External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G04370 227 / 1e-71
NAMT1
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
AT3G11480 219 / 1e-68
BSMT1, ATBSMT1
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT5G66430 218 / 2e-68
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT5G38020 216 / 1e-67
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT5G04380 209 / 1e-64
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT1G19640 190 / 2e-57
JMT
jasmonic acid carboxyl methyltransferase (.1)
AT3G21950 186 / 3e-56
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT2G14060 182 / 8e-55
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT4G36470 140 / 1e-38
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT5G55250 100 / 5e-24
AtIAMT1, IAMT1
IAA carboxylmethyltransferase 1 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.019G022000
487 / 3e-174
AT3G11480 300 / 7e-100
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Potri.019G022400
471 / 5e-168
AT3G11480 311 / 4e-104
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Potri.019G022002
469 / 6e-167
AT3G11480 309 / 3e-103
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Potri.019G022402
438 / 6e-155
AT3G11480 261 / 1e-84
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Potri.007G021300
216 / 1e-67
AT3G11480 318 / 1e-106
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Potri.005G230100
190 / 8e-58
AT1G19640 389 / 1e-134
jasmonic acid carboxyl methyltransferase (.1)
Potri.005G045900
176 / 3e-52
AT1G19640 271 / 3e-88
jasmonic acid carboxyl methyltransferase (.1)
Potri.007G021400
142 / 3e-39
AT4G36470 469 / 4e-166
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Potri.019G016102
120 / 2e-31
AT1G68040 315 / 6e-106
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10025993
191 / 3e-57
AT1G19640 288 / 1e-93
jasmonic acid carboxyl methyltransferase (.1)
Lus10041380
166 / 3e-48
AT4G36470 278 / 4e-91
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10024671
158 / 7e-45
AT5G66430 280 / 2e-91
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10024271
147 / 3e-41
AT1G19640 315 / 2e-105
jasmonic acid carboxyl methyltransferase (.1)
Lus10041776
140 / 2e-38
AT4G36470 421 / 3e-147
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10036550
139 / 3e-38
AT3G11480 249 / 8e-80
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10036548
137 / 3e-36
AT5G66430 250 / 1e-77
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10036547
127 / 1e-33
AT5G66430 243 / 1e-77
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10043177
102 / 2e-24
AT5G55250 532 / 0.0
IAA carboxylmethyltransferase 1 (.1.2)
Lus10021802
76 / 3e-15
AT1G68040 203 / 6e-63
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF03492
Methyltransf_7
SAM dependent carboxyl methyltransferase
Representative CDS sequence
>Potri.019G022200.5 pacid=42774000 polypeptide=Potri.019G022200.5.p locus=Potri.019G022200 ID=Potri.019G022200.5.v4.1 annot-version=v4.1
ATGGTGGTGGAAAGCGTTCTTTGCATGAATCCAGGAGATGGTGAAACCAGCTACGCCAAGAATTCATTCCTTCAAAAAACAGTGCTATCAAAAGCTAGGC
CAATCCTAGAAGATACCATCAAGGACATGTTCAGCACCGCCCTTCCCACTTGCTTCAAACTAGCTGACTTGGGCTGCTTTTCAGGACCCAATACTCTCCT
GTTCGTTTCTGAAATCATGGACGTCGTTTATGAGCTTTGCCAACAACAGAACTGTAAACTGCCCGGAATTTCAGGAGTACCTGGTTCTTTCTATCACAAG
CTCTTCCCAAGCAAGAGATTGCATTTCTTTCATTCCTCTAGCAGCCTACACTGGCTTTCTAAGGTTATGGGAACATATACATGCATGGCAAAGGCAAGTC
CTCCTAATGTATTTAAAGCTTACCTGGAGCAATTCCAGAAGGATTTCTCCTTGTTTCTACGCTTACGCTCAGAGGAAATAATACAAGGAGGACGTGTAGT
TTTTACATTCATCAGCAGGAGTACTGACGATCCAAGAAGCAATGATTGCTGTCTTATATCGGAGCTGCTAGCAAAGTCACTGCTAGACTTGGCAGCAAAG
GGACTTGTTTTAGAGGCCGATATCGATACCTTCAATCTGCCATTCTATCATCCTTACGAAGGAGAAGTGAGGGAAATTATCGAAATGGAGGGGTCATTCG
ATATTAATAAGCTAGAAACTTTTGCCATTAACTGGGATGCTAATGATGATATCAACAACAACAATTTTGTGTTTGACAAGGACCAATGTGGACGAAATGT
GGCAAATATTATAAGAGCTGCTGCAGAACCGATGTTGGTTAGTCATTTTGGAGATGATATAACGGATGACTTGTTCAAGAGGTACGCAGAGTATGTAGGT
GAACATCTGTGCGTGGAGAAGACAAAACACATACACATAGTCTTCACCATGACAAAGAAGGAGTAG
AA sequence
>Potri.019G022200.5 pacid=42774000 polypeptide=Potri.019G022200.5.p locus=Potri.019G022200 ID=Potri.019G022200.5.v4.1 annot-version=v4.1
MVVESVLCMNPGDGETSYAKNSFLQKTVLSKARPILEDTIKDMFSTALPTCFKLADLGCFSGPNTLLFVSEIMDVVYELCQQQNCKLPGISGVPGSFYHK
LFPSKRLHFFHSSSSLHWLSKVMGTYTCMAKASPPNVFKAYLEQFQKDFSLFLRLRSEEIIQGGRVVFTFISRSTDDPRSNDCCLISELLAKSLLDLAAK
GLVLEADIDTFNLPFYHPYEGEVREIIEMEGSFDINKLETFAINWDANDDINNNNFVFDKDQCGRNVANIIRAAAEPMLVSHFGDDITDDLFKRYAEYVG
EHLCVEKTKHIHIVFTMTKKE
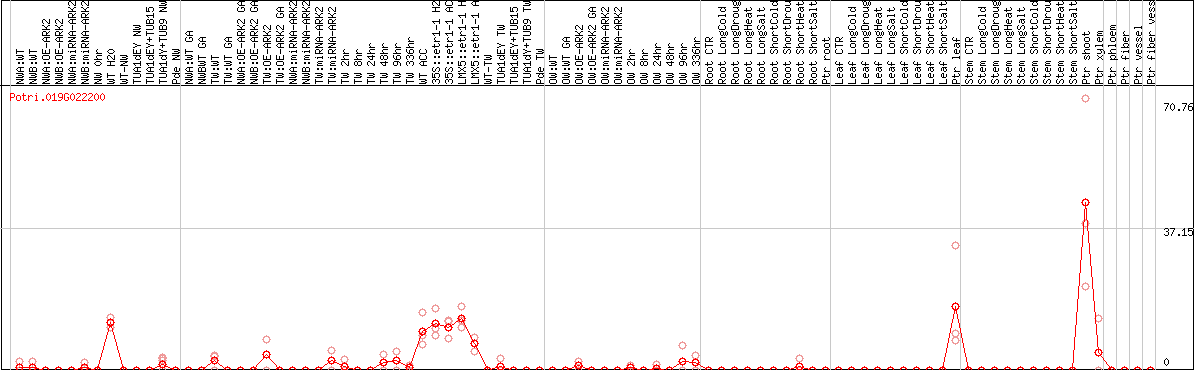
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G022200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.