External link
Symbol
ATNAP10.1
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G63270 387 / 2e-138
ABCI1, ATNAP10
ATP-binding cassette I1, non-intrinsic ABC protein 10 (.1)
AT1G67940 61 / 6e-11
ABCI17, AtSTAR1, ATNAP3
ARABIDOPSIS THALIANA NON-INTRINSIC ABC PROTEIN 3, ATP-binding cassette I17, non-intrinsic ABC protein 3 (.1)
AT3G62150 54 / 3e-08
ABCB21, PGP21
ATP-binding cassette B21, P-glycoprotein 21 (.1)
AT2G41700 54 / 3e-08
AtABCA1, ABCA1
Arabidopsis thaliana ATP-binding cassette A1, ATP-binding cassette A1 (.1.2)
AT4G01830 53 / 7e-08
ABCB5, PGP5
ATP-binding cassette B5, P-glycoprotein 5 (.1)
AT5G14100 52 / 1e-07
ABCI11, ATNAP14
ARABIDOPSIS THALIANANON-INTRINSIC ABC PROTEIN 14, ATP-binding cassette I11, non-intrinsic ABC protein 14 (.1)
AT1G27940 52 / 1e-07
ABCB13, PGP13
ATP-binding cassette B13, P-glycoprotein 13 (.1)
AT2G47000 52 / 2e-07
PGP4 ,MDR4, ABCB4, ATPGP4
MULTIDRUG RESISTANCE 4, ARABIDOPSIS P-GLYCOPROTEIN 4, ATP-binding cassette B4, ATP binding cassette subfamily B4 (.1)
AT5G64840 52 / 2e-07
ABCF5, ATGCN5
ATP-binding cassette F5, general control non-repressible 5 (.1)
AT1G02530 50 / 5e-07
ABCB12, PGP12
ATP-binding cassette B12, P-glycoprotein 12 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G132000
63 / 9e-12
AT1G67940 390 / 3e-138
ARABIDOPSIS THALIANA NON-INTRINSIC ABC PROTEIN 3, ATP-binding cassette I17, non-intrinsic ABC protein 3 (.1)
Potri.014G113500
56 / 5e-09
AT3G62150 1879 / 0.0
ATP-binding cassette B21, P-glycoprotein 21 (.1)
Potri.014G113200
56 / 9e-09
AT3G62150 1871 / 0.0
ATP-binding cassette B21, P-glycoprotein 21 (.1)
Potri.006G049300
56 / 1e-08
AT2G41700 2741 / 0.0
Arabidopsis thaliana ATP-binding cassette A1, ATP-binding cassette A1 (.1.2)
Potri.002G187500
54 / 2e-08
AT3G62150 1860 / 0.0
ATP-binding cassette B21, P-glycoprotein 21 (.1)
Potri.014G113100
54 / 3e-08
AT1G02520 1773 / 0.0
multi-drug resistance 8, ATP-binding cassette B11, P-glycoprotein 11 (.1)
Potri.014G113000
54 / 4e-08
AT3G62150 1767 / 0.0
ATP-binding cassette B21, P-glycoprotein 21 (.1)
Potri.010G045900
53 / 5e-08
AT1G70610 1033 / 0.0
ATP-binding cassette B26, transporter associated with antigen processing protein 1 (.1)
Potri.002G187400
52 / 1e-07
AT3G62150 1702 / 0.0
ATP-binding cassette B21, P-glycoprotein 21 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10006727
382 / 3e-136
AT1G63270 404 / 3e-145
ATP-binding cassette I1, non-intrinsic ABC protein 10 (.1)
Lus10022692
380 / 1e-135
AT1G63270 403 / 8e-145
ATP-binding cassette I1, non-intrinsic ABC protein 10 (.1)
Lus10040034
79 / 1e-19
AT1G63270 79 / 3e-20
ATP-binding cassette I1, non-intrinsic ABC protein 10 (.1)
Lus10010067
62 / 6e-11
AT1G02520 1748 / 0.0
multi-drug resistance 8, ATP-binding cassette B11, P-glycoprotein 11 (.1)
Lus10035453
57 / 2e-09
AT1G67940 371 / 1e-130
ARABIDOPSIS THALIANA NON-INTRINSIC ABC PROTEIN 3, ATP-binding cassette I17, non-intrinsic ABC protein 3 (.1)
Lus10005798
57 / 2e-09
AT5G64840 1049 / 0.0
ATP-binding cassette F5, general control non-repressible 5 (.1)
Lus10006804
57 / 4e-09
AT5G64840 1050 / 0.0
ATP-binding cassette F5, general control non-repressible 5 (.1)
Lus10038050
57 / 5e-09
AT2G47000 1697 / 0.0
MULTIDRUG RESISTANCE 4, ARABIDOPSIS P-GLYCOPROTEIN 4, ATP-binding cassette B4, ATP binding cassette subfamily B4 (.1)
Lus10036857
56 / 1e-08
AT1G70610 953 / 0.0
ATP-binding cassette B26, transporter associated with antigen processing protein 1 (.1)
Lus10006209
55 / 1e-08
AT1G70610 939 / 0.0
ATP-binding cassette B26, transporter associated with antigen processing protein 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF00005
ABC_tran
ABC transporter
Representative CDS sequence
>Potri.018G110200.2 pacid=42800364 polypeptide=Potri.018G110200.2.p locus=Potri.018G110200 ID=Potri.018G110200.2.v4.1 annot-version=v4.1
ATGTTGCTTCGGAAAGCACCCCTCCCTCGGATCCTTCTCGACAATGTATCGTGCATGAGAAATGCCCAGCAAATCTTGCGGCATGTAAATGTCTCTGTTC
ATGATGGTGGGGCGCTTGTACTGACAGGCAGCAACGGTTCGGGCAAGAGCACTTTTTTGCGAATGTTAGCTGGTTTTTCTAAGCCTTCTGCGGGTCAGAT
ACTTTGGAATGGTCATGATGTGACTGAATCTGGAGTGTTCCATCAGTACAAGCTTCAGCTCAACTGGCTCTCCCTGAAAGATGCCATCAAGGAAAAGTTT
ACTGTACTTGACAATGTTCAGTGGTTTGAACTTCTTGAGGGCAAGCAAGGGAATTCTCTGCCTGCTATTGAGTTGATGGGGCTTGGAAGATTAGCAAAGG
ATAAGGCAAGAATGTTGTCCATGGGTCAGCGGAAAAGGCTGCAGCTTGCAAGGTTACTGGCGATTGACAGGCCAATTTGGTTGTTGGATGAGCCTTCTGT
TGCATTAGATGATGACGGGGTGAGGTTGCTTGAGTACATCATCGCAGAGCATAGAAAGAAGGGAGGGATTGTGTTTGTTGCGACTCATCTACCAATTAAC
ATTGAGGATGCCATGTACTTGAGGTTGCCACCAAGGTTCCCCAGAAGGATGACCTTGGTAGATATGCTCGATCGTGCAGAAATCTCATAA
AA sequence
>Potri.018G110200.2 pacid=42800364 polypeptide=Potri.018G110200.2.p locus=Potri.018G110200 ID=Potri.018G110200.2.v4.1 annot-version=v4.1
MLLRKAPLPRILLDNVSCMRNAQQILRHVNVSVHDGGALVLTGSNGSGKSTFLRMLAGFSKPSAGQILWNGHDVTESGVFHQYKLQLNWLSLKDAIKEKF
TVLDNVQWFELLEGKQGNSLPAIELMGLGRLAKDKARMLSMGQRKRLQLARLLAIDRPIWLLDEPSVALDDDGVRLLEYIIAEHRKKGGIVFVATHLPIN
IEDAMYLRLPPRFPRRMTLVDMLDRAEIS
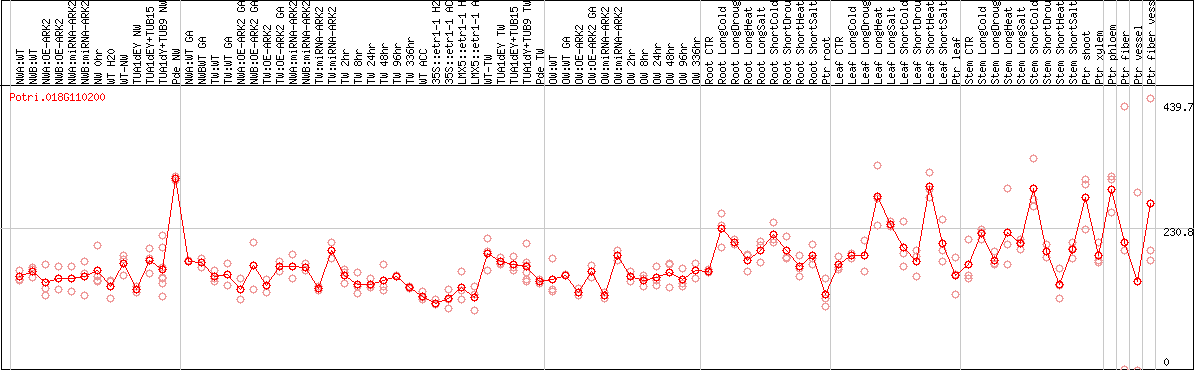
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.018G110200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.