Potri.019G040100 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.019G040100.13 pacid=42774274 polypeptide=Potri.019G040100.13.p locus=Potri.019G040100 ID=Potri.019G040100.13.v4.1 annot-version=v4.1
ATGCTTGCGAATCCTAGCACCAGTACTCTCCATCACAGACCTAATTACACACACAACCCTCCATGTTCACTCTCTAATCATCGCTGCAAGCTCTCTCTCT
CCCCAAATTGCCCTCGCTTCTCCAATTCTGCAAAATCCAACTTTTCTTCGCTTTCTCTGCTTCTATCAAGAAGAAAGCGTGAAAGGTTGTTTACTGTTGT
CAATGCTTTCGAGGAAACTTCTGTGGCACATGACGCCCCTCAGTCTTTGCCTCAGCCTTTTTCTGTCAAAATCCCCGTCGGTGACAGACATATATTGGTT
GAGACGGGTCATATTGGGAGACAAGCTAGTGGTTCTGTCACTGTGACGGATGGGGAAACTATTGTCTATACATCTGTTTGTTTGGATGATGTTCCTAGTG
AGCCTTCTGATTTTTATCCTCTATCTGTAAATTATCAAGAGCGATTTTCTGCAGCAGGTCGAACTAGTGGAGGATTTTTCAAACGAGAAGGGAAGTTAAA
AGATCATGAGGTTCTTATTTGTAGATTGATAGACAGGCCTTTGCGCCCAACCATGTTCAAGGGCTTCTACCATGAAACCCAGATACTATCTTGGGTTTTG
AGCTATGATGGTTTGCATTCTCCTGATTCTTTGGCTGTCACAGCAGCTGGAATAGCTGTAGCTCTTTCAGAAGTACCAAATACAAAAGTAATTGCAGGAG
TTCGAGTTGGCCTTGTTGACAATAAGTTTATTGTGAATCCAACAACCAAGGAGATGGAAGAGTCAGAATTGGATTTATTGCTGGCTGGTACAGACAGTGC
AATATTTATGATAGAGGGTTATTGTAATTTTCTTACAGAAGAAAAGTTGCTTGATGCAGTACAAATTGGCCAGGATGCAGTGCGGGCAATTTGCAATGAG
GTGGGTGTTTTGGTGAAGAAGTGTGGGAAAGCTAAAATGCTTGATGCAATCAAATTACCTCCTGCTGAGCTGTATAAGCATGTGGAGGAAATTGCTGGTG
ATGAATTGGTGAAAATATTGCAAATTACAAGCAAAGTACCAAGAAGAAAAGCCCTTGCATCACTGGAAGAAAAGGTGCTTAGTATACTTACAGAAAAGGG
ATATGTAAGCAAGGATGAAAGATTTGGAACCCCTGAAACAGTTGCAGACTTGCTAGAGGTTGAAGATGAGGATGAGGAAATTGTTGTAGATGGTGAAGTG
GATGAAGGTGATGTCCACATAAAGCCAATTGGACGCAAATTCTCTCCCTTGTTGTTTTCTGAGGTAGATGTGAAGCTGGTATTTAAAGAAGTTACATCCA
AATTCTTGCGGAAACGTATTGTTGAGGGAGGAAAAAGAAGCGACGGGAGAACTCCAGACAGGATACGTCCAATTGATTCCAGATGTGGAATACTTCCCAG
AGCACACGGAAGTGCTCTTTTCACACGTGGGGAAACACAGTCACTAGCAGTTGTTACACTTGGTGATAAACAAATGGCACAAAGAGTAGACAACCTTGTG
GACGAAGAGGAATTCAAGAGGTTCTATCTACAGTACCTGTTTCCTCCATCATGTGTTGGAGAAGTGGGCCGGATAGGAGCACCTAGTAGAAGAGAAATCG
GCCATGGAATGCTCGTGGAGAGAGCTCTCAAACCTATATTGCCTTCTGATGATGATTTTCCTTACACAGTACGGGTTGAGAGTACAATCACAGAAAGCAA
TGGCTCCTCAAGCATGGCCTCTGTTTGTGGAGGCTGCCTGGCATTACAAGATGCTGGTGTTCCTGTAAAATGCATGATTGCTGGGATTGCGATGGGTATG
GTTCTTGACACTGAGGAATTTGGGGGTGATGGAACGCCCCTCATTCTGTCTGACATAACTGGATCAGAAGATGCTTCTGGAGACATGGATTTCAAGGTTG
CTGGAAATGAAGATGGAATAACTGCATTTCAGATGGACATAAAGGTTGGAGGAATCACCTTACCTGTTATGAGAAAGGCATTGCTACAAGCAAGGGATGG
CCGAAAGCATATTCTTGCTGAAATGTTGAAATGCTCACCTTTCCCTTCCGAAAGGCTTTCTAAGTATGCCCCATTGATTCACATTATGAAGGTCAATCCA
AAGAAAGTTAACATCATCATTGGTTCTGGTGGGAAGAAGGTGAAGAGTATAATAGAAGAAACTGGAGTTGAGGCCATTGACACACGAGAAGATGGAATTG
TAAAAATTACTGCAAAGGATTTGTCAAGTATAGAAAAGTCCAAATCCATTATCAATCAGCTAACAATGGTTCCAAATGTCGGTGATATCTACAGGAACTG
TGAGATCAAATCCGTAGCTCCTTATGGAGTTTTTGTTGAGATAGCCCCAGGGCGTGAGGGACTTTGCCATATTAGTGAACTAACTTCGAACTGGTTGCCA
AAGGCTGAAGACGTTTTCAAAGTTGGAGACCGTGTTGATGTTAAACTAATTAAGGTAAATGAAAAGGGACAACTTCGACTTAGTCGCAAAGCATTGCTTC
CTGAAGCCACTCCAGAGAAGTCCAGTGCAGAGCATCATGCAACTGATAAGGCTAGTCCCAGAAGGATTGTGCAAGCACCAAAGGATGGTTTAGGTGGAGA
GTACAAGGATAAAGACAGTGCTGTAAATTCCCCAAGAAGTGATTCAATAAAGGATGCTCCAGTTTCAAAAAAGAAAGTTTACAAGGGATTAACCAGTTCT
GGCACAGAGGGGCCAAAGAACAGTGTTGCATCTGTCAGTAGCATTGCCAGCAAAGACGAAGGTACTTTAGTGAATGGAGATGCTAAGATTGGATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.019G040100.13 pacid=42774274 polypeptide=Potri.019G040100.13.p locus=Potri.019G040100 ID=Potri.019G040100.13.v4.1 annot-version=v4.1
MLANPSTSTLHHRPNYTHNPPCSLSNHRCKLSLSPNCPRFSNSAKSNFSSLSLLLSRRKRERLFTVVNAFEETSVAHDAPQSLPQPFSVKIPVGDRHILV
ETGHIGRQASGSVTVTDGETIVYTSVCLDDVPSEPSDFYPLSVNYQERFSAAGRTSGGFFKREGKLKDHEVLICRLIDRPLRPTMFKGFYHETQILSWVL
SYDGLHSPDSLAVTAAGIAVALSEVPNTKVIAGVRVGLVDNKFIVNPTTKEMEESELDLLLAGTDSAIFMIEGYCNFLTEEKLLDAVQIGQDAVRAICNE
VGVLVKKCGKAKMLDAIKLPPAELYKHVEEIAGDELVKILQITSKVPRRKALASLEEKVLSILTEKGYVSKDERFGTPETVADLLEVEDEDEEIVVDGEV
DEGDVHIKPIGRKFSPLLFSEVDVKLVFKEVTSKFLRKRIVEGGKRSDGRTPDRIRPIDSRCGILPRAHGSALFTRGETQSLAVVTLGDKQMAQRVDNLV
DEEEFKRFYLQYLFPPSCVGEVGRIGAPSRREIGHGMLVERALKPILPSDDDFPYTVRVESTITESNGSSSMASVCGGCLALQDAGVPVKCMIAGIAMGM
VLDTEEFGGDGTPLILSDITGSEDASGDMDFKVAGNEDGITAFQMDIKVGGITLPVMRKALLQARDGRKHILAEMLKCSPFPSERLSKYAPLIHIMKVNP
KKVNIIIGSGGKKVKSIIEETGVEAIDTREDGIVKITAKDLSSIEKSKSIINQLTMVPNVGDIYRNCEIKSVAPYGVFVEIAPGREGLCHISELTSNWLP
KAEDVFKVGDRVDVKLIKVNEKGQLRLSRKALLPEATPEKSSAEHHATDKASPRRIVQAPKDGLGGEYKDKDSAVNSPRSDSIKDAPVSKKKVYKGLTSS
GTEGPKNSVASVSSIASKDEGTLVNGDAKIG
|
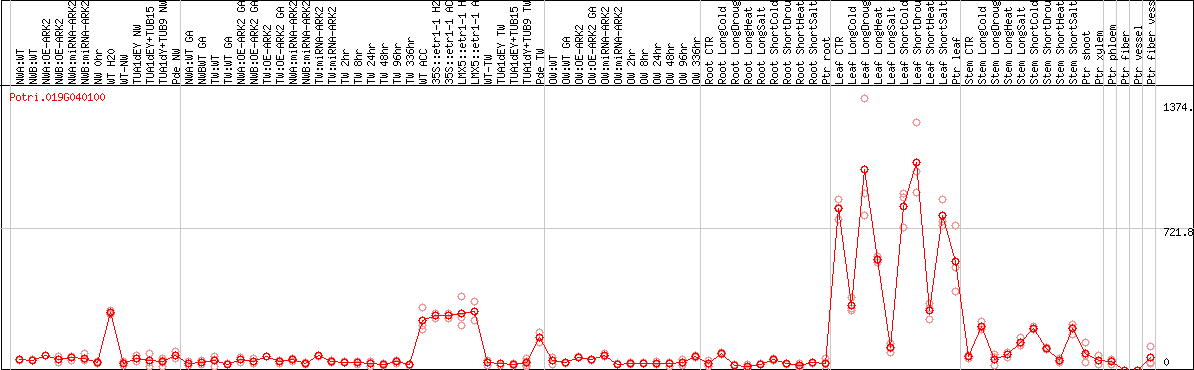
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G040100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
