Potri.018G128400 [POPLAR]

| External link |
|
|||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.018G128400.1 pacid=42800907 polypeptide=Potri.018G128400.1.p locus=Potri.018G128400 ID=Potri.018G128400.1.v4.1 annot-version=v4.1
ATGTCCCTCTCTAGATCACAACCAAGCACCACCACAACTGATACAATCCTCCCTGGCCCACCTTCCCGCAATAACTTTTCCTCCCTCGATCTTTCATCTT
CAAATCTCCTCGCCTTTCCCTCCGGTTCTTCCATTTCCATCGTTGATGCTCTTTCCCTCCAGTTAATCTCCACTTTTCCTCTCCCTCCTCCTCCTTCTTC
CACTTCCTCTCCTTCTCTTTCCCCCTTCATCACTTCCGTACGCTTCACGCCTTCTCCTCTCAATCGCAACCTCCTCTCCACCGAACCTTCCTCCTCCCAC
CTCCTCCTCGCCGCCGCCGACCGCCATGGCCGTATTGCTCTCCTCGATTTCCGCCTCAAGTCCATTGTCCTCTGGCTCGAACCTGATCCCAATCCGAAAT
CTGGTATCCAAGACCTCTGCTGGATCCTCTCTCGTTCCGATTCATACGCCCTCGCTGCCATTTCTGGTCCTTCTTCTCTTTATCTCTACACCACCACCGG
CGCCGCTTCCACTGCCACCGCTTCGAATTGTTGTTTCTTTAAGTACGATGCGTCGCCGGAGTTTTTGTCGTGCATTCGTCGTGATCCGTTTGATTCGCGG
CATTTTTGTGTCATCGGACTTAAAGGATTTTTATTGTCGGTGAAAGTGCTCGCAGAGAGTGAGAATGACGTCATTTTAAAGGAATTTAAGATTCCGACGG
ATTATAGTGATTTGTTGAGGCTAGAGAAAGACGTAACGCCGAGCAGTGGTGGTGTGGGAGGATCCTTGGCGCCGGCATCGGCGGTTTTTCCGTTGTATAG
TGTGAAAATGGCGTTTTCACCACAATGGAGGAATATTCTGTTTGTCACGTTTCCTAGGGAGCTTGTGGTTTTTGATTTGAAATACGAGACCGTGCTTTTT
AGCGCTGCATTGCCTAGAGGCTGTGGGAAGTTTCTAGATGTTTTGCCGGATCCGAATAATGAATTGCTTTACTGCGCTCATCTTGATGGAAAGCTCAGTA
TTTGGCGGCGAAAAGAAGGAGAACAGGTGCATGTAATGTGTGCAATGGAGGAGTTGATGCCGTCAATTGGAACATCGGTCCCTTCTCCTTCAGTTCTAGC
TGTTGCCATTTGTCAGTCAGAGTCTACGCTCCAACATGTTGCTAAGATTTGTTCTGATGCACCTGATTCTCCGTCGGCTGAGGTGGATTTTGACAATCCT
TTTGATTTCTGTGATGATACTGTTGTTCACTCTACTACACATATGATCTCAATTTCTGATGATGGAAAAGTATGGAACTGGCTGTTAACTGCTGAAGGGA
CTGGAGACAATCATAAGGATACGGTTGCTGATTCCCGTGAAATTCCACTTATAGGGGACAATGCCAATGCTGTAGTTGTGACTGATGGACTTGGCAAAGA
AGCAGGCAAGCAACAAGAGCTTGGGAATGGCAATAAAAACCGCCTCTCATCTACTTTGAGCCAGGATTTGTCGTTTAAGATCAGTTTAGTTGGACAGCTG
CAGCTTCTTTCTTCGACAGTTACTATGCTGGCTGTACCCTCTCCTTCTCTAATAGCTACTTTGGCCCGTGGGGGGAATTATCCTGCTGTAGCTGTTCCTC
TGGTTGCTCTGGGTACTCAATCTGGGACAATTGATGTTGTTGATGTTTCTGCAAATGCTGTTGCAGCTAGTTTTTCAGTTCATAATAGTACGGTTAGGGG
GTTACGATGGCTTGGAAATTCTAGACTTGTTTCATTTTCTTACAATCAGGTGAATGAAAAAAATGGAGGTTACAATAATCGGCTTGTTGTAACCTGTCTT
AGAAGCGGTCTTAACAGACCATTTCGAGTTTTACAAAAGCCAGAACGTGCACCCATTAGAGCTCTACGAACTTCTTCATCTGGAAGGTATCTTCTTATTT
TATTCCGTGATGCACCGGTGGAGGTTTGGGCAATGACAAAAACTCCCATCATGCTTAGATCATTAGCTCTTCCATTTACGGTATTGGAATGGACTCTTCC
AACAGTTCCTCGGCCAGTTCAAAATGGTCCTTCTAAACAAGTGTTGTGGTCATCGAAAGATCAAACGCCTGTAGCACAAGATGGAGCATCCACTGCAAAG
GAACCTGCATCAGAATCCACAGCAGGAAGCTCAGATGCTTCTCAGGATGACACTGCTGAAAGTTTTGCATTTGCACTAGTAAATGGTGCTCTTGGAGTTT
TTGAGGTTCATGGGCGGAGGATCCGGGATTTCAGACCGAAATGGCCTTCATCTTCATTTGTGTCTTCTGATGGATTGATTACGGCCATGGCCTACCGCTT
GCCTCATGTTGTCATGGGGGATAGGTCTGGGAACATACGGTGGTGGGATGTAACAACTGGGCATTCTTCATCATTTAACACTCACAGGGAAGGAATCAGG
AGAATCAAATTCTCACCTGTTGTTCCTGGAGACCGCAGCCGAGGACTTATTGCTGTCCTTTTTTATGACAATACCTTTTCTATTTTTGACCTTGATTTGC
CAGACCCGTTAGCTAATTCCCTTTTACAACCTCTATTTCCTGGAACCCTTGTGTTGGAACTCGATTGGTTGCCTCTGCGAACTAATAGAAATGACCCTCT
GGTATTATGTATTGCTGGAGCTGATAGTAGCTTTCGCCTTGTTGAGGTTAACGTAAATGACAAAAAACTTGGCCTCCAACCCCGAGCTATTAAAGAAAAA
TTTCAGCCTATGCCTATATGCTCTCCCATACTGCTTCCCACACCACATGCACTGGCATTGCGGATGATCTTGCAATTAGGTGTCAAACCCTCTTGGTTTA
ACACTTGCAGCACAACTATTGACAAGAGGCCTCACTTAATTCCTGGAACTGCTTCATTCAAAGGGGATCTCCGGAATTACATAATTGATCTTCCACCTGT
TGGTGACTCTGTGGTGCCAGAAATGCTCCTCAAAGTGTTAGATCCTTACAGGAGAGAAGGCTGCATACTTGATGATGAAACGGCTAGACTATATGCTATC
GTAGTCAAGAAAGGCTGTGCTGCGAGGTTTGCATTTGCAGCTGCAATTTTTGGTGAAACCTCAGAAGCACTATTCTGGTTGCAGTTGCCTCGTGCTCTTA
AACATTTGATGGATAAGCTAGTTACGAAGTCAACACAGAAAGCACCTGTCTCAGCATCAACCCCAGAGCTTGATGACGTGACTATGCTGAACAGAATTTC
ATCAAAGGGGAGGTCGGTAATTGGGACTGAGAAAAAGGATCCATTGAGTGAAGGTCAACTTAGATCAATGGCTTTCCAGAAAGAAGAACTATGGGAAAGT
GCTTGTGAGCGCATTCCTTGGCATGAGAAATTGGAGGGAGAAGAGGCTATTCAGAATCGAGTACATGAGCTCGTTTCTATTGGAAACTTAGAAGCAGCAG
TCAGCTTGTTGCTTTCTACTTCTCCTGAGAGCTCTTATTTCTATGTAAATGCTCTACGTGCAGTCGCTCTCTCGTCTGCTGTGTCAAGGTCTCTTCATGA
GCTTGCAGTTAAGGTTGTTGCAGCCAACATGGTCCAGACAGACAGATCATTGTCTGGCACTCATCTTCTTTGTGCTGTTGGAAGGTATCAAGAGGCTTGT
TCACAGTTGCAAGATGCAGGATGCTGGACGGATGCTGCAACTTTGGCGGCTACACATTTAAGTGGATCTGATTATGCTAGGGTGTTGTTAAGATGGGCCA
ACCATGTCCTCCATGCCGAACACAACATCTGGAGGGCTCTTATTCTTTATGTTGCAGCTGGTGCATTGCAAGATGCCCTAGCAGCACTCCGGGAAACACA
ACAACCTGATACAGCTGCCATGTTCATACTTGCTTGCCATGAAGGCCATGCACAATTTATATCAAATTTAGGAAATTCAGATGACGAATCAGGGTCTTCA
ATCAAGGATACTCTCGTTGGCTTGCCAGGGTTGAATCCAGAGAATGAAGATGTCATTGCAGTAGGTGAATATTATGGACAATACCAAAGAAAGTTGGTGC
ATCTGTGCATGGACTCCCAGCCATTTTCTGACTAA
|
|||||||||||||||||||||||||
|
AA sequence
|
>Potri.018G128400.1 pacid=42800907 polypeptide=Potri.018G128400.1.p locus=Potri.018G128400 ID=Potri.018G128400.1.v4.1 annot-version=v4.1
MSLSRSQPSTTTTDTILPGPPSRNNFSSLDLSSSNLLAFPSGSSISIVDALSLQLISTFPLPPPPSSTSSPSLSPFITSVRFTPSPLNRNLLSTEPSSSH
LLLAAADRHGRIALLDFRLKSIVLWLEPDPNPKSGIQDLCWILSRSDSYALAAISGPSSLYLYTTTGAASTATASNCCFFKYDASPEFLSCIRRDPFDSR
HFCVIGLKGFLLSVKVLAESENDVILKEFKIPTDYSDLLRLEKDVTPSSGGVGGSLAPASAVFPLYSVKMAFSPQWRNILFVTFPRELVVFDLKYETVLF
SAALPRGCGKFLDVLPDPNNELLYCAHLDGKLSIWRRKEGEQVHVMCAMEELMPSIGTSVPSPSVLAVAICQSESTLQHVAKICSDAPDSPSAEVDFDNP
FDFCDDTVVHSTTHMISISDDGKVWNWLLTAEGTGDNHKDTVADSREIPLIGDNANAVVVTDGLGKEAGKQQELGNGNKNRLSSTLSQDLSFKISLVGQL
QLLSSTVTMLAVPSPSLIATLARGGNYPAVAVPLVALGTQSGTIDVVDVSANAVAASFSVHNSTVRGLRWLGNSRLVSFSYNQVNEKNGGYNNRLVVTCL
RSGLNRPFRVLQKPERAPIRALRTSSSGRYLLILFRDAPVEVWAMTKTPIMLRSLALPFTVLEWTLPTVPRPVQNGPSKQVLWSSKDQTPVAQDGASTAK
EPASESTAGSSDASQDDTAESFAFALVNGALGVFEVHGRRIRDFRPKWPSSSFVSSDGLITAMAYRLPHVVMGDRSGNIRWWDVTTGHSSSFNTHREGIR
RIKFSPVVPGDRSRGLIAVLFYDNTFSIFDLDLPDPLANSLLQPLFPGTLVLELDWLPLRTNRNDPLVLCIAGADSSFRLVEVNVNDKKLGLQPRAIKEK
FQPMPICSPILLPTPHALALRMILQLGVKPSWFNTCSTTIDKRPHLIPGTASFKGDLRNYIIDLPPVGDSVVPEMLLKVLDPYRREGCILDDETARLYAI
VVKKGCAARFAFAAAIFGETSEALFWLQLPRALKHLMDKLVTKSTQKAPVSASTPELDDVTMLNRISSKGRSVIGTEKKDPLSEGQLRSMAFQKEELWES
ACERIPWHEKLEGEEAIQNRVHELVSIGNLEAAVSLLLSTSPESSYFYVNALRAVALSSAVSRSLHELAVKVVAANMVQTDRSLSGTHLLCAVGRYQEAC
SQLQDAGCWTDAATLAATHLSGSDYARVLLRWANHVLHAEHNIWRALILYVAAGALQDALAALRETQQPDTAAMFILACHEGHAQFISNLGNSDDESGSS
IKDTLVGLPGLNPENEDVIAVGEYYGQYQRKLVHLCMDSQPFSD
|
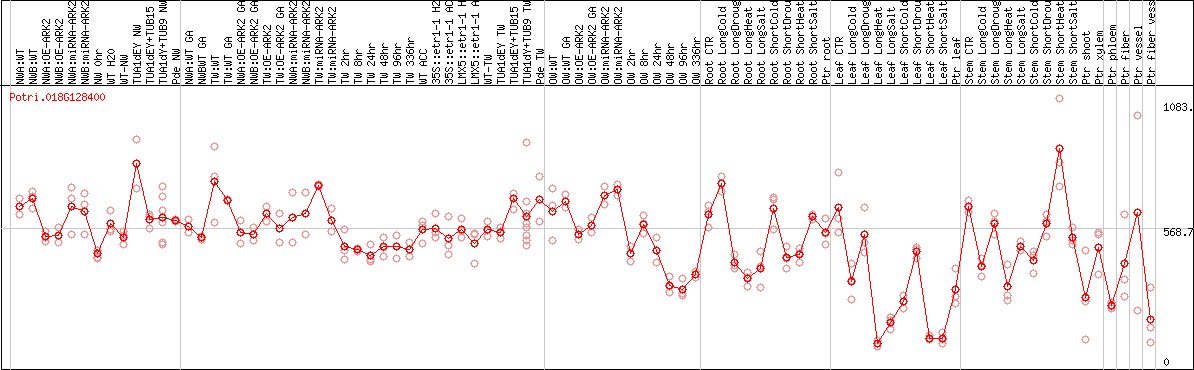
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.018G128400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
