Potri.019G043800 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.019G043800.1 pacid=42774236 polypeptide=Potri.019G043800.1.p locus=Potri.019G043800 ID=Potri.019G043800.1.v4.1 annot-version=v4.1
ATGAGACGTTGCAGTGGAAGCTCTAAAAGCCAAGAGAAGCAGCTGCAGCATCATCATAAGCATAAGCATTGCTGCTTATCACACCTCTTTCTTAAAGCAG
CAGCTCAAAACCCCCCAAAAGTCGCTGTCATACATGCTGCACCATCTTCTTCTTCTTCTCCTGCTGCTGCTTCTAGTGGGCCCCAAACTCAAATCTCTAG
AGAATTGATCACTTCTACTACTCCTCCAATTTATGAAGGCGACCAGTGTTTCACTTTCGCTAATGTGTTCAGCTCCGTTGATTCCCTCTCCTCTCGCCTC
CGCTCCATTCTTGATGGTGCTGATGATCCTCACCTTATCAAACCCCAATCCCCTCCAGGTAAAGGCAGCAACAATCCTGGCAAGAATCAAGCTGAGACCG
CGAGTGCATATAACCCAAAAATAGTGGGAATATATATGCCACCTTCAGTGGAATACATAATTTCTGTTTTCTCAATTTTGAGGTGTGGAGAAGCGTTCTT
GCCTATTGATCCATCGTGGCCGAGAGATAGAGTATTGTCAATTGTTGCTTCAGCCAATGCGGCCCTTATTATTACTTCTCGATCGTCGTTTGGTAAAGGT
GGTAATAAGGATATTAATGAAGCAGATTGGCTTGTGGATCGTAGTGGTTGTCGTGTCTTGTGTTTTTCGATGGAAGACAGCGAGTGTAGTGGTGGTCCGT
CGGAGTTAGCTTGGCCTTGTGAAAATGAGAAAGAGAGATTGTTCTGTTATTTGATGTACACTTCTGGATCGACTGGGAAGCCTAAAGGCGTGTGTGGGAC
TGAACAAGGCCTCTTAAATCGCTTTTGGTGGATGCAAGAATTGTATCCCCTACATGGAGAGGAAGCTCTATTGTTCAAAACATCGATAAGCTTTATCGAC
CACCTGCAAGAATTTCTCAGTGCTATGCTTACTACATGTACACTGGTTATACCTCCTTTCCATGAGCTGAAAGAGTATCCATTTTCTCTTGTCAATGTTC
TACAGGCCTACTCCATAAATAGACTTACAGCTGTCCCTTCACTTATGAGAGCAATCCTTCCTGTTTTGCAAAGGCAACATAGCATGCAGATTCAGACTTC
ATTGAAATTGCTAGTACTGAGTGGTGAAGTTTTTTCTCTATCCTTATGGGATGCGCTTTCGACCTTACTACCTAGGACCACAATTTTGAATTTATATGGG
ACTACCGAGGTATCTGGTGACTGTACATATTTTGATTGCAAGAGGTTGCCTGCGATTTTGGAGACAGAGGCACTAACAAGCATTCCAATTGGTTTGCCTA
TTTCTAACTGTGATGTGGCACTAATTTGTGAATCCGATACATCTAATGAGGGAGAGATATATGTTGGTGGTCTCTGTGTTTCTAATGGTTACTATTCTGA
GTCCACTGTTACATCCTTTATCTCTGCAAATCCACACATGGACAACATCTGTAATAGCTCTGTAGATAACTGGGGATGCCAAGCTTATTATAGAACTGGT
GATTTTGCACAAAGGCTTCAGAATGGTGACTTGTTATTCTTAGGAAGAACAGATCGGACTGTAAAAATCAATGGTCAACGCATTGTTCTAGAGGAGATTG
AAAATACACTCAGGGGACATCCTGATGTTGCTGATGCTGCTGTAATTTCTCGTGAAGGTCCAGGGGAGCTTTTGTTTCTTGATGCAATTTTGCTGTTCAA
AGAGAGAGAAAAATCTGAGGATTTTTTTGTCAGGTCTTCAATAAGAAAGTGGATGGTCGACAAAGTTCCATTAGCAATGGTTCCTAATCGCTTTGTAATC
ACAGAATCGTTGCCCATGTCTTCCACTGGAAAAGTTGATTATGCTTTATTGGCACGTTCAAAGTTTCTCAACTTGCATGTCCAAGATGAGATTGGCAATG
CAACAAGTGATCTCCTGCAAATTATCAAAAAGGCATTTTGTGATGGTTTGATGGTTGAAGAAGTTTCTTGTGATGATGATTTCTTTGCAATGGGTGGTAA
TTCCATTTCTGCAGCACATGTTTCTTATAATCTAGGGATCAATATGAGATTGCTTTATAACTTCCCTACTCCCTCTAAGCTTCATGCTGCTCTTCTAGAG
AAAAAGGAATCTTACTGCATGGAAGTGAGAGTTGATGCTAATTCACAATTGAAGCCAAAAAAAGATAGCTTGGTTTCAGATATGGCTTATTCTCCTAATC
CTACATCTCCTGTTGTTCCTGGTCTCAAATCAATGAAGCAGCCATCAAAAAACCCTCATCAAAATAATGATGATCATACTGTTGCATCTAAACGCTTCAA
GGAAGATTTGGATATTAATATTTCTTCTGCATGTGTTAAGCCTAGTGATGGACAACCATTGTCTAGTTCAATATCAATGTTATGTTCATTCAGCCGGTGC
AACACGGTTATCTATGATGAAAATTGCAGGTCACGAAAATCACATCAAATCAATCGGTTAGCAAAAGTTCCGAGAAATGGAAAAGGTTCTAGTATGCATG
AATTATGGAAGGTTTATATGGAGTCTTGTGTAGATGCATCACCACTGGTTGTGGTTAAACAGCAGGATGTCTATCTCTTCATTGGATCTCATTCGCACAA
ATTTGTTTGTGTTAATGCCCTAAGTGGTTCTATCCAGTGGGAAGTCAAGCTAGAGGGACGAATTGAAAGTTCTGCAGCAATTGTAGGCGACTTTTCTCAG
GTTGTAGTTGGCTGCTACAGTGGGAAAATATATTTTCTTGATTTTCTTGATGGCAGCATTTGCTGGACTTTCCAAACATGTGGTGAGGTGAAGTGCCAGC
CAGTTGTGGACATACACAGACAGTTAATCTGGTGTGGCTCTCATGACCATAATTTATATGCGCTTGACTACAGAAATCATTGCTGTATTTATAAGCTCTC
ATGTGATGGGAGTATATATGGGTCTCCTGCAATTGACGAGGTGCACAACACTCTTTATGTGGCATCTACTAGTGGCCATGTGACAGCTATATCAATAAAG
GCTTTGCCATTTAATACTTTATGGGAGCATGAGCTAAAGGTCCCAGTTTTTGGGTCACTTTCACTTTGTCCTTCAAGTGGAAATGTTATTTGTTGTTTAG
TGGACGGGAACATTGTTGTATTGGATTTCTGTGGCTCCATCATTTGGAGGTGTGGAACTGGTGGTCCTGTTTTTGCTGGAGCCTGTATATCTTGCGTGCT
TCCTTCTCAGGTTCTGATATGCTCTAGAAATGGACGTGTTTATTCATTTGAAATGGAAACAGGAGATTTACTTTGGGAGTATAATGTTGGAGATCCAATT
ACTGCATCTGCATATGTTGACGAGCACTTGCAGCTGCTATCTGATCCCTGCCTTTTGTCAGACAGGTTGGTTTGTGTATGCACCAGTTCTGGACGTGTAC
ATCTGCTTCAGATTAACTTGGATGATTCCGGAAAGCAAAATCAACCTGGTTTAAATATAGTTCAGGAATTTGCAAGGTTGGAGCTCCCCGGAGACATTTT
CTCTTCACCTGTGATGATTGGTGGCAGGATTTTTGTTGGATGCCGGGATGATTATGTCCACTGCATTTCTGTTGAAGATCTAAGTTCGGTGTATGAAGGT
GCTGGTTATTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.019G043800.1 pacid=42774236 polypeptide=Potri.019G043800.1.p locus=Potri.019G043800 ID=Potri.019G043800.1.v4.1 annot-version=v4.1
MRRCSGSSKSQEKQLQHHHKHKHCCLSHLFLKAAAQNPPKVAVIHAAPSSSSSPAAASSGPQTQISRELITSTTPPIYEGDQCFTFANVFSSVDSLSSRL
RSILDGADDPHLIKPQSPPGKGSNNPGKNQAETASAYNPKIVGIYMPPSVEYIISVFSILRCGEAFLPIDPSWPRDRVLSIVASANAALIITSRSSFGKG
GNKDINEADWLVDRSGCRVLCFSMEDSECSGGPSELAWPCENEKERLFCYLMYTSGSTGKPKGVCGTEQGLLNRFWWMQELYPLHGEEALLFKTSISFID
HLQEFLSAMLTTCTLVIPPFHELKEYPFSLVNVLQAYSINRLTAVPSLMRAILPVLQRQHSMQIQTSLKLLVLSGEVFSLSLWDALSTLLPRTTILNLYG
TTEVSGDCTYFDCKRLPAILETEALTSIPIGLPISNCDVALICESDTSNEGEIYVGGLCVSNGYYSESTVTSFISANPHMDNICNSSVDNWGCQAYYRTG
DFAQRLQNGDLLFLGRTDRTVKINGQRIVLEEIENTLRGHPDVADAAVISREGPGELLFLDAILLFKEREKSEDFFVRSSIRKWMVDKVPLAMVPNRFVI
TESLPMSSTGKVDYALLARSKFLNLHVQDEIGNATSDLLQIIKKAFCDGLMVEEVSCDDDFFAMGGNSISAAHVSYNLGINMRLLYNFPTPSKLHAALLE
KKESYCMEVRVDANSQLKPKKDSLVSDMAYSPNPTSPVVPGLKSMKQPSKNPHQNNDDHTVASKRFKEDLDINISSACVKPSDGQPLSSSISMLCSFSRC
NTVIYDENCRSRKSHQINRLAKVPRNGKGSSMHELWKVYMESCVDASPLVVVKQQDVYLFIGSHSHKFVCVNALSGSIQWEVKLEGRIESSAAIVGDFSQ
VVVGCYSGKIYFLDFLDGSICWTFQTCGEVKCQPVVDIHRQLIWCGSHDHNLYALDYRNHCCIYKLSCDGSIYGSPAIDEVHNTLYVASTSGHVTAISIK
ALPFNTLWEHELKVPVFGSLSLCPSSGNVICCLVDGNIVVLDFCGSIIWRCGTGGPVFAGACISCVLPSQVLICSRNGRVYSFEMETGDLLWEYNVGDPI
TASAYVDEHLQLLSDPCLLSDRLVCVCTSSGRVHLLQINLDDSGKQNQPGLNIVQEFARLELPGDIFSSPVMIGGRIFVGCRDDYVHCISVEDLSSVYEG
AGY
|
DESeq2's median of ratios [POPLAR]
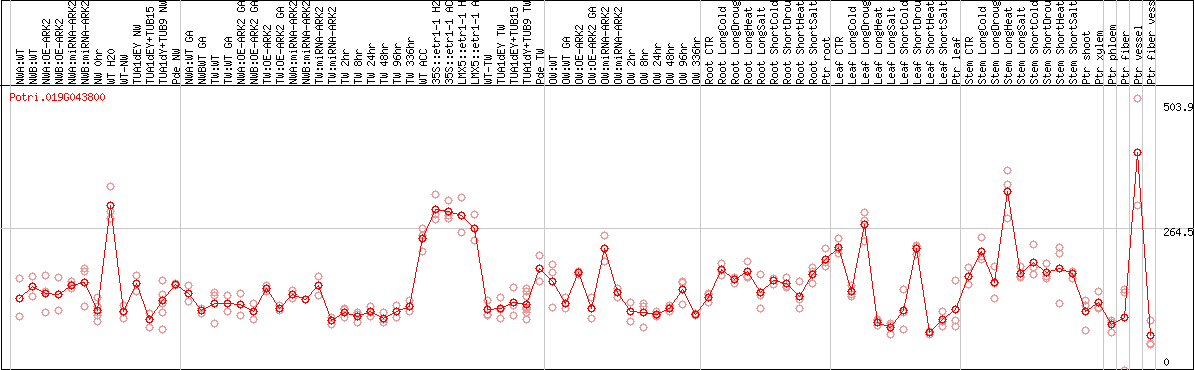
Coexpressed genes
Potri.019G043800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
