Pt-ACA8.6 (Potri.018G139800) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | Pt-ACA8.6 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.018G139800.8 pacid=42801615 polypeptide=Potri.018G139800.8.p locus=Potri.018G139800 ID=Potri.018G139800.8.v4.1 annot-version=v4.1
ATGACAAGTTTGTTTAAAGGCTCGCCGTGTATACGCCAGCAAGATGATTTGGAAGCTGGTGAAAATCGTTCTACTGACGTCGGCCGTGATGCCAACTCTT
CGTCTGGTCCTTTTGATATCGTCAGCACCAAAAACGCTCCCATCGACAGCCTCCGCAGATGGCGGAAAGCTGCGCTCGTGCTTAATGCTTCTAGAAGATT
CCGATATACTCTGGACTTGAAGAAGGAAGAGGAGAAGAGGAGAATATTAAGCAAGATAAGAGCACATGCGCAAGTGATTTGGGCTGCACATCTTTTCAAG
GAAGCTGGGAATAATAGAGTAAATGGAGACACAGAACCACATCCACCCCCCACTGGTGATTTTGGAATTAGCGTGGGGCAAATTTCTGTAATCACGAGGG
ATCATGATCATAATGCTTTGGAGGCACTCGGTGGGGTAAAAGGGGTTGCAGATGCATTGAAAACTGATATAGAGAAGGGAATTCATGAAGATGATGCTGA
TTTATTGAAACGGAAGAATGCATTTGGATCAAATACATATCCTCAGAAAAAAGGAAGGAGTTTTTGGATGTTCCTTTGGGAAGCTTGGCAAGATCTTACT
TTGATCATATTGATGGTAGCTGCAGTGGCCTCTTTGGTGCTGGGTATGAAGACAGAGGGTGTTAAGGAAGGATGGTATGAAGGGGCCAGCATTGCTTTTG
CAGTTATTCTTGTCATTGTTGTGACAGCTATAAGTGACTACAAACAATCTCTTCAGTTTCAAAATTTAAACGAGGAGAAGAGAAACATACATTTGGAGGT
TACCAGAGGAGGTAGAAGAGTTGAAGTTTCAATATATGATATTGTTGCTGGTGATGTTATACCCCTTAACATTGGTGATCAGGTTCCTGCTGATGGAATT
TTAATTACTGGTCACTCCCTTGCTATTGATGAATCAAGCATGACTGGAGAAAGCAAGATTGTCCAAAAGAATTCCAGGGAACCGTTTCTAATGTCTGGCT
GCAAAGTTGCAGATGGTAGTGGCACTATGCTGGTAACAGGTGTTGGAATTAATACTGAATGGGGGTTGCTCATGGCTAGTATATCAGAAGACAATGGTGA
AGAAACACCTTTGCAGGTGCGCTTGAATGGGGTCGCAACTTTCATTGGTATTGTGGGGCTGACAGTAGCTTTGCTTGTCTTGGTAGTCCTCTTGGTCAGA
TATTTCACTGGACATACAAAAAACTTTGATGGAAGTCCTGAGTTCGTAGCGGGTAAAACAAAAGTTAGTAAAGCAGTAGATGGAGCCGTTAAAATTCTGA
CTGTCGCGGTCACCATTGTTGTAGTTGCAGTGCCTGAAGGGCTTCCCTTAGCAGTTACTTTAACTCTTGCATACTCGATGAGAAAAATGATGAGAGATAA
GGCTTTGGTGCGCCGGCTTTCTGCTTGTGAAACCATGGGCTCTGCCACCACTATTTGCAGTGATAAGACTGGAACTTTGACCTTGAATCAGATGACCGTT
GTGGAAGCTTTTTCTGGAGGGAAAAAAATGGATCTTCCTGAAAGTAAATCACAATTGCCTCCTATTTTGTCTTCTTTACTTATTGAAGGTATTGCACAAA
ACACAACCGGCAGTGTTTTTGTTCCTGAGGGAGGTGGAGATCTAGAGATTTCTGGATCGCCAACAGAAAAGGCCATTATGGGATGGGCAATCAAGCTGGG
GATGAATTTTGATGCTGTCCGATCTGAATCGAATGTCATTCATGTATTCCCATTCAACTCTGAGAAGAAAAAAGGCGGGGTGGCACTACAGCTGCCCAAC
TCTCAAGTGCATATACATTGGAAGGGGGCTGCCGAAATAGTCTTAGCCTCATGTACAAAATATGTTGATGCAAGTGGTAACACAGTGCCATTGGATCAAG
ACAAGGTGTCGTTTTTTAAGAAAGCTATTGAAGATATGGCTTGCAGTAGTTTGCGTTGTGTTTCCATTGCGTATAGAACATATGACATGGACAAAGTTCC
AGCTGATGAACAACAACTAGCTCAATGGGTGATTCCTCAAGATGATCTTGTTTTGCTTGCAATTATTGGCATAAAGGACCCATGTCGTCCAGGTGTGAGA
GATGCTGTTCGACTGTGCCAAAATGCTGGTGTTAAGGTTCGCATGGTAACTGGTGACAATCCTCAAACTGCTAAAGCAATTGCTTTGGAATGTGGGATAC
TGAGCTCAGAAGAAGATGCTGTGGAGCCTAATGTTATTGAAGGAAGAGTGTTTCGTGAATATTCAGATTCAGAAAGAGAAGACATCGCAGAGAAGATCTC
AGTGATGGGGAGGTCTTCTCCGAATGACAAGCTTTTGCTTGTGCAAGCATTGAAAAGGAGGGGACATGTTGTTGCTGTAACTGGAGATGGAACAAATGAT
GCTCCTGCATTGCATGAGGCAGACATTGGTCTCTCAATGGGTATTCAAGGCACAGAAGTTGCTAAAGAGAGCTCAGATATCATAATTTTGGACGACAATT
TTGCTTCAGTCGTGAAGGTTGTCCGTTGGGGTAGATCTGTATATGCAAATATCCAGAAATTTATTCAGTTTCAGCTTACAGTTAATGTTGCAGCTCTCAT
TATAAATGTTGTGTCTGCAATGTCTTCTGGTGAAGTCCCATTGAATGCAGTGCAGCTCCTTTGGGTTAATCTTATCATGGATACTCTTGGAGCACTGGCA
CTGGCTACTGAGCCTCCAACAGATCACCTGATGAATAGATCTCCAGTTGGTCGCCGGGAACCACTAATAACAAATATCATGTGGAGGAACCTGCTGGTAC
AGGCAGCATATCAAGTGACCGTGCTCCTTGTCCTTAATTTTCGAGGAGAGAGTATACTAGGTTTGGAGCATGAGACCCCTCAGCGTGCGATCGAAGTGAA
GAATACCTTGATATTCAATGCATTTGTGCTCTGTCAGATCTTCAACGAATTTAATGCCCGGAAACCAGATGAAATCAATATTTTCAAAGGGATTTCAAAA
AACCACCTATTCATTGCTATAATTGGAATAACACTTGTACTTCAGGTAATCATAGTTGAGTTCGTTGGAAAGTTTACTTCTACCGTCAAGCTCAATTGGA
AACAGTGGCTGATTTCAATTATTATTGGTTTTATCGGCTGGCCGCTTGCTGCTCTCGCTAAACTGATACCTGTTCCACAAACTCCTTTGCACAAGTTCTT
TACAAATATGTGTAATCGACGTGCCAAGTCTTCCAAGTCTTCCAAGTCTTCCAAGTCATCATCAGTAGAGGTCACTAACAATTCACAGCACTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.018G139800.8 pacid=42801615 polypeptide=Potri.018G139800.8.p locus=Potri.018G139800 ID=Potri.018G139800.8.v4.1 annot-version=v4.1
MTSLFKGSPCIRQQDDLEAGENRSTDVGRDANSSSGPFDIVSTKNAPIDSLRRWRKAALVLNASRRFRYTLDLKKEEEKRRILSKIRAHAQVIWAAHLFK
EAGNNRVNGDTEPHPPPTGDFGISVGQISVITRDHDHNALEALGGVKGVADALKTDIEKGIHEDDADLLKRKNAFGSNTYPQKKGRSFWMFLWEAWQDLT
LIILMVAAVASLVLGMKTEGVKEGWYEGASIAFAVILVIVVTAISDYKQSLQFQNLNEEKRNIHLEVTRGGRRVEVSIYDIVAGDVIPLNIGDQVPADGI
LITGHSLAIDESSMTGESKIVQKNSREPFLMSGCKVADGSGTMLVTGVGINTEWGLLMASISEDNGEETPLQVRLNGVATFIGIVGLTVALLVLVVLLVR
YFTGHTKNFDGSPEFVAGKTKVSKAVDGAVKILTVAVTIVVVAVPEGLPLAVTLTLAYSMRKMMRDKALVRRLSACETMGSATTICSDKTGTLTLNQMTV
VEAFSGGKKMDLPESKSQLPPILSSLLIEGIAQNTTGSVFVPEGGGDLEISGSPTEKAIMGWAIKLGMNFDAVRSESNVIHVFPFNSEKKKGGVALQLPN
SQVHIHWKGAAEIVLASCTKYVDASGNTVPLDQDKVSFFKKAIEDMACSSLRCVSIAYRTYDMDKVPADEQQLAQWVIPQDDLVLLAIIGIKDPCRPGVR
DAVRLCQNAGVKVRMVTGDNPQTAKAIALECGILSSEEDAVEPNVIEGRVFREYSDSEREDIAEKISVMGRSSPNDKLLLVQALKRRGHVVAVTGDGTND
APALHEADIGLSMGIQGTEVAKESSDIIILDDNFASVVKVVRWGRSVYANIQKFIQFQLTVNVAALIINVVSAMSSGEVPLNAVQLLWVNLIMDTLGALA
LATEPPTDHLMNRSPVGRREPLITNIMWRNLLVQAAYQVTVLLVLNFRGESILGLEHETPQRAIEVKNTLIFNAFVLCQIFNEFNARKPDEINIFKGISK
NHLFIAIIGITLVLQVIIVEFVGKFTSTVKLNWKQWLISIIIGFIGWPLAALAKLIPVPQTPLHKFFTNMCNRRAKSSKSSKSSKSSSVEVTNNSQH
|
DESeq2's median of ratios [POPLAR]
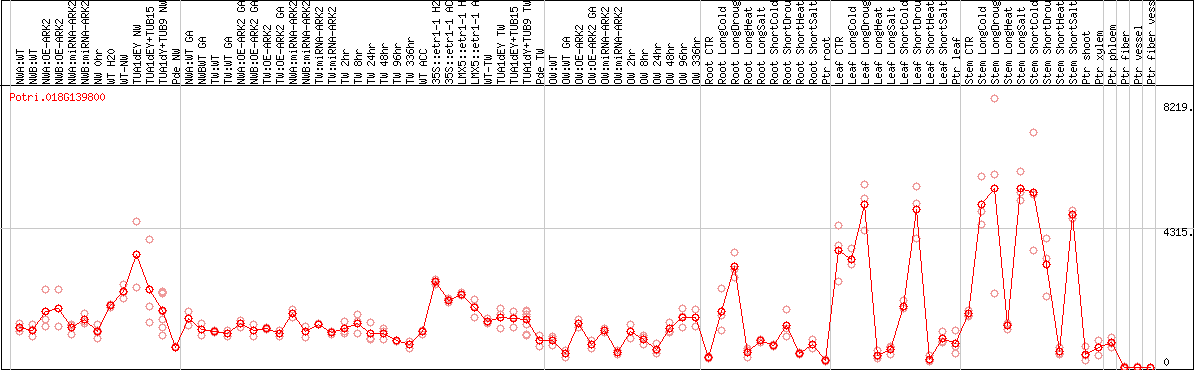
Coexpressed genes
Potri.018G139800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
