Potri.018G149400 [POPLAR]

| External link |
|
||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||
|
Representative CDS sequence |
>Potri.018G149400.1 pacid=42801246 polypeptide=Potri.018G149400.1.p locus=Potri.018G149400 ID=Potri.018G149400.1.v4.1 annot-version=v4.1
ATGGCGGATTCTTCGTCGGGGACGACGTTGATGGATCTGATAACAGCGGACCCCGGGCCGGCTCCGAAGTCATCGGGATCCTCGGAAGCGCCAACTCCTC
CGGCATCGCAGCAACCGACTGGATCGATGTCATATTCCACGCCGACGACGACGACGGCAAGTTCGTCCTCGGGGTCAGGGAAGACAATGCTAGGAGAGAG
AAAGTCGAAGAGAGCTACACTTATGCAGATCCAAAACGACACCATTTCTGCTGCTAAAGCTGCCATGAAAACTACTGCCGGTATCAATATCATGCCTCAA
AAACAAAAGAAGAATCCGGTTTCGTATTCGCAACTAGCTAGGAGTATTCATGAACTAGCTGCTACATCTGATCAAAAAAGTTCTCAAAAGCAACTAGTGC
ATCATGTATTTCCGAAACTTGCTGTTTACAATTCAGTCGATCCGTCGTTGGCACCGTCTCTTCTCATGCTGGATCAGCAATGTGAAGACAGGACGATTCT
GCGCTATGTGTATTATTACTTGGCTAGGATTTTATCAGATACTGGTTCTCAAGGTTTAAATCCAGGTGGTGGAATACCGACTCCTAATTGGGATGCTTTG
GCTGATATTGATGCTGTTGGAGGGGTGACTAGAGCTGATGTTGTACCAAGGATTGTAGATCAGCTTTCGAAAGAGGCTTCAGATGCTAATGTTGAATTTC
ATGCACGGAGATTGCAAGCATTGAAGGCTCTTACATATGCTCCCGAAAGCAACACTGGTATTTTATCCAGATTGTATGAGATTGTATTTGGCATTCTTGA
CAAGGTTGGTGACAACCCTCAAAAACGAAAGAAAGGTGTTTTTGGGACTAAAGGTGGTGATAAGGAGTCTATCGTTCGGAGTAATTTGCAATATGCTGCA
CTAAGTGCATTAAGAAGACTCCCTCTTGATCCTGGAAACCCAGCATTTCTGCACCGTGCTGTACAGGGGGTTTCATTTGCTGATCCTGTTGCTGTAAGGC
ATGCTTTGGAAATTCTCTCTGAGTTGGCAACAAAAGATCCTTATGGAGTTGCAATGGCTTTAGGAAAGCTTGTAGTCCCTGGAGGGGCATTGCAGGATGT
TCTTCATTTGCATGATGTGCTTGCTAGAGTTTCGCTGGCCAGGTTGTGTCATACAATATCTAGAGCTCGTGCATTAGATGAGAGGCCTGACATTAAATCT
CAGTTCAACTCAGTGCTTTATCAACTTCTCCTAGATCCCAGTGAAAGAGTCTGTTTTGAGGCAATCTTTTGTGTCTTGGGAAAACACGATAATACAGAGA
GGACTGAGGAGCGTGCTGCTGGCTGGTACCGTTTAACAAGGGAGATACTGAAGTTGCCAGAAGCACCCTCTCTATCATCGAAGGGATCTATTGCTGATTC
AAATGATATGTCAAAGGCTTCTAAAGATAAATCCCACAAGACTAGGCGTCCACAACCTCTTATTAAACTTGTAATGAGAAGGTTGGAAAGTTCTTTTCGT
AATTTCTCCAGGCCAGTACTTCATGCAGCAGCAAGAGTTGTGCAGGAGATGGGAAAAAGTCGCGCTGCTGCTTATGCTGTAGGCTTACAGGACATCGATG
AAGGAGTTAATGTGAATTCATTTTCTGAGAGTGCAGATCCAGTCGATTCGGATTTTAATGAAAATCCATATGCAGATGGTGCCCGGAAGGTCTCTGCTGT
ATCCAGTGCAACAGGTAGCAAGGATACAATTGCAGGTTTATTAGCTTCATTAATGGAGGTGGTACGGACAACTGTAGCCTGTGAATGTGTCTATGTTCGA
GCAATGGTAATCAAGGCTTTGATATGGATGCAACTTCCACATGAATCTTTTGAGGAACTGGAATCCATTATCGCATCCGAGCTTTCTGATCCATCATGGC
CAGCAACATTGTTGAATGATGTTTTGCTGACTTTGCATGCTCGTTTCAAGGCAACACCAGATATGGCTGTCACCCTGCTTGAAATTGCGAGGATCTTTGC
TACTAAGGTTCCAGGAAAGATTGATGCTGATGTGTTGCAGCTTTTGTGGAAAACGTGCCTTGTTGGTGCCGGTCCTGATGGGAAGCACACAGCATTGGAA
GCAGTCACTATAGTCCTTGACCTACCACCACCACAGCCTGGGTCAATGTTAGGGCTTACATCAGTGGACAGGGTATCTGCATCTGATCCGAAGTCAGCTT
TGGCATTGCAGAGATTAGTCCAAGCAGCAGTGTGGTTCCTTGGAGAAAATGCAAATTATGCTGCTTCCGAATATGCTTGGGAATCTGCAACGCCTCCTGG
CACTGCATTGATGATGTTAGATGCAGATAAAATGGTTGCTGCTGCCAGTTCTCGTAATCCTACTCTAGCTGGTGCTTTGACTCGACTACAGAGGTGTGCT
TTCAGTGGCAGCTGGGAGGTTCGAATTGTTGCTGCTCAAGCCCTCACAACTATGGCAATCAGGTCTGGTGAGCCCTTTAGGCTGCAGATCTATGAATTTT
TAAATGCTTTAGCACAGGGTGGAGTGCAGTCCCAGCTATCAGAAATGCATCTTAGTAATGGGGAAGATCAAGGAGCCAGCGGTACAGGCCTTGGAGTCTT
AATAAGTCCCATGGTAAAGGTCCTTGATGAAATGTACAGAGCACAAGATGAACTGATCAGAGATATAAGAAACCATGACAATACAAATAAAGAATGGACA
GATGAAGAACTGAAGAAACTTTATGAAACGCATGAGAGATTGTTAGATATTGTTTCATTGTTTTGTTATGTTCCAAGAGCAAAGTATCTACCTTTGGGAC
CAATAAGTGCTAAGCTTATTGATATTTATCGCACCAAGCACAACATTAGTGCATCAACTGGCTTAAGTGATCCAGCTGTTGCCACTGGAATTTCTGACCT
AATGTATGAATCAAAGCCGGCCCCTGTTGAGTCTGATGCGCTTGATGATGACCTAGTAAATGCTTGGGCAGCAAACCTTGGTGATGATGGTTTATTGGGA
AATAGCGCACCTGCAATGAGTAGGGTCAACGAATTTCTTGCTGGTATGGGAACTGAAGCTCCGGATGTTGAAGAGGAAAATATAATATCTAGACCTTCAG
TCAGCTATGATGATATGTGGGCAAAGACACTTTTGGAGTCTTCAGAGTTAGAGGAAGACGTGAGGTCATCTGGGTCCTCATCCCCAGACTCAATAGGGTC
AGTTGAAACTTCTATATCGTCCCACTTTGGTGGAATGAACTACCCATCACTGTTCAGTTCTCGGCCCACCAGTTATGGAGCATCACAAATATCGGAGAGG
TCAGGTGGAAACAGGTATAGCGGTCCCTCGTCCTTTTATGAGGGTGCAGGTTCTCCGATTAGGGAAGAGCCCCCGCCATACACATCACCTGACAGGTCAT
TTGAAAACCCCTTGGCTGGGCATGGGTCCCGGAGTTTTGAATCACAGGAATCAGGACGAGCATCATCTGCAAATCCACAATATGGCTCTGCACTCTATGA
CTTCAGTGCGGGTGGAGATGATGAGTTGAGTTTAACAGCAGGAGAAGAGCTTGAAATTGAGTACGAGGTTGATGGATGGTTTTATGTCAAGAAGAAACGC
CCTGGCAGGGATGGCAAAATGGCCGGACTAGTTCCTGTACTCTATGTCAATCAAAGTTAA
|
||||||||||||||||||||
|
AA sequence
|
>Potri.018G149400.1 pacid=42801246 polypeptide=Potri.018G149400.1.p locus=Potri.018G149400 ID=Potri.018G149400.1.v4.1 annot-version=v4.1
MADSSSGTTLMDLITADPGPAPKSSGSSEAPTPPASQQPTGSMSYSTPTTTTASSSSGSGKTMLGERKSKRATLMQIQNDTISAAKAAMKTTAGINIMPQ
KQKKNPVSYSQLARSIHELAATSDQKSSQKQLVHHVFPKLAVYNSVDPSLAPSLLMLDQQCEDRTILRYVYYYLARILSDTGSQGLNPGGGIPTPNWDAL
ADIDAVGGVTRADVVPRIVDQLSKEASDANVEFHARRLQALKALTYAPESNTGILSRLYEIVFGILDKVGDNPQKRKKGVFGTKGGDKESIVRSNLQYAA
LSALRRLPLDPGNPAFLHRAVQGVSFADPVAVRHALEILSELATKDPYGVAMALGKLVVPGGALQDVLHLHDVLARVSLARLCHTISRARALDERPDIKS
QFNSVLYQLLLDPSERVCFEAIFCVLGKHDNTERTEERAAGWYRLTREILKLPEAPSLSSKGSIADSNDMSKASKDKSHKTRRPQPLIKLVMRRLESSFR
NFSRPVLHAAARVVQEMGKSRAAAYAVGLQDIDEGVNVNSFSESADPVDSDFNENPYADGARKVSAVSSATGSKDTIAGLLASLMEVVRTTVACECVYVR
AMVIKALIWMQLPHESFEELESIIASELSDPSWPATLLNDVLLTLHARFKATPDMAVTLLEIARIFATKVPGKIDADVLQLLWKTCLVGAGPDGKHTALE
AVTIVLDLPPPQPGSMLGLTSVDRVSASDPKSALALQRLVQAAVWFLGENANYAASEYAWESATPPGTALMMLDADKMVAAASSRNPTLAGALTRLQRCA
FSGSWEVRIVAAQALTTMAIRSGEPFRLQIYEFLNALAQGGVQSQLSEMHLSNGEDQGASGTGLGVLISPMVKVLDEMYRAQDELIRDIRNHDNTNKEWT
DEELKKLYETHERLLDIVSLFCYVPRAKYLPLGPISAKLIDIYRTKHNISASTGLSDPAVATGISDLMYESKPAPVESDALDDDLVNAWAANLGDDGLLG
NSAPAMSRVNEFLAGMGTEAPDVEEENIISRPSVSYDDMWAKTLLESSELEEDVRSSGSSSPDSIGSVETSISSHFGGMNYPSLFSSRPTSYGASQISER
SGGNRYSGPSSFYEGAGSPIREEPPPYTSPDRSFENPLAGHGSRSFESQESGRASSANPQYGSALYDFSAGGDDELSLTAGEELEIEYEVDGWFYVKKKR
PGRDGKMAGLVPVLYVNQS
|
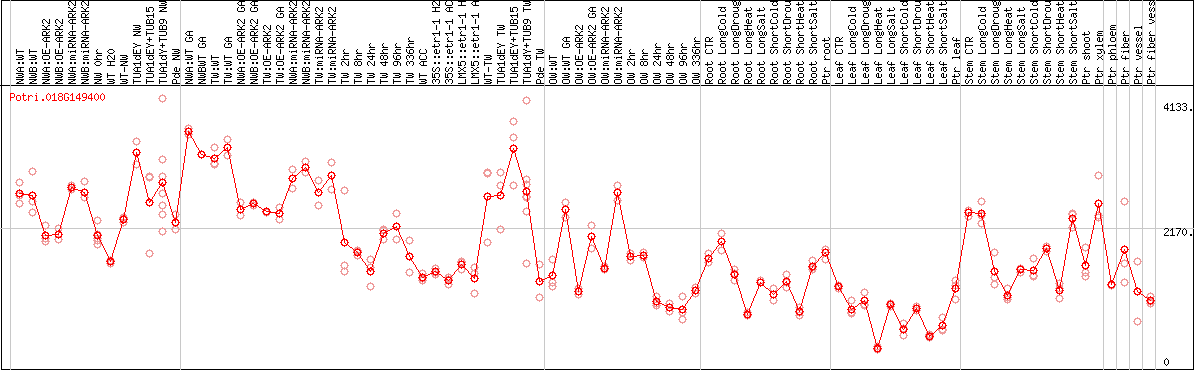
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.018G149400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
