External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G02630 212 / 4e-68
Nucleoside transporter family protein (.1.2)
AT1G70330 163 / 1e-48
"ENT1,AT", ENT1,AT, ATENT1
equilibrative nucleotide transporter 1 (.1)
AT1G61630 60 / 6e-11
ATENT7
equilibrative nucleoside transporter 7 (.1)
AT4G05120 60 / 7e-11
FUR1, ENT3, FLUOROURIDINEINSENSITIVE1, ATENT3
FUDR RESISTANT 1, EQUILIBRATIVE NUCLEOSIDE TRANSPORTER 3, Major facilitator superfamily protein (.1)
AT3G09990 58 / 3e-10
Nucleoside transporter family protein (.1)
AT4G05110 58 / 3e-10
ATENT6
equilibrative nucleoside transporter 6 (.1)
AT4G05130 57 / 9e-10
ATENT4
equilibrative nucleoside transporter 4 (.1)
AT4G05140 51 / 1e-07
Nucleoside transporter family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G130000
314 / 5e-108
AT1G02630 452 / 5e-159
Nucleoside transporter family protein (.1.2)
Potri.006G068100
259 / 3e-86
AT1G02630 432 / 6e-151
Nucleoside transporter family protein (.1.2)
Potri.012G058900
160 / 1e-47
AT1G70330 530 / 0.0
equilibrative nucleotide transporter 1 (.1)
Potri.019G118400
64 / 2e-12
AT4G05120 520 / 0.0
FUDR RESISTANT 1, EQUILIBRATIVE NUCLEOSIDE TRANSPORTER 3, Major facilitator superfamily protein (.1)
Potri.004G032300
60 / 7e-11
AT4G05120 586 / 0.0
FUDR RESISTANT 1, EQUILIBRATIVE NUCLEOSIDE TRANSPORTER 3, Major facilitator superfamily protein (.1)
Potri.004G032400
57 / 8e-10
AT4G05120 584 / 0.0
FUDR RESISTANT 1, EQUILIBRATIVE NUCLEOSIDE TRANSPORTER 3, Major facilitator superfamily protein (.1)
Potri.004G032500
57 / 1e-09
AT4G05120 457 / 2e-160
FUDR RESISTANT 1, EQUILIBRATIVE NUCLEOSIDE TRANSPORTER 3, Major facilitator superfamily protein (.1)
Potri.011G041112
56 / 2e-09
AT4G05120 540 / 0.0
FUDR RESISTANT 1, EQUILIBRATIVE NUCLEOSIDE TRANSPORTER 3, Major facilitator superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10030104
234 / 2e-76
AT1G02630 526 / 0.0
Nucleoside transporter family protein (.1.2)
Lus10029162
167 / 2e-50
AT1G70330 551 / 0.0
equilibrative nucleotide transporter 1 (.1)
Lus10013002
159 / 4e-47
AT1G70330 534 / 0.0
equilibrative nucleotide transporter 1 (.1)
Lus10018413
71 / 1e-14
AT4G05120 606 / 0.0
FUDR RESISTANT 1, EQUILIBRATIVE NUCLEOSIDE TRANSPORTER 3, Major facilitator superfamily protein (.1)
Lus10002589
71 / 2e-14
AT4G05120 609 / 0.0
FUDR RESISTANT 1, EQUILIBRATIVE NUCLEOSIDE TRANSPORTER 3, Major facilitator superfamily protein (.1)
Lus10002167
64 / 5e-14
AT1G02630 95 / 3e-25
Nucleoside transporter family protein (.1.2)
Lus10018523
64 / 3e-12
AT4G05120 579 / 0.0
FUDR RESISTANT 1, EQUILIBRATIVE NUCLEOSIDE TRANSPORTER 3, Major facilitator superfamily protein (.1)
Lus10039741
64 / 3e-12
AT4G05120 580 / 0.0
FUDR RESISTANT 1, EQUILIBRATIVE NUCLEOSIDE TRANSPORTER 3, Major facilitator superfamily protein (.1)
Lus10020079
62 / 1e-11
AT4G05120 392 / 7e-135
FUDR RESISTANT 1, EQUILIBRATIVE NUCLEOSIDE TRANSPORTER 3, Major facilitator superfamily protein (.1)
Lus10018414
55 / 4e-09
AT4G05120 548 / 0.0
FUDR RESISTANT 1, EQUILIBRATIVE NUCLEOSIDE TRANSPORTER 3, Major facilitator superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0015
MFS
PF01733
Nucleoside_tran
Nucleoside transporter
Representative CDS sequence
>Potri.018G130200.1 pacid=42802091 polypeptide=Potri.018G130200.1.p locus=Potri.018G130200 ID=Potri.018G130200.1.v4.1 annot-version=v4.1
ATGGAACAGCGTTACAAACTTGCTCCTGATGATTCACTGTTTCCAAAACCAAAATTTCGGGCTGTAGCAAGAAAAATCCGATGGCCAGCTTTTGGTGTTC
TCATGATTTATATCGTGACTTTGTCGATATTTCCTGGTTTCATAGAAGATCTCTCGTCCAAGTTACTAAAAGATTGGTATCGCGTTTTGCTGATAACAAT
ATACAATGTTGCAGACTTCACAGGGAAATCTTTAACCGCAATATATGTTCTGCAGAGCATTAAGAAGGCAACATGGGGTTGCATTCTCAGGCTTGTGTTC
TATCCTCTCTTTGCAGCTTGTCTCAATGGACCCAAGTGGCTGAAAACTGAAGTACCCGTGGCAATTCTAACTTTCATGCTCGGAGTGACTAATGGCTATC
TGACAAGTGTTCTCATGATCCTTGCCCCCATGGCAGTATCAGTTTCAGAAGCAGAGTTATCTGCAATTGCAATGGTTGTGTTCCTAGGAATAGGGTTGGT
TGGTGGATCAGTTATTGGCTGGTTTTGGATCATTTGA
AA sequence
>Potri.018G130200.1 pacid=42802091 polypeptide=Potri.018G130200.1.p locus=Potri.018G130200 ID=Potri.018G130200.1.v4.1 annot-version=v4.1
MEQRYKLAPDDSLFPKPKFRAVARKIRWPAFGVLMIYIVTLSIFPGFIEDLSSKLLKDWYRVLLITIYNVADFTGKSLTAIYVLQSIKKATWGCILRLVF
YPLFAACLNGPKWLKTEVPVAILTFMLGVTNGYLTSVLMILAPMAVSVSEAELSAIAMVVFLGIGLVGGSVIGWFWII
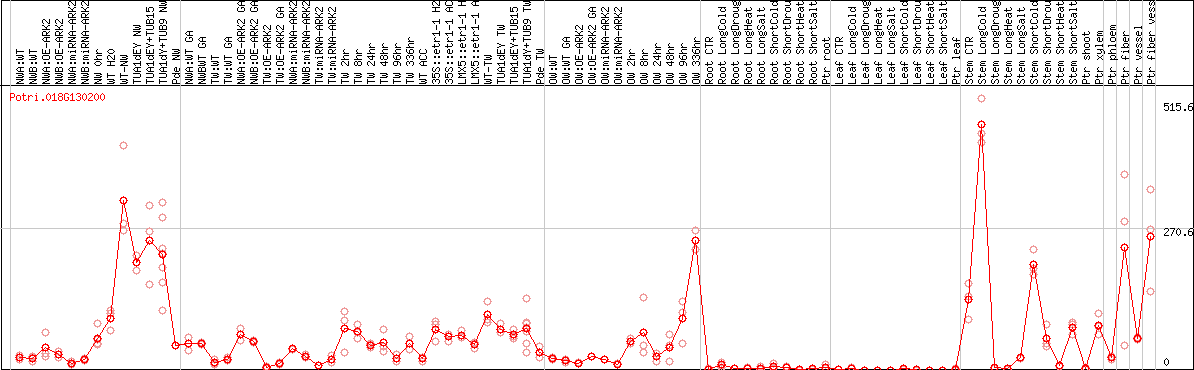
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.018G130200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.