTCATAAAGCAAAAGGAGAGAAGAAGAAGAAGGAAGAAAAAGTGTTGCCTACAGTCATTGAAGCCACCGTTG
AAACGCCAGACGACTCTCAAGTGACTCTCAAGGGCATCTCTACGGACAGGATCTTAGATGTAAGAAAGCTTCTTGGGGTGCATGTCGAGACGTGCCATTT
GACCAATTTCTCCTTATCTCACGAGGTACGGGGACCCAGACTCAAGGATTCGGTGGACATCATCTCTCTCAAACCCTGCCACCTCACCATTATCGAAGAG
GACTACACAGAAGATCTTTCCATAGCTCACATTCGCCGTCTCCTCGACATCGTCGCCTGCACCACCTCATTTGGGGCCTCCTCCACTTCTCCGACCAAAC
CAGCGGGTCGAATCGGAAATTCAAAGGAGTCCGGTTCAAAAGAGATCTCTTCCACCGAGACAAGAGGAGATAACAAGAAAAGCGTGAGCAAGAGCGGGAA
TGATGATTGTACGGACGCGATGGAGAAAGCCGATGCGGAGGTTTCCATGTGTCCGCCACCGCGGCTTGGCCAGTTTTATGACTTCTTTTCTTTCTCGCAC
CTCACGCCGCCGGTTCAATATATCAGGAGATCAAATCGGTCTTTCGTTGAAGATAAAACGGAAGATGATTACTTCCAGATAGACGTTAGGGTTTGCAGTG
GGAAGCCAATGAAAATTGTGGCTTCGCGAAAAGGATTTTATCCAGCTGGAAAACGCCTTCTTCTTTGCCACTCTTTGGTTAGTTTGTTGCAGCAGATTAG
CAGAGTTTTTGATGCTGCATACAAAGCTCTTATGAAAGCTTTCACTGAGCATAATAAATTTGGTAATCTTCCTTATGGATTTCGAGAGAACACGTGGGTT
GTTCCACCGGTTGTTGCTGATAATCCGTCTGGTTTTCCACCACTTCCTGTGGAGGATGAGAATTGGGGAGGTAATGGAGGAGGACATGGAAAAGATGGGA
AACATGATTACAGGCCATGGGCGAAGCAGTTTGCGATTCTGGCAGCAATGCCTTGTAAAACATCAGAAGAGAGGCAAATCCGGGACAGGAAAGCCTTTCT
ACTTCATAGTTTATTTGTCGACATTTCTGTTTTCAAAGCTGTTGCAGCAATCAAGCATATCGTTGAAAGTAATCAGTGCTTTTTAAGTGATCTTGGGAAG
TCGGTTTTGCACGAGGAAAGAGTTGGAGATTTAATTATTATTGTTATGAGAGATGCGTCTGATGCTAGCACCAAGTTGGATTGTAAAAATGATGGGTGTC
TGGTTCTTGGAGTGTCACAGGAAGAGCTTGCTCAGAGAAATTTACTTAAGGGCATAACTGCCGATGAAAGTGCAACTGTTCATGATACTCCCACTTTAGG
TGTTGTGGTTGTTCAACATTGTGGCTTCACAGCTGTTGTTAAAGTTTCTTCTGAGGTGAACTGGGAGGGGAATCGTATTCCCCAGGACATTTCCATAGAG
GATCAGACAGAAGGAGGCGCCAATGCCTTAAATGTTAATAGCTTGAGAATGCTATTGCATAATTCATCTACGCCTCAATCATCTAGTACCCCCCAACGAC
TACAGGGTGGAGATCATGAAAGTTTACGCTCTGCCAGGTCTTTGGTGCGGAAAATACTAGAGGATAGTTTGTTGAAGCTGCAGGAAGAATCCAGTAGATG
CACGAAGTCTATCAGATGGGAGCTAGGAGCATGTTGGATTCAACACTTGCAAAATCAGGCTTCAGGGAAAGCTGAGGCTAAGAAAACTGAAGAAACTAAG
CCTGAGCCTGCTGTCAAGGGGCTTGGAAAGCAAGGTGCATTACTACGGGAAATAAAGAAGAAAACAGATGTCAGAACCAGCAAAACTGAAGAGGGAAAAG
ATGTTTCTTCTGGTACCAACCTTGACACGAGCAAGAAATCTGATAGCACTAATCAGAAAGAATCGGAGAAAATGGATGAGAAAATGGAAGTCATGTGGAA
AAAGCTGCTTCCCGAAGCAGCATATCTGCGACTTAAAGAATCAGAAACCGGTCTTCATCTGAAGACACCTGATGAGTTGATTGAAATGGCACATAAATAC
TATGCTGACATTGCTCTTCCAAAACTGGTGGCAGATTTTGGCTCCCTAGAACTTTCCCCAGTTGATGGAAGAACATTGACAGATTTTATGCATACTCGGG
GTTTGCAGATGTGCTCTCTGGGGCGTGTGGTTGAACTTGCAGACAAGCTCCCTCATGTCCAATCACTCTGTATACATGAAATGATTGTCCGAGCCTTCAA
ACACATATTACAAGCTGTTGTTGCATCAGTTAACAATGTTGCTGACTTGGCTGCTTGCATTGCATCATGTCTAAATATATTGTTAGGAACGCCTTCAACT
GAAAATGAAGACTCGGATATTATAAATGATGAGAAACTAAAGTGGAAGTGGGTGGAAACCTTCCTTGCAAAGAGGTTTGGGTGGCGGTGGAAGCATGAGA
ACTGCCAGGATCTAAGAAAGTTTGCCATTCTTCGTGGGCTGTCCCACAAGGTTGGCCTTGAGCTTCTTCCTAGGGACTATGATATGGACAATGCATCTCC
TTTCAAGAAGTCTGATATTATAAGCATGGTCCCTGTATACAAGCATGTTGCATGTTCATCTGCGGATGGACGTACACTGCTAGAATCATCCAAAACTTCC
TTAGATAAGGGTAAATTGGAGGATGCTGTTAACTATGGAACTAAGGCACTGTTGAAACTCGTGTCAGTCTGTGGCCCTTTCCATCGAATGACTGCAGGAG
CATATAGTCTTTTGGCTGTAGTGCTCTACCATACTGGGGATTTTAATCAGGCTACCATTTATCAACAAAAAGCGTTGGATATTAATGAGAGGGAGCTCGG
ACTTGATCATCCTGATACAATGAAAAGTTATGGGGATCTAGCTGTCTTCTACTACCGTCTTCAACACACAGAATTAGCCTTGAAGTATGTCAATCGTGCG
CTGTATCTTTTGCATCTAACATGCGGGCCATCTCATCCAAATACTGCTGCCACATATATCAATGTTGCAATGATGGAAGAAGGATTAGGGAACGTGCATG
TTGCTCTTAGATATCTTCATGAGGCTCTCAAGTGTAACCAGAGACTCCTTGGAGCGGATCATATACAGACTGCTGCCAGCTACCATGCCATAGCAATCGC
TCTATCATTGATGGAAGCTTACTCCTTAAGTGTTCAGCATGAGCAGACCACACTGCAAATACTGCAGGCCAAACTTGGATCTGAGGATCTACGGACACAG
GATGCAGCTGCGTGGCTTGAATATTTTGAGTCAAAGGCTCTGGAACAGCAAGAAGCTGCACGCAACGGTACTCCAAAGCCAGATGCCTCAATTTCCAGTA
AAGGTCATTTAAGTGTATCAGACCTTTTGGATTATATAACACCGGATGCAGATATGAAAGCAAGAGAAGCTCAGAAGAAAGCTCGTGCAAAGGTTAAAGG
CAAACCAGGCCAGAATGAGGACACAGTCTCAGATGAATACCAAAAGGATGAAATTTTGTCACCCACTTATCCTGTTGCAGAGAACTCAAGTGATAAAGAA
AACAAATCTGAAACACAATTCGTAGAGCCTAGGAATGATAAATCTGATTTAGGTCTTCCAGATGAATCATTATTGAAGAATGATGATATGACACTAGAGG
ACAACTCAGAAGAGGGATGGCAAGAAGCTGTTCCTAAAGGCCGTTCACCTACAAGTCGCAAATCTTCTGGTTCAAGGAGGCCAAGCCTAGCAAAATTAAA
TACTAACTTTATGAATGTACCTCAGTCATCAAGATTCCGAGGAAAACCTAGCAATTTTGCATCCCCTAAAACATCGCCAAATGATCCTGCTGCTTCCAAT
GCAATGACAGTTCCAGTTCGTAAAAAGTTTGTTAAGAGTGCCAGCTTCGGTCCAAAAGTAAATAACTCTGGCGCCTCAACTGGTGGAGCAGAGAAATCAA
GTAATGCTAAATCAGCCCCAGCTACCCCTGCCTCAACTGAGCAAGCTGCCAAAGCAGCTCCAATGGCAAGTCCAATCAGTGTTCAGGCAGCTGGAAAAAT
GTTTTCCTATAAAGAAGTTGCTTTGGCTCCTCCTGGGACAATTGTGAAGGCAGTAGCAGAACAGTTGCCAAAAGGAAACCCTACAAAGGAACCAAGTCCT
CAGGGAAGCCATGAGACAGCTGCAACTGATGTCAAATCAGAAGGGGTGACAGCATTAAAAGCTGTGGAGGTAGGAAAACTTCAGAAACCTGAAGGAGAGA
GACAGCTTCCAGCTTCAGAGGGAATGAAAAGTCCTGTTGATCAAGAGAGAGAAACAGGAGGGGTTCTGGTGGCGACAGAGAAACTTGAAGAAATAAAATT
TGCTGATGAGGATCACATAGATACAGAAGATGGAGGTGCTGAAATTAAAGTAGTTACTGTTAAAGATACTACCGCAGAAGCTGAAACTATTTCAGACTTG
GGTCATGAGAATTTGGATACTTCCAAGGATTCAAATACCATGTCTTCCCCAACTGAAGTGCCTGATACAAGAGCATCAGATGGTTTCCCTTCAGCTTGCC
CTGATCTGAAACCTCAATCTACTTCAATTGAAAAGGCTGGTTTGTTGGAAAAAGATTCTTCCAGCACAAATGAAAAGGTAGAAGATGAGAATACTCCAGA
TCTTTCTAATGACAATACAAATGCTAAGCTATTGTCAACTGGAGGAGTAAAGCAAGATGATGCGGAAACTGGTAAGGAGGCGACCAAGAAACTTTCTGCA
GCTGCACCACCATTTAATCCATCCACAATTCCAGTTTTCAGCTCAGTTACTGTCCCAGGATTCAAAGATCACGGACTTCTTCCTCCACCTGTGAACATTC
CTCCAATGCTTACTGTCAATCCTGTTCGTAGATCACCTCATCAATCAGCTACAGCTCGGGTTCCATATGGTCCGAGGTTGTCTGGTGGCTATAACAAATC
TGGAAATCGGGTTCCACGTAACAAGCCGAGCTTCCACAACGGTGAGCATACAGGGGATGGGAATCATTTCAGCCCTCCTAGAATCATGAACCCACATGCA
GCTGAATTTGTGCCTTGCCAACCTTGGGTTCCCAATGGCTATCCATTGCAACACAATGGCTATATGGCTACCACAAATGGTATGCCAGTATCTCCAAATG
GCTATCCCATTTCCCCAACCAGCATTCCTGTGTCACCAAATGGATATCCAGCATCGTTGAATGGTATTGAAGTGACTCAAAATGGGTTTCCAGCCTCTCT
AGTCGGTTCAGAAGAAACACCCACGTCAGTTTCTGTTGATGTTGGTGGTGAAAACAAGAGTGAGGCTGCAGCAGAAAATGGCACTGAAAATTCTGAGATT
GAAGTAGGAGTTGAAAACCATTCAAGTGATTATGAAAATCAAAAATATCAAGAAGAAAACGTAAATCCTGAAATAGGAGAGAAGCCTGCAGAGGTCGCGG
TGACTAGTGATACTGTTGTAGCAAAAGAAACTTGTGATAGCTTGCCAACTGAGGAGAAACCAAGCAAGTGTTGGGCAGATTACAGCGACAACGAAGCCGA
GATCGTTGAGGTTGCAAGTTGA
|
|||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.018G102500.1 pacid=42801004 polypeptide=Potri.018G102500.1.p locus=Potri.018G102500 ID=Potri.018G102500.1.v4.1 annot-version=v4.1
MAPRTGKAKPHKAKGEKKKKEEKVLPTVIEATVETPDDSQVTLKGISTDRILDVRKLLGVHVETCHLTNFSLSHEVRGPRLKDSVDIISLKPCHLTIIEE
DYTEDLSIAHIRRLLDIVACTTSFGASSTSPTKPAGRIGNSKESGSKEISSTETRGDNKKSVSKSGNDDCTDAMEKADAEVSMCPPPRLGQFYDFFSFSH
LTPPVQYIRRSNRSFVEDKTEDDYFQIDVRVCSGKPMKIVASRKGFYPAGKRLLLCHSLVSLLQQISRVFDAAYKALMKAFTEHNKFGNLPYGFRENTWV
VPPVVADNPSGFPPLPVEDENWGGNGGGHGKDGKHDYRPWAKQFAILAAMPCKTSEERQIRDRKAFLLHSLFVDISVFKAVAAIKHIVESNQCFLSDLGK
SVLHEERVGDLIIIVMRDASDASTKLDCKNDGCLVLGVSQEELAQRNLLKGITADESATVHDTPTLGVVVVQHCGFTAVVKVSSEVNWEGNRIPQDISIE
DQTEGGANALNVNSLRMLLHNSSTPQSSSTPQRLQGGDHESLRSARSLVRKILEDSLLKLQEESSRCTKSIRWELGACWIQHLQNQASGKAEAKKTEETK
PEPAVKGLGKQGALLREIKKKTDVRTSKTEEGKDVSSGTNLDTSKKSDSTNQKESEKMDEKMEVMWKKLLPEAAYLRLKESETGLHLKTPDELIEMAHKY
YADIALPKLVADFGSLELSPVDGRTLTDFMHTRGLQMCSLGRVVELADKLPHVQSLCIHEMIVRAFKHILQAVVASVNNVADLAACIASCLNILLGTPST
ENEDSDIINDEKLKWKWVETFLAKRFGWRWKHENCQDLRKFAILRGLSHKVGLELLPRDYDMDNASPFKKSDIISMVPVYKHVACSSADGRTLLESSKTS
LDKGKLEDAVNYGTKALLKLVSVCGPFHRMTAGAYSLLAVVLYHTGDFNQATIYQQKALDINERELGLDHPDTMKSYGDLAVFYYRLQHTELALKYVNRA
LYLLHLTCGPSHPNTAATYINVAMMEEGLGNVHVALRYLHEALKCNQRLLGADHIQTAASYHAIAIALSLMEAYSLSVQHEQTTLQILQAKLGSEDLRTQ
DAAAWLEYFESKALEQQEAARNGTPKPDASISSKGHLSVSDLLDYITPDADMKAREAQKKARAKVKGKPGQNEDTVSDEYQKDEILSPTYPVAENSSDKE
NKSETQFVEPRNDKSDLGLPDESLLKNDDMTLEDNSEEGWQEAVPKGRSPTSRKSSGSRRPSLAKLNTNFMNVPQSSRFRGKPSNFASPKTSPNDPAASN
AMTVPVRKKFVKSASFGPKVNNSGASTGGAEKSSNAKSAPATPASTEQAAKAAPMASPISVQAAGKMFSYKEVALAPPGTIVKAVAEQLPKGNPTKEPSP
QGSHETAATDVKSEGVTALKAVEVGKLQKPEGERQLPASEGMKSPVDQERETGGVLVATEKLEEIKFADEDHIDTEDGGAEIKVVTVKDTTAEAETISDL
GHENLDTSKDSNTMSSPTEVPDTRASDGFPSACPDLKPQSTSIEKAGLLEKDSSSTNEKVEDENTPDLSNDNTNAKLLSTGGVKQDDAETGKEATKKLSA
AAPPFNPSTIPVFSSVTVPGFKDHGLLPPPVNIPPMLTVNPVRRSPHQSATARVPYGPRLSGGYNKSGNRVPRNKPSFHNGEHTGDGNHFSPPRIMNPHA
AEFVPCQPWVPNGYPLQHNGYMATTNGMPVSPNGYPISPTSIPVSPNGYPASLNGIEVTQNGFPASLVGSEETPTSVSVDVGGENKSEAAAENGTENSEI
EVGVENHSSDYENQKYQEENVNPEIGEKPAEVAVTSDTVVAKETCDSLPTEEKPSKCWADYSDNEAEIVEVAS
|
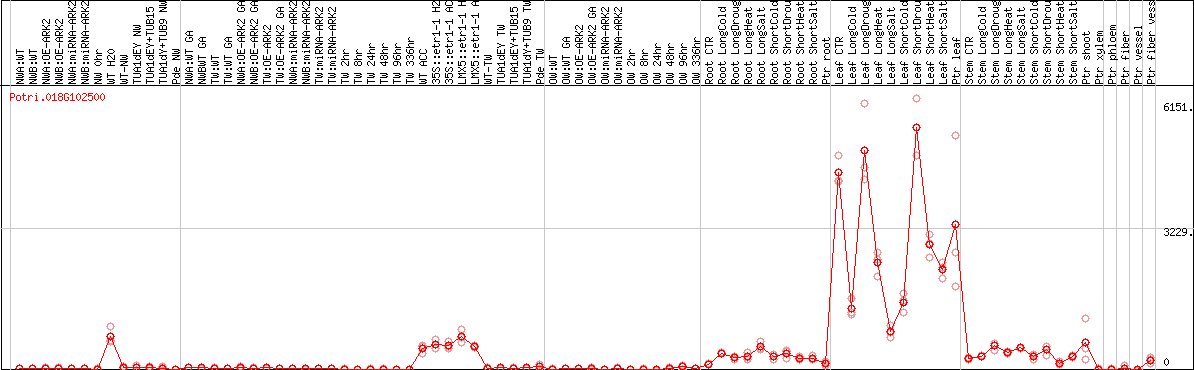
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.018G102500 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
