External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G55340 232 / 9e-78
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
AT3G03880 218 / 2e-72
Protein of unknown function (DUF1639) (.1)
AT4G20300 107 / 2e-27
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
AT1G68340 58 / 5e-10
Protein of unknown function (DUF1639) (.1)
AT1G25370 58 / 6e-10
Protein of unknown function (DUF1639) (.1)
AT3G60410 57 / 2e-09
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2), Protein of unknown function (DUF1639) (.3)
AT4G17440 47 / 3e-06
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G057600
410 / 7e-148
AT1G55340 212 / 1e-69
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Potri.001G004000
238 / 4e-80
AT1G55340 268 / 4e-92
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Potri.003G221000
226 / 2e-75
AT1G55340 261 / 3e-89
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Potri.018G059200
127 / 9e-36
AT1G55340 133 / 3e-38
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Potri.006G233100
123 / 4e-34
AT1G55340 130 / 9e-37
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Potri.001G073400
111 / 5e-29
AT4G20300 226 / 3e-71
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Potri.003G157500
111 / 7e-29
AT4G20300 201 / 2e-61
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Potri.010G122700
59 / 2e-10
AT1G25370 110 / 1e-28
Protein of unknown function (DUF1639) (.1)
Potri.008G122600
54 / 2e-08
AT1G25370 100 / 6e-25
Protein of unknown function (DUF1639) (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10010312
229 / 1e-76
AT1G55340 254 / 1e-86
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Lus10010650
159 / 9e-50
AT1G55340 148 / 3e-46
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Lus10013412
142 / 3e-43
AT1G55340 165 / 1e-52
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Lus10013616
119 / 9e-35
AT1G55340 111 / 1e-32
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Lus10038379
99 / 2e-25
AT4G20300 188 / 1e-58
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Lus10036240
100 / 3e-24
AT4G20300 217 / 3e-67
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Lus10035144
53 / 3e-08
AT1G25370 87 / 2e-20
Protein of unknown function (DUF1639) (.1)
Lus10004378
49 / 1e-06
AT4G17440 96 / 4e-23
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Lus10010984
45 / 2e-05
AT3G60410 89 / 3e-20
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2), Protein of unknown function (DUF1639) (.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF07797
DUF1639
Protein of unknown function (DUF1639)
Representative CDS sequence
>Potri.019G035300.6 pacid=42773019 polypeptide=Potri.019G035300.6.p locus=Potri.019G035300 ID=Potri.019G035300.6.v4.1 annot-version=v4.1
ATGCAATTATCTATTGATATGGAGAAGGAAGAAAGAAACCAGAGAGCTTCTTGTAAAAGTTTAGAGCCCGAGTTTTTCTTGCAGTGGGGAAATAAAAAGA
GGCTTAGATGTGTTAGAGTTAGAGACCCTCAAATCATTTCACGGAGATCTGATGGTGTTTTTCGCAGAAAAATCACTTCTCGGATTGATAGATTTGTGGT
CTCTTCTGCTACTACAGAGAAAGACACATCTCTTCTTCAATCTAATCGTATCACAAGGAATTCTGAAGCAGCAGTACTTAGGTCCAGTGTGGCTGAGAAC
AGGAAATCATCATCGCCGGAGAAGGAGGATCGTTATTACACTACAAGGGGGTCAGTGGGTCTGGATGAGAATGGCAAGGTCTCAATGGATGGTAACAATG
GAGATGATAAAGCCCACGTTTGGCCTAAACTATATACTACTTTGTCAAGCAAAGAAAAAGAGGAAGATTTTATGGCTATGAAGGGGTGTAAGCTACCTCA
AAGGCCAAAAAAGAGAGCCAAGATTATACAAAGAAGTTTACTTCTGGTAAGCCCTGGGGCATGGTTGACTGATATGTGCCAAGAGAGGTATGAAGTTAGA
GAGAAGAAGAGTTCCAAGAAGAGACCAAGAGGTTTGAAGGCCATGGGGAGCATGGAAAGCGATTCAGAATAG
AA sequence
>Potri.019G035300.6 pacid=42773019 polypeptide=Potri.019G035300.6.p locus=Potri.019G035300 ID=Potri.019G035300.6.v4.1 annot-version=v4.1
MQLSIDMEKEERNQRASCKSLEPEFFLQWGNKKRLRCVRVRDPQIISRRSDGVFRRKITSRIDRFVVSSATTEKDTSLLQSNRITRNSEAAVLRSSVAEN
RKSSSPEKEDRYYTTRGSVGLDENGKVSMDGNNGDDKAHVWPKLYTTLSSKEKEEDFMAMKGCKLPQRPKKRAKIIQRSLLLVSPGAWLTDMCQERYEVR
EKKSSKKRPRGLKAMGSMESDSE
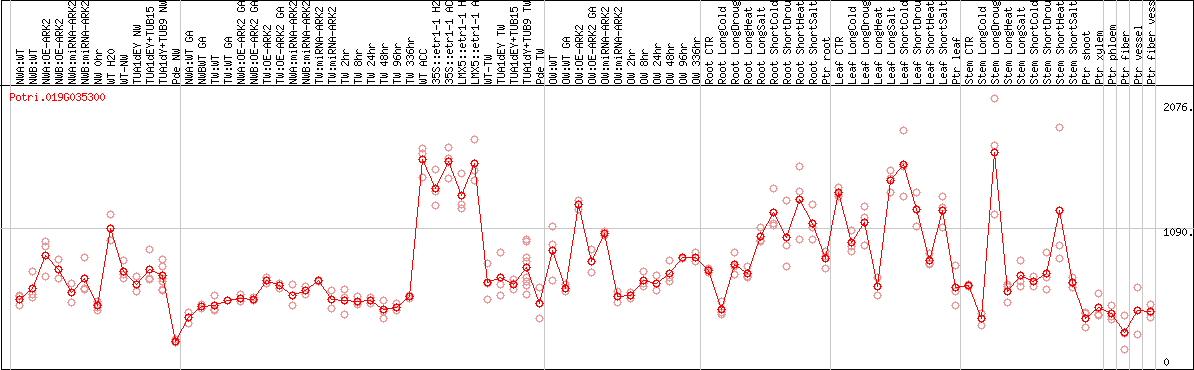
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G035300 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.