External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G33970 345 / 2e-118
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
AT1G33950 251 / 6e-82
Avirulence induced gene (AIG1) family protein (.1), Avirulence induced gene (AIG1) family protein (.2)
AT1G33900 238 / 9e-77
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT1G33960 238 / 5e-76
AIG1
AVRRPT2-INDUCED GENE 1, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT1G33930 234 / 7e-75
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT4G09940 235 / 2e-74
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT4G09950 225 / 1e-71
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT1G33890 224 / 3e-71
Avirulence induced gene (AIG1) family protein (.1)
AT2G26820 213 / 4e-65
ATPP2-A3
phloem protein 2-A3 (.1)
AT1G33910 207 / 5e-65
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G104800
540 / 0
AT1G33970 314 / 3e-106
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
Potri.008G224900
50 / 2e-06
AT4G02510 847 / 0.0
TRANSLOCON AT THE OUTER ENVELOPE MEMBRANE OF CHLOROPLASTS 86, TRANSLOCON AT THE OUTER ENVELOPE MEMBRANE OF CHLOROPLASTS 160, PLASTID PROTEIN IMPORT 2, translocon at the outer envelope membrane of chloroplasts 159 (.1)
Potri.010G014300
49 / 6e-06
AT4G02510 812 / 0.0
TRANSLOCON AT THE OUTER ENVELOPE MEMBRANE OF CHLOROPLASTS 86, TRANSLOCON AT THE OUTER ENVELOPE MEMBRANE OF CHLOROPLASTS 160, PLASTID PROTEIN IMPORT 2, translocon at the outer envelope membrane of chloroplasts 159 (.1)
Potri.009G131300
48 / 7e-06
AT3G16620 1111 / 0.0
ARABIDOPSIS THALIANA TRANSLOCON OUTER COMPLEX PROTEIN 120, translocon outer complex protein 120 (.1)
Potri.008G225000
48 / 8e-06
AT4G02510 790 / 0.0
TRANSLOCON AT THE OUTER ENVELOPE MEMBRANE OF CHLOROPLASTS 86, TRANSLOCON AT THE OUTER ENVELOPE MEMBRANE OF CHLOROPLASTS 160, PLASTID PROTEIN IMPORT 2, translocon at the outer envelope membrane of chloroplasts 159 (.1)
Potri.009G131200
44 / 0.0001
AT3G16620 1129 / 0.0
ARABIDOPSIS THALIANA TRANSLOCON OUTER COMPLEX PROTEIN 120, translocon outer complex protein 120 (.1)
Potri.004G171600
44 / 0.0002
AT3G16620 1128 / 0.0
ARABIDOPSIS THALIANA TRANSLOCON OUTER COMPLEX PROTEIN 120, translocon outer complex protein 120 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10005300
376 / 1e-129
AT1G33970 321 / 7e-109
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
Lus10005299
348 / 9e-119
AT1G33970 339 / 5e-115
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
Lus10003732
224 / 7e-71
AT1G33970 210 / 1e-65
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
Lus10028023
205 / 1e-64
AT4G09940 181 / 2e-55
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10026891
174 / 3e-53
AT2G26820 156 / 2e-45
phloem protein 2-A3 (.1)
Lus10009485
167 / 4e-51
AT1G33970 142 / 2e-42
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
Lus10009482
134 / 4e-37
AT1G33970 148 / 9e-43
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
Lus10005301
107 / 2e-28
AT1G33970 121 / 3e-34
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
Lus10009483
104 / 1e-27
AT1G33970 91 / 3e-23
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
Lus10009484
84 / 8e-18
AT4G26220 255 / 1e-83
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF01926
MMR_HSR1
50S ribosome-binding GTPase
Representative CDS sequence
>Potri.019G077300.6 pacid=42773282 polypeptide=Potri.019G077300.6.p locus=Potri.019G077300 ID=Potri.019G077300.6.v4.1 annot-version=v4.1
ATGGGTGGAAGTGCAATGGATGACGATTGGGAGTTTGCTTCACCATCAAATGGGGCTCGAACAATTGTTTTAGTTGGACGCACAGGCAATGGAAAAAGTG
CGACTGGTAACAGTATTCTTGGAAGAAAGGCCTTCAAGTCAAGGGCTAGCTCATCCGGTGTTACCAGTAGTTGTGAATTGCAGAGAACTGTACTGAGAGA
TGGTCAGATTATCAATGTCATTGATACTCCTGGACTTTTTGATTTTTCTGCCGGATCTGAATTTGTTGGCAGAGAAATTGTCAAATGTATTAATATGGCA
AAGGATGGGATCCATGCTGTCCTAGTAGTTTTCTCAGTTAGGACCCGCTTTTCACAAGAAGAAGAAGCAGCCCTTCGCAGCTTACAAACTTTGTTTGGAA
GTAAGATACTTGACTACATGATTGTAGTCTTCACTGGGGGAGATGAGCTTGAAGATAATGATGAGACTTTAGAAGACTATTTGGGTCGCGAGTGTCCACA
GCCTTTAAAGGAAGTTCTCACACTGTGTGAAAATCGACGAGTTCTTTTTAATAACAAAACTAAAGATGTACTTAAGGGAGTTGAGCAGGTGCAGGAGCTT
CTATCCCTTGTAAACAGAGTCATAGAACAAAATGGAGGCCAACCATACAGTGACGAATTATTTGCTGAAATACAGAAAGGGGAAATGAACTTCCGTGGTC
AACAAGAAGAGGTTGATTCCTTGAAAGGGAATTTTTCTATAGGAGAAATATCAGAGTTGCAGGAGCAGATGAAAAGGCAATATGAAGATCAATTGAAACG
AGTAACTGATATGGTTGAGATGAAGCTTAAAGAGGCAACTGGTAATCTTGAGCGGCGGTTAGCAGAAGAGCAAGCTGCACGATTACGGGCAGAAGAGAGC
GCACAATTAGAGCAAAGGAAATCAAACGAAGAAATTCGCATGCTTAGAGAGAGGCTAGAGAAGGCTCACGAGGAGCTCCGCAATAAGGGAGGCTGTGCCA
TTCTATGA
AA sequence
>Potri.019G077300.6 pacid=42773282 polypeptide=Potri.019G077300.6.p locus=Potri.019G077300 ID=Potri.019G077300.6.v4.1 annot-version=v4.1
MGGSAMDDDWEFASPSNGARTIVLVGRTGNGKSATGNSILGRKAFKSRASSSGVTSSCELQRTVLRDGQIINVIDTPGLFDFSAGSEFVGREIVKCINMA
KDGIHAVLVVFSVRTRFSQEEEAALRSLQTLFGSKILDYMIVVFTGGDELEDNDETLEDYLGRECPQPLKEVLTLCENRRVLFNNKTKDVLKGVEQVQEL
LSLVNRVIEQNGGQPYSDELFAEIQKGEMNFRGQQEEVDSLKGNFSIGEISELQEQMKRQYEDQLKRVTDMVEMKLKEATGNLERRLAEEQAARLRAEES
AQLEQRKSNEEIRMLRERLEKAHEELRNKGGCAIL
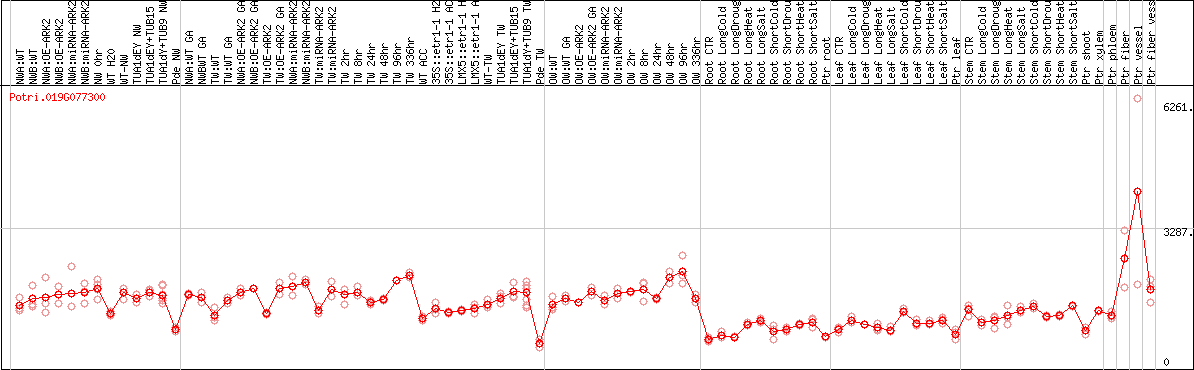
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G077300 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.