External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G16800 334 / 2e-116
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.3)
AT3G02980 300 / 2e-103
MCC1
MEIOTIC CONTROL OF CROSSOVERS1 (.1)
AT5G11340 67 / 1e-13
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1)
AT2G39000 48 / 2e-06
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G079900
459 / 6e-166
AT5G16800 346 / 3e-121
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.3)
Potri.006G248900
67 / 8e-14
AT5G11340 280 / 3e-98
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1)
Potri.018G032400
66 / 3e-13
AT5G11340 270 / 4e-94
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1)
Potri.008G039300
49 / 2e-06
AT2G39000 387 / 1e-135
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10013457
395 / 9e-141
AT5G16800 337 / 1e-117
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.3)
Lus10041016
392 / 2e-139
AT5G16800 334 / 1e-116
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.3)
Lus10013120
70 / 2e-14
AT5G11340 256 / 8e-89
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1)
Lus10008088
60 / 1e-10
AT5G11340 249 / 2e-85
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1)
Lus10040401
45 / 2e-05
AT2G39000 386 / 3e-136
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.3)
Lus10009229
45 / 2e-05
AT2G39000 320 / 5e-112
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.3)
Lus10023517
45 / 2e-05
AT2G39000 388 / 4e-137
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.3)
Lus10037996
42 / 0.0003
AT2G39000 335 / 2e-115
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0257
Acetyltrans
PF00583
Acetyltransf_1
Acetyltransferase (GNAT) family
Representative CDS sequence
>Potri.019G050700.1 pacid=42773590 polypeptide=Potri.019G050700.1.p locus=Potri.019G050700 ID=Potri.019G050700.1.v4.1 annot-version=v4.1
ATGGTGAACCAAAAAATTTCTCATCGTCCAACCATATGCTACAGACCTATACAACCTTCTGATTTGGAGGTATTAGAGCGATTACATGCTGATATATTCC
CTATCAGGTACGAGGAAGAGTTTTTCCAAAGTGTTGTTCATGCACGTGATATTGTTTCCTGGGCTGCAGTCGACCGCAGTCGACCTAATGGTCACAGTGA
TGAACTTATTGGATTTGTTACTGCACGGATTGTACTGGCAAAAGAAACTGAGATAGGAGATTTGCTCATATACGACCCCTTGAAACCAGATCAAACACTA
GTTTACATCATGACACTTGGAGTAGTAGAAACTTACAGAAATCTTGGAATAGCAAGGTCTCTTATTCGTCAGGTCATTAAATATGCTTCAAGTTTTCCGA
CATGTCATGCAGTTTACCTGCATGTGATCTCTTACAACATTCCTGCTATTCATTTGTACAAAAAAATGTCATTCAAGTGTATACGAAGGTTGCAAGGCTT
TTATCTAATCAACGGCCAGCATTATGATTCATTCTTGTTTGTTTACTATGTAAATGGTGGTTGTTCTCCTTGCTCACCATTAGAGCTTCTTGTAGCTGCA
GTGAGTTACATGACAAGCAGTCTCAAGACCGTAGCTGCGAGGCTAAGGAAAAATGAAGAGGAGTCACCAAAATTGCCCAAATGTAAAGAAACCAATTTTC
TTATATCAATGCAAGCCAAGAGAAACCTCACAACTGAATGCACTGGTTATGAATGTGTGTAA
AA sequence
>Potri.019G050700.1 pacid=42773590 polypeptide=Potri.019G050700.1.p locus=Potri.019G050700 ID=Potri.019G050700.1.v4.1 annot-version=v4.1
MVNQKISHRPTICYRPIQPSDLEVLERLHADIFPIRYEEEFFQSVVHARDIVSWAAVDRSRPNGHSDELIGFVTARIVLAKETEIGDLLIYDPLKPDQTL
VYIMTLGVVETYRNLGIARSLIRQVIKYASSFPTCHAVYLHVISYNIPAIHLYKKMSFKCIRRLQGFYLINGQHYDSFLFVYYVNGGCSPCSPLELLVAA
VSYMTSSLKTVAARLRKNEEESPKLPKCKETNFLISMQAKRNLTTECTGYECV
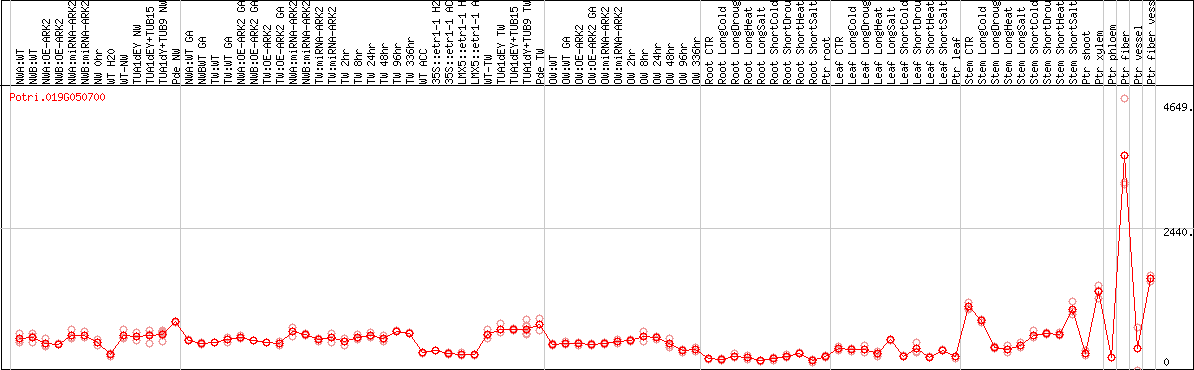
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G050700 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.