External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G26594 120 / 4e-36
ARR24
response regulator 24 (.1)
AT3G04280 119 / 3e-35
ARR22
response regulator 22 (.1.2.3)
AT2G47430 57 / 3e-10
CKI1
CYTOKININ-INDEPENDENT 1, Signal transduction histidine kinase (.1)
AT5G10720 52 / 1e-08
CKI2, AHK5
CYTOKININ INDEPENDENT 2, histidine kinase 5 (.1)
AT2G01830 51 / 4e-08
CRE1, AHK4, WOL1, WOL, ATCRE1
WOODEN LEG 1, WOODEN LEG, CYTOKININ RESPONSE 1, ARABIDOPSIS HISTIDINE KINASE 4, CHASE domain containing histidine kinase protein (.1.2.3)
AT3G04580 47 / 1e-06
EIN4
ETHYLENE INSENSITIVE 4, Signal transduction histidine kinase, hybrid-type, ethylene sensor (.1.2)
AT5G35750 46 / 2e-06
AHK2
histidine kinase 2 (.1)
AT2G17820 44 / 6e-06
AHK1, ATHK1
histidine kinase 1 (.1)
AT1G27320 44 / 1e-05
AHK3
histidine kinase 3 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10004775
112 / 4e-33
AT5G26594 98 / 2e-27
response regulator 24 (.1)
Lus10005407
95 / 1e-25
AT5G26594 135 / 2e-41
response regulator 24 (.1)
Lus10028021
76 / 2e-18
AT5G26594 99 / 3e-27
response regulator 24 (.1)
Lus10031600
66 / 2e-13
AT2G47430 624 / 0.0
CYTOKININ-INDEPENDENT 1, Signal transduction histidine kinase (.1)
Lus10033746
63 / 2e-12
AT2G47430 615 / 0.0
CYTOKININ-INDEPENDENT 1, Signal transduction histidine kinase (.1)
Lus10035114
57 / 2e-10
AT2G47430 132 / 2e-34
CYTOKININ-INDEPENDENT 1, Signal transduction histidine kinase (.1)
Lus10031985
57 / 4e-10
AT5G63890 752 / 0.0
HISTIDINE BIOSYNTHESIS 8, histidinol dehydrogenase (.1.2)
Lus10029849
55 / 2e-09
AT2G47430 489 / 2e-153
CYTOKININ-INDEPENDENT 1, Signal transduction histidine kinase (.1)
Lus10028992
54 / 3e-09
AT2G01830 1337 / 0.0
WOODEN LEG 1, WOODEN LEG, CYTOKININ RESPONSE 1, ARABIDOPSIS HISTIDINE KINASE 4, CHASE domain containing histidine kinase protein (.1.2.3)
Lus10003686
53 / 7e-09
AT2G01830 1330 / 0.0
WOODEN LEG 1, WOODEN LEG, CYTOKININ RESPONSE 1, ARABIDOPSIS HISTIDINE KINASE 4, CHASE domain containing histidine kinase protein (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0304
CheY
PF00072
Response_reg
Response regulator receiver domain
Representative CDS sequence
>Potri.019G024900.1 pacid=42774214 polypeptide=Potri.019G024900.1.p locus=Potri.019G024900 ID=Potri.019G024900.1.v4.1 annot-version=v4.1
ATGGCTGGTGGAGAAGAGAACAATAATGGTGGTAAAAATAGTTTTTCTGTGCTTGTTGTCGATGATGATACTATCATTCGAATGGTCCATAGGATGCTTG
TGACAGGTCTTGGGCTGAAAGTTCAAGAAGCCAAAAATGGAAAAGAAGCTGTTGATCTTCATATCAATGGAGCTTCCTTTGACCTTATTTTAATGGATAT
GGAAATGCCCATCATGAATGGTCCCGCGGCGACAAGGGAGCTTCGAGCAATGGGTGTGAAGAGCATCATTGTTGGTGTCACATCTCATACTTCTGAGTCT
ATCCACAAAGATTTCATGGAAGCAGGCCTAAACCATTGTGTTGCGAAGCCAATGACTATAGCCAAGATTGCCCCCTTCCTTCCGAAATCCAACAACAACT
AA
AA sequence
>Potri.019G024900.1 pacid=42774214 polypeptide=Potri.019G024900.1.p locus=Potri.019G024900 ID=Potri.019G024900.1.v4.1 annot-version=v4.1
MAGGEENNNGGKNSFSVLVVDDDTIIRMVHRMLVTGLGLKVQEAKNGKEAVDLHINGASFDLILMDMEMPIMNGPAATRELRAMGVKSIIVGVTSHTSES
IHKDFMEAGLNHCVAKPMTIAKIAPFLPKSNNN
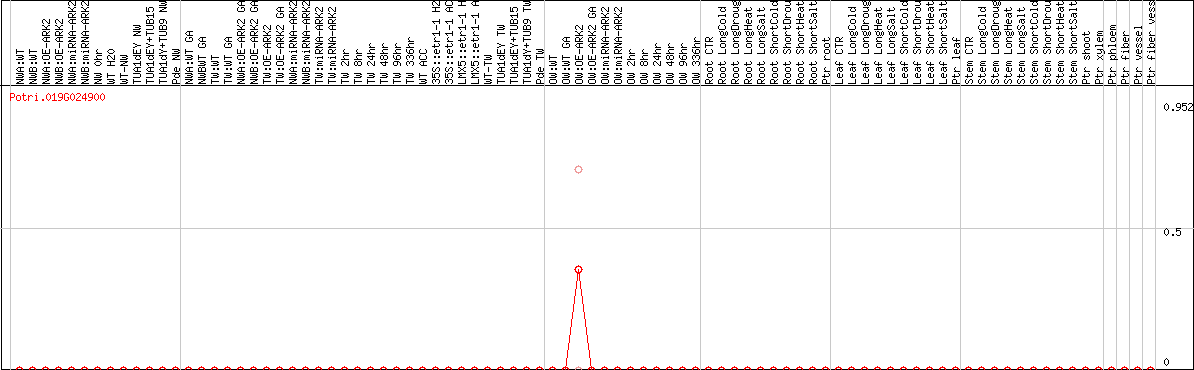
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G024900 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.