Potri.019G021500 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.019G021500.5 pacid=42773791 polypeptide=Potri.019G021500.5.p locus=Potri.019G021500 ID=Potri.019G021500.5.v4.1 annot-version=v4.1
ATGGAGGAAAAAAATGAGGAAGTTCAGGACATTGACAGTGCTTCAAGTGATTCATTTATTGATGATGATGAGAATGATGAGCCTTCTACATCTGGACAAG
ATGATGGGACCCGTATTCAGGTGCCTTTGACCGATCAAGAAGTAGAGGAATTGGTTGCCGAGTTCTTGGAAGTTGAGAGTAAGGCTGCAGAGGCTCAAGA
AGCACTGGAAAAAGAGTCTCTTGCAAAAGTTGAGAGTGATGTGAGAGAGGAGTTGGCACAAAGTCTTCAGGGTGATGATCTGGAGGCGGCTGTTGAAGAT
GAGATGACAACTTTCAGGGAAGAGTGGGAAAATGTGCTTGATGAGCTTGAGACTGAGAGCTACCATTTGTTGGAACAACTTGATGGATCTGGTATTGAGC
TACCAAGCCTTTATAAGTGGATTGAAAGTCAGGCTCCTAATGGTTGCTGCACTGAAGCTTGGAAAAGGAGGGCACACTGGGTGGGATCTCAGGTGACCAA
GGAAACTATTGATTCAGTTGCTGATGCTGAGAAATATCTTCAAATCCATAGGCCTGCTAGACGACGTCATGGTAAATTATTGGAGGAGGGTGCAAGTGGA
TTTCTGCAGAAAAAACTTTCCATGGATGGAAGTGAGGCAATTGTAGAAAATGGAGAAGTAGATTGGGTCTCTATGAAGAAACTATTCTCAACTAGCAGTG
GGGAGGATGTTGCTTCATTTGGTAGTAAGCATTGGGCTTCTGTTTACCTGGCCAATACTCCCCAGGAAGCTGCACTTATGGGACTTAAATTTCCCGGAGT
CAATGAGGTTGAGGAGATTGAAGACATTGATGAGGATTCCATTGATCCATTTGTTGCAGAGGCCGTGGCAAATGAAAAGGAACTTGTTCTTTCAGAGGAA
CAGAGAAAAAGTTATAGAAAGGTCAAAGAGGAAGATGATGCAAAGATTGATCAGAAACTTCAACTTCATTTGAAACAGAGGAGACAGCGTAAGAGATGTA
AACAGGGTGTTAGCTCAGTCATCCAGGAAATGGGAAGGAACATGGATGAGCCACTGCCTCTGGATGACGATTACAATGAAGTCACATGCCAAGATTTGAA
GAAAGACAAGCTTTCTGTAGACCTTGTAATGGAACATTCTACAGGTAAGAGCAATTCAGTTTTCCCAGAGTCTGCGCTGCCTGATGCCACTGAACCTAGA
AGGTCCAAGCGTCCAAATGAGAGTGAGGATCTAAGCATCAATGATAAGAAGATTCGGACTGTTATCATAGACAGTGATGATGAAGCGGGCATTCTGGAGG
ATAAATCAGTACACAACATTAAAGTTGAGGATCAATCAACTTTGCAAGAGAACACTGGGGACCCTACAACTGATTGCAATCCTTCACAGGGTTCAAATGA
GAAGTTTCTGTGCACTGCCTGTGATAAAGTGGCTGTTGAAGCGCACTCACATCCACTTTTGAAAGTAATAGTTTGCAAAGACTGTAAATTCTTAATGGAG
GAAAAGATGCATGCGAAGGATCCTGACTGTTCTGAGTGTTACTGTGGATGGTGTGGACGGAACATTGAATTGGTAAGTTGTAAGTCATGCAGAACATTAT
TCTGCACTGCTTGTATAAAACGGAACATTGGTGAAGAGTACTTGCCTAAGGTTCCAGCTTCTGGTTGGCAATGTTGTTGTTGCTCTCCAAGTCTGCTACA
AATGTTTACATTACAGTTAGAGAAAGCCGTGGGGTCTGGCGATACAATGATTACGAGCTCTGATAGTGATTCAGAGTCCTCAGATACAGATGGTGGTGTT
ACAATCAGATCTAAGAGAAAGAAGAAAAAGAAGATTAGAAGGATCATTGACGATGCTGAATTAGGAGAAGAGACCAAGAGAAAAATTGCTATTGAAAAGG
AACGTCAAGAACGCTTGAAATCCCTGAAAGTACAGTTTTCTGACAAATCCAAGATGATCAATCCTGCAAGCTGCAGTGGAAACTTAACTGAAGGTGCTAG
TGTTGAAGTGCTTGGTGATGCCACAACAGGTTATATTGTGAATGTCGTGAGGGAAAAAGGTGAAGAAGCTGTGAGGATTCCTCCAAGCATTTCGTCTAAA
TTGAAAGCCCATCAGGTGACAGGGATAAGATTTTTGTGGGAAAATATTATACAGTCAATTGGAAAAGCAAGGTCTGGTGATAAAGGTCTTGGTTGCATTT
TAGCGCATATGATGGGCCTTGGTAAAACTTTTCAGGTCATAGCTTTTCTGTACACTGCCATGAGAAGTGTTGATCTGGGTTTGAGAACAGTGCTCCTTGT
GACTCCTGTGAATGTGCTGCATAACTGGCGCAAGGAGTTCATGAAGTGGACACCTTCAGAAGTAAAACCACTCCGTGTTTTCATGCTGGAGGATGTGTCG
AGGGAGAGAAGAGCGGAATTGCTTGCAAAGTGGAGAGCTAAAGGTGGTGTATTCTTGATTGGCTACTCTGCCTTTCGGAACTTAACCCTTGGAAAGAATG
TGAAGGAACCAAAATTGGCTAGAGAAATTTGTAATGCCTTGCAGGATGGACCTGATATACTTGTTTGTGATGAAGCTCATATAATCAAGAATACCAGGGC
TGATACAACCCAAGCACTGAAACTCGTGAAATGCCAGAGAAGGATAGCATTAACTGGATCACCTCTTCAGAACAATCTAATGGAGTATTATTGTATGGTT
GATTTTGTAAGAGAAGGTTTTCTTGGCAGCAGCCACGAGTTCAGGAATCGTTTCCAAAATCCTATAGAAAATGGACAACATACTAATTCAATGGTTGACG
ATGTAAAGATCATGAACCAAAGATCTCATATTCTCTATGAACAATTGAAAGGGTTTGTTCAAAGAATGGACATGAGTGTGGTGAAGAAAGACCTGCCACC
CAAAACAGTATTTGTAGTGGCTGTAAAGCTTTCTCCATTACAGAGGAAATTGTACAAGAGATTCCTTGATGTGCATGGGTTCACAAATGGCAGGGTTTCG
AATGAAAAGATGAGGAAGAGCTTTTTTGCTGGTTACCAGGCATTGGCTCAGATATGGAACCATCCTGGAATATTGCAATTGAGGAAAGGTAGAGATTACA
TTGGCCGTGAGGACAATGTTGAGAATGTTCTTGCAGATGACTGCTCTAGTGATGAAAATGTAGATTACAATACAATTGTTGGAGAGAAGTCAAGGAATCA
AAATGATTTCGTTCAAGGGAAAAGTGATGATGGATTTTTTCAAAAGGATTGGTGGAATGATCTTCTGCATGAAAATAATTATAAAGTGATTGATTATAGT
GGCAAAATGGTATTGCTGCTTGACATTTTAGTCATGTCTTCTAATGTGGGTGATAAGACATTGGTTTTTAGCCAGAGCATACCCACTTTAGATCTTATAG
AGCTTTATCTTTCAAGATTAACTCGACATGGGAAAAAGGGAAAATTCTGGAGAAAAGGAAAAGACTGGTATAGGCTAGATGGAAGAACAGAGAGCTCTGA
AAGGCAAAGGTTGGTTGAGAGGTTCAATGATCCCGAGAATAAGCGGGTCAAATGTACTTTGATTTCTACTAGGGCTGGTTCTCTCGGGATTAATCTCTAT
GCTGCTAACCGGGTGATTATTGTTGATGGTTCTTGGAATCCGACATATGATCTTCAGGCCATATATCGGGCTTGGAGGTATGGCCAGACTAAGCCAGTGT
TTGCTTACAGATTAATGGCACATGGAACCATGGAGGAAAAAATCTATAAGCGTCAGGTCACAAAGGAAGGACTTGCTGCAAGGGTGGTTGATAGACAACA
AGTATATAGGACTATGTCCAGAGAAGAAATGTTGCATCTTTTTGAATTTGGTGATGATGAGAAATCTGACACACTAAATGACATTGGCCAAGAATATAGA
CATGCAGATACCAGGAATGTTACCTGCCAAACCGTAAATTCTTTGAAAGAGAACATTCCTTGTTCTCAAGGAAGCTGCTCATCTGATAAGTTAATGGAAA
GCTTGCTTGACAAGCACCGTCAGAGGTGGATTTTTGATTACCATGAACATGAGACCCTTTTGCAAGAGAATGAAGAGGAAAAACTAACAAAGGAGGAGCA
AGACATGGCTTGGGAAGTATACAAGAGATCATTGGAATGGGAAGAAGTGCAGCGAGTTTCCGTTGATGACTCTACATTTGAGCGAAAGCCACAAATGTCA
AACGGGGCATCTTCTGCCCTAGACACTAGCAGCATACCTGTGCCAAGCATGGCCCCTCCTGCATCTGAGGCTTCCAATGTAGCACCATCCAAGAGCATTT
TAAGGAGTCGTGTGGTGCAGCGGAAATGCACTAATCTCTCTCATTTGCTAACCCTAAGAAGTCAGGGCACAAAAGCTGGGTGCACTACCGTCTGCGGGGA
ATGTGCCCAGGAGATAAGCTGGGAAGATCTGAATCGGGAGGGTAAGGCAGCACGGTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.019G021500.5 pacid=42773791 polypeptide=Potri.019G021500.5.p locus=Potri.019G021500 ID=Potri.019G021500.5.v4.1 annot-version=v4.1
MEEKNEEVQDIDSASSDSFIDDDENDEPSTSGQDDGTRIQVPLTDQEVEELVAEFLEVESKAAEAQEALEKESLAKVESDVREELAQSLQGDDLEAAVED
EMTTFREEWENVLDELETESYHLLEQLDGSGIELPSLYKWIESQAPNGCCTEAWKRRAHWVGSQVTKETIDSVADAEKYLQIHRPARRRHGKLLEEGASG
FLQKKLSMDGSEAIVENGEVDWVSMKKLFSTSSGEDVASFGSKHWASVYLANTPQEAALMGLKFPGVNEVEEIEDIDEDSIDPFVAEAVANEKELVLSEE
QRKSYRKVKEEDDAKIDQKLQLHLKQRRQRKRCKQGVSSVIQEMGRNMDEPLPLDDDYNEVTCQDLKKDKLSVDLVMEHSTGKSNSVFPESALPDATEPR
RSKRPNESEDLSINDKKIRTVIIDSDDEAGILEDKSVHNIKVEDQSTLQENTGDPTTDCNPSQGSNEKFLCTACDKVAVEAHSHPLLKVIVCKDCKFLME
EKMHAKDPDCSECYCGWCGRNIELVSCKSCRTLFCTACIKRNIGEEYLPKVPASGWQCCCCSPSLLQMFTLQLEKAVGSGDTMITSSDSDSESSDTDGGV
TIRSKRKKKKKIRRIIDDAELGEETKRKIAIEKERQERLKSLKVQFSDKSKMINPASCSGNLTEGASVEVLGDATTGYIVNVVREKGEEAVRIPPSISSK
LKAHQVTGIRFLWENIIQSIGKARSGDKGLGCILAHMMGLGKTFQVIAFLYTAMRSVDLGLRTVLLVTPVNVLHNWRKEFMKWTPSEVKPLRVFMLEDVS
RERRAELLAKWRAKGGVFLIGYSAFRNLTLGKNVKEPKLAREICNALQDGPDILVCDEAHIIKNTRADTTQALKLVKCQRRIALTGSPLQNNLMEYYCMV
DFVREGFLGSSHEFRNRFQNPIENGQHTNSMVDDVKIMNQRSHILYEQLKGFVQRMDMSVVKKDLPPKTVFVVAVKLSPLQRKLYKRFLDVHGFTNGRVS
NEKMRKSFFAGYQALAQIWNHPGILQLRKGRDYIGREDNVENVLADDCSSDENVDYNTIVGEKSRNQNDFVQGKSDDGFFQKDWWNDLLHENNYKVIDYS
GKMVLLLDILVMSSNVGDKTLVFSQSIPTLDLIELYLSRLTRHGKKGKFWRKGKDWYRLDGRTESSERQRLVERFNDPENKRVKCTLISTRAGSLGINLY
AANRVIIVDGSWNPTYDLQAIYRAWRYGQTKPVFAYRLMAHGTMEEKIYKRQVTKEGLAARVVDRQQVYRTMSREEMLHLFEFGDDEKSDTLNDIGQEYR
HADTRNVTCQTVNSLKENIPCSQGSCSSDKLMESLLDKHRQRWIFDYHEHETLLQENEEEKLTKEEQDMAWEVYKRSLEWEEVQRVSVDDSTFERKPQMS
NGASSALDTSSIPVPSMAPPASEASNVAPSKSILRSRVVQRKCTNLSHLLTLRSQGTKAGCTTVCGECAQEISWEDLNREGKAAR
|
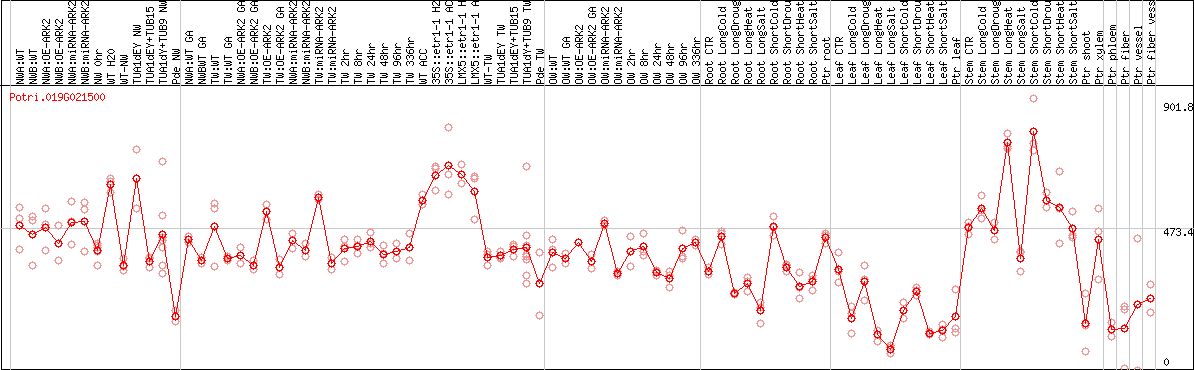
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G021500 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
