External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G32450 401 / 2e-142
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
AT5G46870 207 / 1e-65
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT4G17720 204 / 3e-64
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT5G16840 195 / 1e-61
BPA1
binding partner of acd11 1 (.1.2.3)
AT1G67950 173 / 6e-53
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2), RNA-binding (RRM/RBD/RNP motifs) family protein (.3), RNA-binding (RRM/RBD/RNP motifs) family protein (.4)
AT1G14340 146 / 1e-42
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT3G01210 125 / 1e-34
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT2G16940 44 / 7e-05
Splicing factor, CC1-like (.1.2.3)
AT5G09880 41 / 0.0005
Splicing factor, CC1-like (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G055100
502 / 0
AT5G32450 389 / 9e-138
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.001G138400
213 / 2e-68
AT4G17720 365 / 2e-127
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.003G095400
204 / 6e-65
AT4G17720 355 / 9e-124
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.010G103600
188 / 3e-58
AT1G67950 280 / 6e-94
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2), RNA-binding (RRM/RBD/RNP motifs) family protein (.3), RNA-binding (RRM/RBD/RNP motifs) family protein (.4)
Potri.017G081400
143 / 1e-41
AT1G14340 229 / 3e-75
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.017G081500
142 / 2e-41
AT1G14340 239 / 6e-80
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.001G070200
62 / 2e-11
AT5G32450 65 / 8e-13
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.007G009800
47 / 1e-05
AT5G09880 519 / 9e-180
Splicing factor, CC1-like (.1)
Potri.004G177000
46 / 2e-05
AT2G16940 538 / 0.0
Splicing factor, CC1-like (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10013651
385 / 3e-133
AT5G32450 343 / 5e-117
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10010660
372 / 4e-130
AT5G32450 352 / 3e-122
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10040095
206 / 3e-65
AT4G17720 379 / 8e-133
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10030952
202 / 1e-63
AT4G17720 382 / 7e-134
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10011063
187 / 8e-58
AT4G17720 333 / 2e-114
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10025743
178 / 1e-54
AT4G17720 295 / 4e-100
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10000676
170 / 4e-51
AT4G17720 313 / 1e-106
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10030491
154 / 2e-45
AT1G14340 300 / 1e-103
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10012843
145 / 4e-42
AT1G14340 301 / 5e-104
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10035919
134 / 1e-37
AT4G17720 226 / 3e-73
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0221
RRM
PF00076
RRM_1
RNA recognition motif. (a.k.a. RRM, RBD, or RNP domain)
Representative CDS sequence
>Potri.019G037600.2 pacid=42774234 polypeptide=Potri.019G037600.2.p locus=Potri.019G037600 ID=Potri.019G037600.2.v4.1 annot-version=v4.1
ATGCTTGTTACAGCAATGCAGACAAGAACAGTAGAAGTGAGGAATGTATCAGATCTAGCAAGTGAGAGAGAGGTTCACGAGTTCTTCTCCTTCTCTGGCG
AAATCGAACACATTCACATCCAGCGTGAAAATGGGCAATCAAAAACTGCGTTTGTCACATTCAAAGATCCTAAAGCACTTGAAATTGCGTTGTTGTTATC
GGGAGCAACTTTAGTAGACCGGATAGTGACTATAACCCCTGCTGAAAATTATGTACTGAATCGTGAGCTACAGGAAGTAAGGAACGTGGAAAATGCTGTC
TCTGTAGTTCCTTCGGAAAACTTCCCCTCAAATGTTGAGCAGGGGAAAACTAGCCCTTCAGGCAGTGGAAGAGTCTATGTCAGCAGAGCACAAGAAGTAG
TCACGAGTGTGCTAGCAAAAGGTTCGGCTATTAGCCAAGATGCAATGAATAAGGCCAAGGCATTTGATGAGAAACATCGATTGAGTGCGAGTGCATCTGA
GAAGGTAACTTCTTTTGACCGGAGAGTTGGGCTAACAGAGAAATTGACGATTGGAATTTCAGTAGTTAATGAGAAAGTGAAGTCTGTTGACCAAAGGCTT
CATGTGTCGGATAAAACAATGGCAGCTATATTTGCAGCAGAAAGGAAGCTAAATGACACAGGATCAGCTGTAAAATCGAGCAGGTATGTGAGTGCTGGAA
CAGCCTGGCTAAATGGTGCCTTTAGTAAGGTGGCGAGGGCTGGGCAGGTCGCTGGTACAAAAACAAGAGAAAAGTTCAATTTGGCTGTTTCGAATTTGAC
TGCTAAGGAATCTCCAATTGCTGTATGA
AA sequence
>Potri.019G037600.2 pacid=42774234 polypeptide=Potri.019G037600.2.p locus=Potri.019G037600 ID=Potri.019G037600.2.v4.1 annot-version=v4.1
MLVTAMQTRTVEVRNVSDLASEREVHEFFSFSGEIEHIHIQRENGQSKTAFVTFKDPKALEIALLLSGATLVDRIVTITPAENYVLNRELQEVRNVENAV
SVVPSENFPSNVEQGKTSPSGSGRVYVSRAQEVVTSVLAKGSAISQDAMNKAKAFDEKHRLSASASEKVTSFDRRVGLTEKLTIGISVVNEKVKSVDQRL
HVSDKTMAAIFAAERKLNDTGSAVKSSRYVSAGTAWLNGAFSKVARAGQVAGTKTREKFNLAVSNLTAKESPIAV
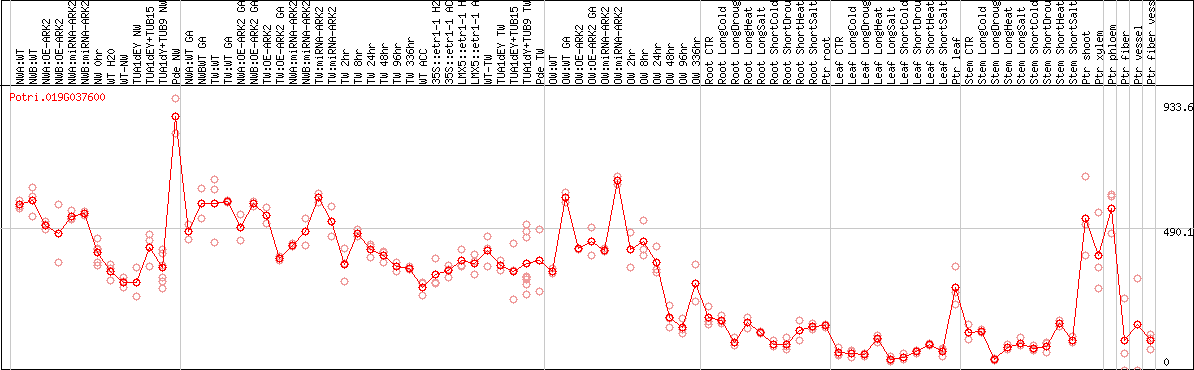
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G037600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.