Potri.019G011826 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.019G011826.1 pacid=42774084 polypeptide=Potri.019G011826.1.p locus=Potri.019G011826 ID=Potri.019G011826.1.v4.1 annot-version=v4.1
ATGAGTCGACCAAATGATCCTCCTTTTTGGGAATATGTTGAAAAGATGGATGGTGGTGGCATGACGTGTACATTTTGTGGGCATTTTTTTTCCGAAGGTA
CTTCCATTACGAGGATCAAATTGCATTTGGCAAGAAAGAAAGGGCGTGGTGTTAAAATTTGTGAAAACGTTCCACCAGAAGTTCATGATGCAGCTTGTGA
AGCAGCAGTTGATAATAGCCCTCCAAAAAAGAAACTTAAAACTGTTGCGGGCTCAAGCAGTAACGAGGCCGCTAATGCAATTTCAGCTTCAACGCAAGAA
CAGAACAATGAAGTGACGCATGTAGAGATGGCACAACAAGGAGGACCTTTTTTCACTGGAGAACTTGCATGGGCAAATGATTTGATTGGAGCGGCTGAAG
TTGTGCAGCTGGAGAGAGGTAGTTCTCATGAGCCTCGAGGAGATTCATCCCGACCAACAGATGATCAACTCTGTTCTCCATCAGTAAACAATGATGCCAA
TATGAACGATGTGCAGAACATGGTTGCAGTCAGGATAGAACCAGTACCGGTTCAAGTGTTGGAACAAAGCAATGCAGAACTTGACAGTTTCGCAGGACAT
GCTGGAAGGATACAAGTGGGAGTTCATGGCATGGAACAAGGTGCGGAGGAAGAGAGAATTTGTCCGAACTCAGCGGAAAATGGCATGGAAAACACTGGTG
ATGGATCTAGTCAGCATTCCAAGAGACTCATAATTGATGCACATTATAATACAGGGGAAGCAACACAAGGAATAGATCTAGCGCAACAATTAGAAGGGAA
AACTTGGGATCAGATTAATGCAATCGCTACTTCATTAATGGGGGAGGAGGACGTGGAGAACAATAGTGGAAGATCAGAGCAACCTGGCGCAGGAGCTAGC
TCTTCTGGAGGGGTTGCAGGCAACACAAATAAGATTAAAGGAGATGCACTGCCAACGAGAAAAATGGTGGGTCAAGCATTCGAAGAACACAAGAAGACTA
TCTCATCTTTGTTGATGCGCAATGAAGTTTCAAGTATTGGCATCTATGGGATGGGCGGAGTGGGAAAGACGGCATTGGTGACACATATACACAATCAGCT
TCTAGAAAGACGAGACACTCATGTTTACTGGATTACTGTTTCACAAAATACCAGCATTCATAGATTGCAGACAAGTCTCGCAGGACGAATTGGTTTAGAT
CTTTCCAAGGTAGATGAGGAGGTGCATAGAGCGGTAGCGTTGAATAAAGAATTAATGAAGAAACAGAAATGGGTTCTGATTTTAGATGATTTGTGGAAGG
CTTTTGACCTCCAAAAGCTGGGAGTTCCTGACCAAGTTGAGGGATGCAAGCTGATTCTTACATCTCGATCAGCAAAGGTTTGTCAACAGATGAAAACCCA
ACACACCATCAAAGTGCAGCCTATTTCGGAGAGAGAAGCTTGGACTTTGTTTATTGAGAGACTTGGACATGACAGAGAACTTTCTCCAAAAGTGAAACGA
ATTGCTGTGGAAGTTGTAAGGGAATGTGCTGGTTTGCCATTGGGAATTATTACGATGGCAGGAAGCATGAGGGGAGTGGATGAGCCACATGAGTGGAGGA
ATACATTGAATAAGTTGAAAGGATCAAAATACAGAGACATGGAAGATGACGTATTCCGGTTGCTGAGGATTAGTTATGATCAGTTGGATAATGATTTAGC
ATTACAACAATGTCTCTTATATTGTGCATTGTATCCTGAAGATTATCAGATTGAAAGGGAGGAGCTGATAGGTTATTTGATAGATGAGGGAATAATTGAA
GAAATGAGGAGCAGGCAAGCAGCATTTGACGAGGGCCACACGATGCTTGATAAACTTGAAAAAGTCTGTTTATTGGAAAGAGCTTGTTATGGTGATCATA
ATACAAGTGTCAAGATGCATGACTTGATTAGGGACATGGCCCACCAAATACTCCAAACAAACTCTCCAGTCATGGTTGGGGGATATTATGATGAATTACC
TGTGGATATGTGGAAAGAGAATCTAGTGAGAGTCTCTCTGAAGCATTGTTATTTCAAGGAAATTCCTTCTAGTCATTCACCAAGATGTCCCAATCTTTCA
ACTCTATTGCTGTGTGATAATGGACAGTTGAAATTTATAGAAGATTCTTTTTTCCAACATTTGCATGGGCTCAAGGTTCTTGATCTGTCTCGTACAGACA
TCATAGAATTGCCTGGTTCTGTCTCTGAACTGGTGAGTCTCACTGCATTATTGCTCGAAGAATGTGAGAATTTAAGGCATGTACCATCATTAGAAAAGCT
CAGAGCACTGAAGAGGCTAGATCTCTCTGGTACTTGGGCACTTGAAAAGATACCTCAAGACATGCAATGTCTATCCAACCTGAGGTATCTGAGGATGAAT
GGATGTGGTGAAATGGAGTTTCCTAGTGGGATATTACCAATACTCTCTCACCTGCAAGTCTTCATATTAGAGGAGATTGATGATGACTTCATTCCAGTAA
CAGTTACAGGAGAGGAAGTAGGATGCTTGAGGGAGTTGGAAAATTTGGTATGCCATTTTGAAGGTCAATCTGACTTTGTGGAGTATCTCAATTCTCGAGA
TAAGACTCGATCACTAAGTACATACAGTATTTTCGTAGGACCACTGGATGAATATTGTAGTGAAATAGCTGATCATGGCGGAAGTAAAACAGTTTGGTTG
GGTAATTTGTGTAACAATGGAGATGGAGATTTTCAAGTCATGTTCCCAAATGACATTCAAGAACTGTTTATTTTTAAATGTTCATGTGATGTTTCCTCTC
TAATAGAGCATTCAATAGAACTGGAGGTCATCCACATTGAGGATTGCAATAGCATGGAGAGCTTGATTTCATCTTCTTGGTTCTGCCCTTCTCCAACACC
ATTGTCATCTTATAATGGTGTATTTTCTGGTCTTAAAGAGTTCAATTGCTCTGGATGTAGTAGTATGAAGAAGTTGTTCCCTCTTGTCTTGCTACCAAAC
CTCGTAAACCTAGAAAATATTTCAGTTTTTGGTTGTGAGAAAATGGAGGAGATAATAGTTGGAACAAGATCAGATGAAGAAAGCAGCAGCAACAGCACCG
AATTCAAACTACCAAAGTTGAGATATCTGGCATTGGAAGATTTACCAGAACTGAAAAGAATTTGTAGTGCAAAATTGATTTGCGATTCTCTCCAACAAAT
TGAAGTAAGGAATTGCAAAAGCATGGAGAGCTTGGTTCCATCTTCTTGGATTTGCCTCGTAAACCTAGAAAGGATTATAGTTACAGGATGTGGGAAAATG
GAGGAGATAATAGGTGGAACAAGAGCAGATGAAGAAAGCAGCAACAACACCGAATTCAAACTCCCAAAGTTAAGATCGCTGGAATCGGTAGATTTACCAG
AACTGAAAAGAATTTGTAGTGCAAAATTGATTTGCGATTCTCTCCGAGAAATTGAAGTAAGGAATTGCAATAGCATGGAGATCTTGGTTCCATCTTCTTG
GATTTGCCTCGTAAACCTAGAAAGGATTATAGTTGCAGGATGTGGGAAAATGGATGAGATAATATGCGGAACAAGATCAGATGAAGAAGGGGATATAGGT
GAAGAAAGCAGCAACAACAACACCGAATTCAAACTCCCAAAGTTAAGATCTCTGCTATTGTTTGAATTACCAGAACTGAAAAGCATTTGTAGTGCAAAAT
TGATTTGTGATTCTCTCGGAACTATTTCAATAAGAAATTGTGAGAATCTGAAGAGGATGCCAATTTGTTTTCCGTTGCTTGAAAATGGCCAGCCATCTCC
TCCCCCTTCTCTTACATATATCTATATAGAACCGAAAGAATGGTGGGAGTCTGTAGTGGAGTGGGATCATCCTAATGCTAAGAACATCCTTCGTCCCTTT
GTAAAGTTTTTTGGTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.019G011826.1 pacid=42774084 polypeptide=Potri.019G011826.1.p locus=Potri.019G011826 ID=Potri.019G011826.1.v4.1 annot-version=v4.1
MSRPNDPPFWEYVEKMDGGGMTCTFCGHFFSEGTSITRIKLHLARKKGRGVKICENVPPEVHDAACEAAVDNSPPKKKLKTVAGSSSNEAANAISASTQE
QNNEVTHVEMAQQGGPFFTGELAWANDLIGAAEVVQLERGSSHEPRGDSSRPTDDQLCSPSVNNDANMNDVQNMVAVRIEPVPVQVLEQSNAELDSFAGH
AGRIQVGVHGMEQGAEEERICPNSAENGMENTGDGSSQHSKRLIIDAHYNTGEATQGIDLAQQLEGKTWDQINAIATSLMGEEDVENNSGRSEQPGAGAS
SSGGVAGNTNKIKGDALPTRKMVGQAFEEHKKTISSLLMRNEVSSIGIYGMGGVGKTALVTHIHNQLLERRDTHVYWITVSQNTSIHRLQTSLAGRIGLD
LSKVDEEVHRAVALNKELMKKQKWVLILDDLWKAFDLQKLGVPDQVEGCKLILTSRSAKVCQQMKTQHTIKVQPISEREAWTLFIERLGHDRELSPKVKR
IAVEVVRECAGLPLGIITMAGSMRGVDEPHEWRNTLNKLKGSKYRDMEDDVFRLLRISYDQLDNDLALQQCLLYCALYPEDYQIEREELIGYLIDEGIIE
EMRSRQAAFDEGHTMLDKLEKVCLLERACYGDHNTSVKMHDLIRDMAHQILQTNSPVMVGGYYDELPVDMWKENLVRVSLKHCYFKEIPSSHSPRCPNLS
TLLLCDNGQLKFIEDSFFQHLHGLKVLDLSRTDIIELPGSVSELVSLTALLLEECENLRHVPSLEKLRALKRLDLSGTWALEKIPQDMQCLSNLRYLRMN
GCGEMEFPSGILPILSHLQVFILEEIDDDFIPVTVTGEEVGCLRELENLVCHFEGQSDFVEYLNSRDKTRSLSTYSIFVGPLDEYCSEIADHGGSKTVWL
GNLCNNGDGDFQVMFPNDIQELFIFKCSCDVSSLIEHSIELEVIHIEDCNSMESLISSSWFCPSPTPLSSYNGVFSGLKEFNCSGCSSMKKLFPLVLLPN
LVNLENISVFGCEKMEEIIVGTRSDEESSSNSTEFKLPKLRYLALEDLPELKRICSAKLICDSLQQIEVRNCKSMESLVPSSWICLVNLERIIVTGCGKM
EEIIGGTRADEESSNNTEFKLPKLRSLESVDLPELKRICSAKLICDSLREIEVRNCNSMEILVPSSWICLVNLERIIVAGCGKMDEIICGTRSDEEGDIG
EESSNNNTEFKLPKLRSLLLFELPELKSICSAKLICDSLGTISIRNCENLKRMPICFPLLENGQPSPPPSLTYIYIEPKEWWESVVEWDHPNAKNILRPF
VKFFG
|
DESeq2's median of ratios [POPLAR]
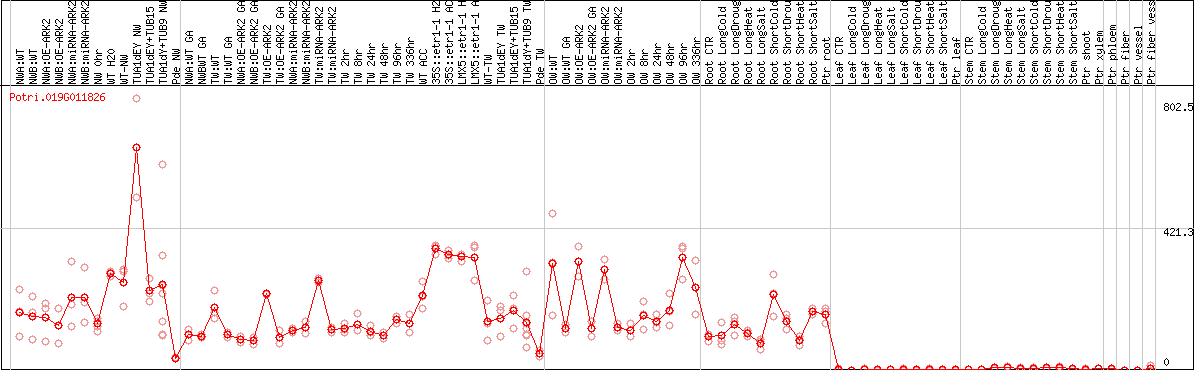
Coexpressed genes
Potri.019G011826 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
