External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G12010 180 / 3e-52
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT5G41750 176 / 1e-50
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
AT1G65850 173 / 7e-50
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
AT1G69550 170 / 1e-48
disease resistance protein (TIR-NBS-LRR class) (.1)
AT3G44480 168 / 6e-48
COG1, RPP10, RPP1
recognition of peronospora parasitica 1, Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT2G20142 159 / 7e-48
Toll-Interleukin-Resistance (TIR) domain family protein (.1)
AT1G31540 166 / 1e-47
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
AT3G04220 166 / 2e-47
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT3G44630 166 / 4e-47
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2), Disease resistance protein (TIR-NBS-LRR class) family (.3)
AT1G72920 156 / 4e-47
Toll-Interleukin-Resistance (TIR) domain family protein (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10029722
209 / 2e-62
AT5G17680 608 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10013729
202 / 2e-61
AT5G36930 310 / 2e-92
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10011104
200 / 5e-59
AT5G17680 574 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10042752
176 / 2e-56
AT4G16990 145 / 5e-41
RESISTANCE TO LEPTOSPHAERIA MACULANS 3, disease resistance protein (TIR-NBS class), putative
Lus10041060
189 / 2e-55
AT5G17680 641 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10014829
186 / 2e-54
AT5G17680 441 / 1e-136
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10005588
183 / 2e-53
AT5G17680 499 / 2e-156
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10026201
183 / 3e-53
AT5G17680 511 / 1e-160
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10014582
166 / 8e-51
AT1G27170 186 / 9e-56
transmembrane receptors;ATP binding (.1.2)
Lus10030839
174 / 5e-50
AT4G12010 552 / 7e-176
Disease resistance protein (TIR-NBS-LRR class) family (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0173
STIR
PF13676
TIR_2
TIR domain
Representative CDS sequence
>Potri.019G069866.1 pacid=42773119 polypeptide=Potri.019G069866.1.p locus=Potri.019G069866 ID=Potri.019G069866.1.v4.1 annot-version=v4.1
ATGGCATCTTCATCTGCTGTTGCTCATAAATGGAAGTATGATGTGTTCCTTAGTTTTAGAGGCCAGGATACGCGCAATAATTTTACGAGCCATCTCTATG
ATGCTTTGTGTCGTAAGAAAATCAAGACCTTCATTGATAATGGGCTTGAAAGAGGAGAAGAAATTACCCCTGCCCTTCTGAAAACGATTGAAGAATCAAG
AATTTCTGTAGTAATTTTCTCCAAGAACTATGCATCTTCTCCGTGGTGTGTGGATGAACTGGTGAAAATACTTGAATGCAAAGAGACTTATGGGCAGATT
GTTTTACCGGTCTTCTATCATGTGGACCCATCTGAAGTTGATGAACAGACTGGGAGTTTTGAAAATGCATTTGCCGAACTTGAAAAAAATTTCAAGGGGA
AGATGGACAAGTTGCCAAGGTGGAGGGCTGGCTTGACATATGCAGCCAGTATATCTGGATGGGATTCACAGGTTACTAGTCCGGAGTCCAAACTAGTAAG
AGAAGTTGTGCAAACTATCTGGAAAAGGTTAAATGATGCATCCCCAAGCAAATTAAGAGGTCTGGTTGGAGTAGATTCACGCATTGAGCAAATCAACAAA
TTACTATCTACTGTGCCATCTGAT
AA sequence
>Potri.019G069866.1 pacid=42773119 polypeptide=Potri.019G069866.1.p locus=Potri.019G069866 ID=Potri.019G069866.1.v4.1 annot-version=v4.1
MASSSAVAHKWKYDVFLSFRGQDTRNNFTSHLYDALCRKKIKTFIDNGLERGEEITPALLKTIEESRISVVIFSKNYASSPWCVDELVKILECKETYGQI
VLPVFYHVDPSEVDEQTGSFENAFAELEKNFKGKMDKLPRWRAGLTYAASISGWDSQVTSPESKLVREVVQTIWKRLNDASPSKLRGLVGVDSRIEQINK
LLSTVPSD
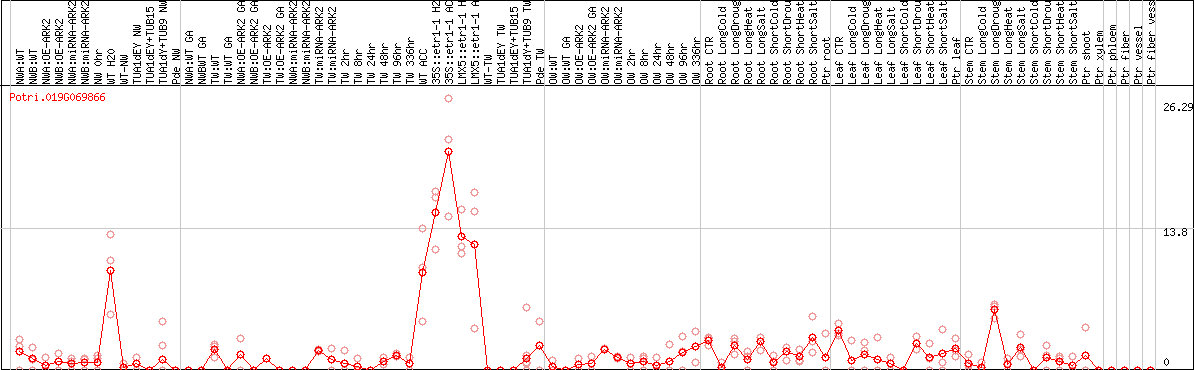
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G069866 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.