Potri.019G060400 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.019G060400.1 pacid=42773927 polypeptide=Potri.019G060400.1.p locus=Potri.019G060400 ID=Potri.019G060400.1.v4.1 annot-version=v4.1
ATGACGGCGTCCTTACTAAACAAACCATTCTTCTCCTTCTTTCTCCACATCCTGTTTTTGCTCCTTCACATATTCAACTCTTCTTCTTTCTTTGCTTTAG
CTGAACACACTCCCTCCACAACTTCACTATTTGGTAATAATAATTCAGAAGCTGAGGCTCTTCTCCAATGGAAAGCCAGTCTTGACAACCAAAGCCAATC
TCTCCTTTCTTCTTGGGTCGGCATCAGCCCTTGCATTAACTGGATTGGAATCACCTGTGACAATTCTGGAAGCGTCACCAATTTGACACTTCAAAGTTTT
GGTTTGAGAGGTACGCTTTATGATTTAAACTTCTCATCCTTCCCCAATCTTTTGTTTCTTGTCCTTCCGAACAATTCTCTTTCTGGAACCATCCCGCATG
AAATTGGAAAATTAAGAAACTTATCTTTTCTAGCTCTTTCTTGGAACCAACTCTCTGGTTCCATCCCTTCTTCTATTGGAAACTTGAAAAGTCTATCTGT
GCTCTATCTATGGGATAACCAACTCTCTGGTTCCATCCCTTTTTCTATTGGAAACATGACCATGCTCACTGGTTTAGCACTATATCAAAATAATCTCACT
GGTTCCATCCCTTCTTTTATTGGAAACTTAACAAGCCTATCCGAACTCAATCTTTGGGGTAACAAACTCTCTGGTTCCATTCCTCAGGAAATTGGGTTGC
TAGAGTCTCTCAATATTCTTGACTTGGCAGATAATGTTTTAACTGGTAGAATCCCTTATTCCATTGGAAAATTAAGAAATTTATTTTTCCTTGGTCTTTC
TTATAACCAACTCTCTGGTCTCATCCCTTCTTCTATTAAGAACTTGACAAGTGTTTCTGAGTTCTATCTTGAAAAAAACAAACTCTCTAGTCCTATTCCT
CAGGAAATTGGGTTGCTAGAGTCTCTCCATGTACTTGCCTTGGCAGGTAACAAATTTCACGGCCCATTGCCCTCAGAGATGAACAATCTCACCCATTTGC
ATGGTTTGGCTTTGGATGGCAACGAATTCACGGGTCATTTGCCAGTAGATTTATGCCATGGAGGAGTATTGAAAATCTGTACTGCTTCCAACAACTACTT
CTCAGGTTCAATCCCGGAAAGCTTGAAAAACTGTACTGGTTTATACCGAGTAAGGCTTGATCGTAATCAGTTAACAGGAAATATTTCTGAGGTTTTTGGT
ATCTATCCACATCTGAATTATATTGATTTGAGTTACAATAATTTTTATGGTGAGCTTTCTTCAAAATGGGGAGATTGTCGTAACATGACAAGCCTTCAAA
TCTCAAAAAATAATGTTTCTGGTGAGATACCACCAGAGCTTGGGAAGGCTACTCAATTGCACTTGTTAGATCTGTCATCAAATCAGTTAAAAGGAGGCAT
CCCAAAAGATTTAGGGGGTTTGAAGTTGTTGTACAAGCTTATTCTTAACAACAACCATCTTTCAGGTGCCATTCCCTTAGATATTAAGATGCTTTCCAAT
CTTCAAATCCTTAATTTAGCATCTAATAATCTGAGTGGATTGATTCCAAAACAACTTGGGGAATGCTCGAATTTACTGTTGCTGAACCTCAGCGGTAATA
AATTTAGAGAAAGCATTCCAGGTGAGATAGGCTTTCTACTTTCTCTTCAAGATCTTGATCTTAGCTGCAATTTCCTGACTCGAGATATACCGCGGGAACT
TGGGCAACTGCAGAAGTTGGAAACTCTGAATGTCTCCCACAACATGCTTTCTGGACGGATTCCGAGCACTTTCAAAGACATGCTAAGCTTAACAACTGTG
GATATCTCATCCAATAAGCTACAAGGCCCCATTCCCGATATCAAAGCCTTCCACAACGCTTCATTTGAGGCATTAAGGGATAATATGGGTATATGTGGCA
ACGCCAGTGGTCTGAAGCCGTGCAATCTTCCAACAAGCAGCAAAACTGTTAAGAGAAAGAGCAACAAGCTTGTGGTTTTGATTGTACTCCCTCTTTTAGG
AAGCTTGCTTTTAGTGTTTGTTGTGCTTGGAGCTTTATCCATTCTCTGCAAAAGAGCTAGGAAGACAAACGCTGAGCCAGAAAATGAACAAGATCGAAAC
ATGTTCACGATATTGGGCCATGATGGGAAGAAGTTCTATGAGAATATCGTTGAAGCTACAGAGGAATTCAACTCCAACTACTGCATTGGCGAAGGAGGAT
ATGGAACTGTTTATAAAGCAGTGATGCCAACAGAACAAGTGGTCGCTGTGAAGAAACTTCACCGGTCACAAACAGAAAAGTTGTCCGATTTTAAGGCTTT
TGAAAAGGAAGTTTGTGTGCTGGCAAATATTCGACATCGAAATATTGTGAAAATGTATGGATTTTGCTCACATACAAAGCATTCTTTTTTGGTTTACGAG
TTCATAGAAAGAGGAAGCTTGAGAAAGATTATAAGCAGTGAGGAACAAGCAATTGAGTTCGATTGGACGAAAAGGCTAAACGTTGTGAAAGGGGTGGGTG
GAGCTTTATCCTATCTACACCATTCTTGCTCTCCTCCGATCATTCATCGAGACATTACCAGCAATAATATCCTTCTGGACTTGGAATATGAAGCACACGT
CTCCGACTTTGGCACCGCTAGACTGTTGATGCCTGACTCATCCAATTGGACCTCATTTGCAGGTACCTTTGGATATACTGCTCCAGAGCTAGCTTACACA
ATGAAGGTGACTGAAAAATGCGATGTCTATAGCTTCGGAGTGGTAACAATGGAGGTGATGACAGGAAGGCACCCCGGCGATCTCATTTCAGCACTCTTAT
CACCAGGATCTTCGTCGTCATCATCGATGCCACCGATTGCTCAACATGCACTATTGAAGGATGTGTTAGACCAACGCATCTCGCTTCCTAAAAAAGGAGC
AGCAGAGGGCGTTGTCCACATGATGAAAATTACACTTGCATGCTTGCATCCCAATCCCCAATCTCGGCCTACAATGGAAAAAATTTCTTTCGAGCTTACA
ACTAAGTGGCCTCCACTGCCTCAGGCATTCGGCACAATAAGTTTGGGAGATCTCTTCTCATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.019G060400.1 pacid=42773927 polypeptide=Potri.019G060400.1.p locus=Potri.019G060400 ID=Potri.019G060400.1.v4.1 annot-version=v4.1
MTASLLNKPFFSFFLHILFLLLHIFNSSSFFALAEHTPSTTSLFGNNNSEAEALLQWKASLDNQSQSLLSSWVGISPCINWIGITCDNSGSVTNLTLQSF
GLRGTLYDLNFSSFPNLLFLVLPNNSLSGTIPHEIGKLRNLSFLALSWNQLSGSIPSSIGNLKSLSVLYLWDNQLSGSIPFSIGNMTMLTGLALYQNNLT
GSIPSFIGNLTSLSELNLWGNKLSGSIPQEIGLLESLNILDLADNVLTGRIPYSIGKLRNLFFLGLSYNQLSGLIPSSIKNLTSVSEFYLEKNKLSSPIP
QEIGLLESLHVLALAGNKFHGPLPSEMNNLTHLHGLALDGNEFTGHLPVDLCHGGVLKICTASNNYFSGSIPESLKNCTGLYRVRLDRNQLTGNISEVFG
IYPHLNYIDLSYNNFYGELSSKWGDCRNMTSLQISKNNVSGEIPPELGKATQLHLLDLSSNQLKGGIPKDLGGLKLLYKLILNNNHLSGAIPLDIKMLSN
LQILNLASNNLSGLIPKQLGECSNLLLLNLSGNKFRESIPGEIGFLLSLQDLDLSCNFLTRDIPRELGQLQKLETLNVSHNMLSGRIPSTFKDMLSLTTV
DISSNKLQGPIPDIKAFHNASFEALRDNMGICGNASGLKPCNLPTSSKTVKRKSNKLVVLIVLPLLGSLLLVFVVLGALSILCKRARKTNAEPENEQDRN
MFTILGHDGKKFYENIVEATEEFNSNYCIGEGGYGTVYKAVMPTEQVVAVKKLHRSQTEKLSDFKAFEKEVCVLANIRHRNIVKMYGFCSHTKHSFLVYE
FIERGSLRKIISSEEQAIEFDWTKRLNVVKGVGGALSYLHHSCSPPIIHRDITSNNILLDLEYEAHVSDFGTARLLMPDSSNWTSFAGTFGYTAPELAYT
MKVTEKCDVYSFGVVTMEVMTGRHPGDLISALLSPGSSSSSSMPPIAQHALLKDVLDQRISLPKKGAAEGVVHMMKITLACLHPNPQSRPTMEKISFELT
TKWPPLPQAFGTISLGDLFS
|
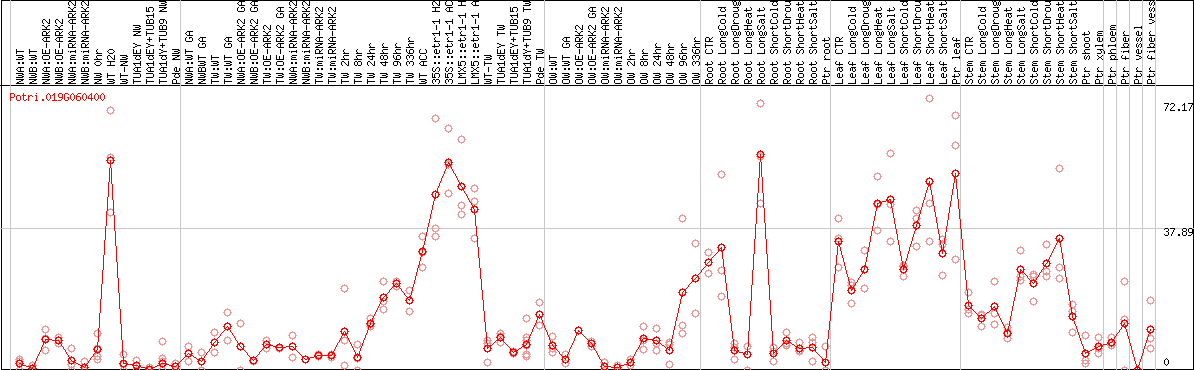
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G060400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
