External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G40770 419 / 5e-135
zinc ion binding;DNA binding;helicases;ATP binding;nucleic acid binding (.1)
AT5G43530 98 / 2e-22
Helicase protein with RING/U-box domain (.1)
AT5G22750 92 / 2e-20
RAD5A, RAD5
DNA/RNA helicase protein (.1)
AT1G05120 92 / 2e-20
Helicase protein with RING/U-box domain (.1)
AT1G02670 90 / 8e-20
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT5G05130 85 / 5e-18
DNA/RNA helicase protein (.1)
AT1G61140 84 / 7e-18
EDA16
embryo sac development arrest 16, SNF2 domain-containing protein / helicase domain-containing protein / zinc finger protein-related (.1.2.3)
AT3G20010 82 / 6e-17
SNF2 domain-containing protein / helicase domain-containing protein / zinc finger protein-related (.1)
AT1G50410 81 / 1e-16
SNF2 domain-containing protein / helicase domain-containing protein / zinc finger protein-related (.1)
AT2G02090 80 / 2e-16
CHA19, ETL1, CHR19
CHROMATIN REMODELING 19, SNF2 domain-containing protein / helicase domain-containing protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.019G060126
631 / 0
AT2G40770 414 / 4e-133
zinc ion binding;DNA binding;helicases;ATP binding;nucleic acid binding (.1)
Potri.019G060900
617 / 0
AT2G40770 1921 / 0.0
zinc ion binding;DNA binding;helicases;ATP binding;nucleic acid binding (.1)
Potri.019G060300
561 / 0
AT2G40770 360 / 6e-114
zinc ion binding;DNA binding;helicases;ATP binding;nucleic acid binding (.1)
Potri.019G060210
411 / 1e-146
AT2G40770 292 / 4e-91
zinc ion binding;DNA binding;helicases;ATP binding;nucleic acid binding (.1)
Potri.004G189400
96 / 8e-22
AT5G22750 1506 / 0.0
DNA/RNA helicase protein (.1)
Potri.014G154600
94 / 3e-21
AT1G05120 1058 / 0.0
Helicase protein with RING/U-box domain (.1)
Potri.003G083400
94 / 5e-21
AT5G43530 1342 / 0.0
Helicase protein with RING/U-box domain (.1)
Potri.007G000700
88 / 4e-19
AT1G50410 1083 / 0.0
SNF2 domain-containing protein / helicase domain-containing protein / zinc finger protein-related (.1)
Potri.011G046400
87 / 9e-19
AT1G61140 958 / 0.0
embryo sac development arrest 16, SNF2 domain-containing protein / helicase domain-containing protein / zinc finger protein-related (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10012867
412 / 2e-131
AT2G40770 1843 / 0.0
zinc ion binding;DNA binding;helicases;ATP binding;nucleic acid binding (.1)
Lus10030512
319 / 1e-104
AT2G40770 445 / 2e-139
zinc ion binding;DNA binding;helicases;ATP binding;nucleic acid binding (.1)
Lus10043365
97 / 3e-22
AT5G22750 1445 / 0.0
DNA/RNA helicase protein (.1)
Lus10041156
92 / 2e-20
AT5G43530 1222 / 0.0
Helicase protein with RING/U-box domain (.1)
Lus10041008
90 / 1e-19
AT1G50410 1060 / 0.0
SNF2 domain-containing protein / helicase domain-containing protein / zinc finger protein-related (.1)
Lus10030044
87 / 8e-19
AT1G05120 1047 / 0.0
Helicase protein with RING/U-box domain (.1)
Lus10013452
87 / 1e-18
AT1G50410 1003 / 0.0
SNF2 domain-containing protein / helicase domain-containing protein / zinc finger protein-related (.1)
Lus10018465
86 / 3e-18
AT1G61140 546 / 2e-175
embryo sac development arrest 16, SNF2 domain-containing protein / helicase domain-containing protein / zinc finger protein-related (.1.2.3)
Lus10039001
86 / 4e-18
AT5G05130 1002 / 0.0
DNA/RNA helicase protein (.1)
Lus10019523
82 / 4e-18
AT5G22750 257 / 3e-79
DNA/RNA helicase protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF00271
Helicase_C
Helicase conserved C-terminal domain
Representative CDS sequence
>Potri.019G060000.2 pacid=42773211 polypeptide=Potri.019G060000.2.p locus=Potri.019G060000 ID=Potri.019G060000.2.v4.1 annot-version=v4.1
ATGGTTTTCCCATGCGGGCACGTCACTTGCTGTAAATGTTTCTTTGCAATGACTGAGCGCAAAATGCATGATAACAGGTTCCAACGTAAATGGGTAATGT
GCCCAACATGTCGGCAGCATACTGATTTTGGGAATATTGCTTATGCCGATGATAAACGGGACAAATCTTGTAGTTCTGCTATGCTGGATGCAATTCAAGG
CTGTGAAAAAACTGAAGCGTCTTTGGCTGTTCAAGGTTCATATGGAACAAAGGTTGAAGCAATCACTAGAAGAATCTTGTGGATAAAGTCTTCAGATCCT
AAAGCAAAGGTTCTCGTATTCTCAAGTTGGAATGATGTCCTTGATGTGTTAGAGCATGCTTTCAATGCTAATGAAGTCACCTATATCCGGATGAAAGGAG
GAAGGAAATCACATGTTGCCATCAGTGAATTCAGAGCGCAGAATGGCAGTCCCAAAAGAACTGATAGACAACAACAAGAAACAAAATCTGTTCAAGTGTT
GCTGCTCTTGATCCAGCATGGAGCCAATGGACTCAATCTTTTAGAAGCACAGCATGTTGTTCTTGTGGAGCCACTGCTCAATCCAGCAGCTGAGGCTCAA
GCAGTCAGCCGGGTGCACCGAATTGGGCAGGAAAAGAGGACATTAGTCCATCGATTTATAGTTAAAGACACTGTGGAAGAGAGCATATATGAACTGAACA
GATGTAGAAGCACCAGTTCATTCATAAGTGGGAACACGAAGAATCAGGATCAGACTTTGTTAACCCTTAAGGATGTTGAGTCCCTATTCGCAACAGTGCC
GTCCACTGTACCAGAAAGTGATGGAAAGCCAACTGAAAACCTAAGACATCTGCCCCCGTCCGTGGCTGCTGCTCTAGCAGCTGAGAGGAGACTAAAGGAA
AATACAGCAGGAATTTCCGTGTGA
AA sequence
>Potri.019G060000.2 pacid=42773211 polypeptide=Potri.019G060000.2.p locus=Potri.019G060000 ID=Potri.019G060000.2.v4.1 annot-version=v4.1
MVFPCGHVTCCKCFFAMTERKMHDNRFQRKWVMCPTCRQHTDFGNIAYADDKRDKSCSSAMLDAIQGCEKTEASLAVQGSYGTKVEAITRRILWIKSSDP
KAKVLVFSSWNDVLDVLEHAFNANEVTYIRMKGGRKSHVAISEFRAQNGSPKRTDRQQQETKSVQVLLLLIQHGANGLNLLEAQHVVLVEPLLNPAAEAQ
AVSRVHRIGQEKRTLVHRFIVKDTVEESIYELNRCRSTSSFISGNTKNQDQTLLTLKDVESLFATVPSTVPESDGKPTENLRHLPPSVAAALAAERRLKE
NTAGISV
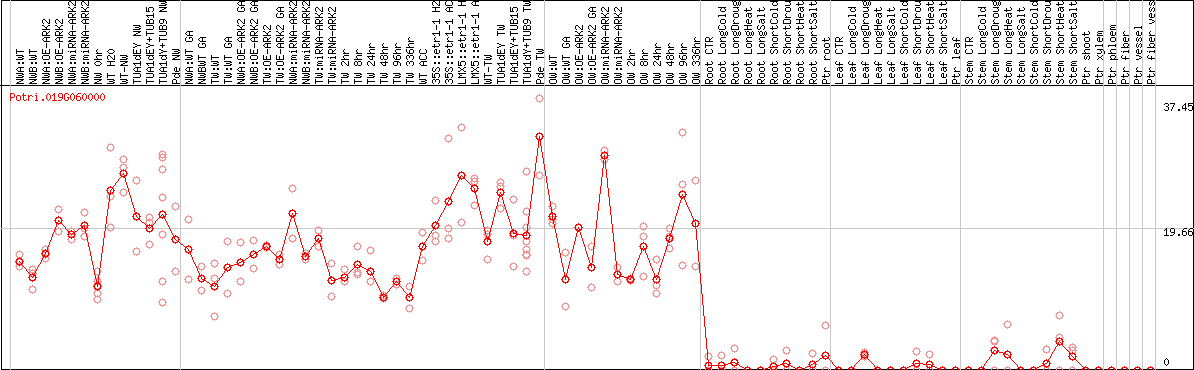
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G060000 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.