Potri.019G083100 [POPLAR]

| External link |
|
|||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||
|
Paralogs
|
|
|||||||||||||||
|
Flax homologues |
|
|||||||||||||||
| PFAM info | ||||||||||||||||
|
Representative CDS sequence |
>Potri.019G083100.1 pacid=42773056 polypeptide=Potri.019G083100.1.p locus=Potri.019G083100 ID=Potri.019G083100.1.v4.1 annot-version=v4.1
ATGGAGATGGAGAAACAATCACCGTCACCGTCGCCGCCGTCTCTGCAATCAGAATCGAGCTCGTTAAGAGAACTACAACGCGTCGGTACACTGGAAATCG
CTAGACCTAAACCACCTGTCGGTTTTCTCTGCGGTTCTATCCCTGTTCCTACTGACAAATCTTTCCACGCCTTCAATTCCGCTCTCGTTCCTTCTTCTCG
CCAAACATGTAGTGTTAGTGCACCGAGGTACCGGATGTTGCCGACGGAGACTGATTTGAATACGCTCCCTGTGGTTTCTAACTTGCCGGAGAAAGTTCTT
CCGATTAGTGCTGCTGTACAGTCTAAGTTCAAAGGAGAATTTCCTTGGGATGCTGATGCCATCTCATCAAATCTTACAAGGAAATGTGAAGCCCTTGCTG
TATCTGGTTTGGTGGAGTATGGAGATGAAATAGACGTGATTGCTTCAGCTGACATTCTTAAGCAGATCTTTAAAATACCATATTCCAAGGCTCGATTATC
AATTGCAGTACGCCGTATTGGACAGACTCTTGTTCTCAACAAAGGACCCGATGCAGAAGAGGGAGAAAGGTTGGTCAGAAGACATAAAAATCAATCAAAA
AAATGCACAGATCAATCTTTATTTTTGAATTTTGCTATGCACTCAGTTAGAATGGAGGCTTGTGATTGTCCGCCCACCCATCCTGCTTCATCAACGGGGC
AATCAAATTCATCTGTTCTTCCAGGGGGAGATGCATCCCAGTTTGTGGGGCAGAGTGATGATGTCACTAGAAATGAAGGATTCAACCATTGCTCAGAATA
TCCACATGTTAAACAGGATAACTTTTTTTGGGAGAGTAAAAAAAATAAAAGAAATAAGGGCCATCATCCTGTAAAGAAGTCTTCGCACCTTGGTGAGAAG
CCCAGAAGCTCAATGCAAGAAACTGAAAAGCATAAAAGAGTCAGTAATGATGGATTTCTAAGGGTTTTATTTTGGCAATTTCACAATTTCCGCATGCTTT
TAGGAAGTGACTTGCTTCTGTTCAGTAATGAAAAATACGTTGCTGTCAGCTTGCATTTATGGGATGTAACTCGACAGGTCACTCCGTTAACTTGGCTTGA
AGCTTGGCTTGACAATGTCATGGCTAGTGTTCCTGAGCTGGCCATATGTTATCATCAAGATGGTGTTGTTCAGGGTTATGAGCTACTTAAAACAGATGAT
ATATTTCTGTTAAAAGGCATCTCTGAAGATGGCACTCCTGCTTTCCATCCTCATGTCGTCCAACAAAATGGCCTTTCTGTTTTGAGGTTCCTTGAAGAAA
ACTGCAAGCAAGATCCTGGTGCTTACTGGCTTTATAAAAGTGCTGGAGAAGACATGATTCAACTTTTTGATCTTTGTGTTATTCCCAAAACTCATTCCTC
CAACGATTGTGATGATGGCACAAGCTCCCTGCCTTCTTTAATGCATAGAGGGAGAAGCGATTCCTTATTTTCTTTAGGAACTCTCCTTTATCGTATAGCT
CACAGGCTTTCCCTTTCAATGGCTCCTAATAATAGGGCCAAGTGTGCCAGATTCTTCCAACAATGTTTGGAATTTCTAGATGATCCGGACCATCTGGTAG
TGCGCGCATCTGCTCATGAACAATTTGCAAGGCTCCTTTTGAATCATGATGAGGAGCTGGAATTGACGTTTGAATCACTCCCAGGAGAATGTGAAGTTAC
AGTTCCAGTTGACTCCTCAGACCCCTTGAGTAGATTCTCTGAATCAGTTGCTTATGAAAATGTCTCTTCAGTTGCAGAAGACAGATGGAGTGAGGAGGGA
AAAGCTTTTCAGGAAGTAATATCAGAAGCCTCGGTAAAAATGACTTTGGAGTCAAATATTTCGACTCCAGGAAATTTGATAGCATTGGATGATACTGAAT
CAAAAGATTCAGGAGTTTTGCCTAGTTCCAGTAGTGACGAAATGGTTGCTGTGTGCAAAGTTTCCCCAACACCTCCCCATGCGGTGCAAACTGTGGCTGA
GCCAGTCTCTTCTAAGTTAGCTGCAGTACATCATGTTTCTCAAGCTATCAAGTCGCTCAGATGGATGCACCAGCTGCAAAGTTCTGATTCAGAGTTGCTT
GATGAAGGCAGTTATTTTGATGGGCCACCTTCCTCAATGAACTTTTCTGTTTGTGCATGTGGTGATGCTGATTGTATCGAAGTTTGCGACATCCGACAGT
GGCTACCAACATCAAAAGTGGATGAAAAATTGTGGAAACTAGTTCTTCTTCTTGGAGAATCTTATTTGGCACTAGGACAAGCTTATAAGGAGGATAAACA
GCTGCATCAAGCTTTGAAGGTAGTAGAATTAGCATGTGCTGTATACGGATCAATGCCACAGTTCTTGGAAGACAGTAGGTTTATTTCCTCAATGGTTACT
TATTCATCCTCAATAAAATGCAATGATGGAGATGAGAAGATGATATCGTGTGTTAGCAACAGAAAAGAAGTGAAATCCAGCTCCAATGACAGGTTTCTTG
CTTATGAGCAGTTTTCTTCTACGTACCTCTTTTGGGCCAAGGCATGGACACTGGTTGGTGATGTTTATGTTGAGTTCCATTTCATGAAGGGTAAAGTGCT
CTCAAATCAATCAGAGACAAAATCTTCTGCTAGAGAGTTGAGAATATCAACAGAGGTCGTGAAAGAAGTCCAGAGGCTGAAGAAGAAGCTGGGGCAGCAT
AATCAGAACTGCAGTTCATGCTCCTTGGTGAATTGCAGCTGCCAGAGTGATAGGGCAAGTAGCGGTAGCAGTGCAAGCAGTAGCAGTGGAGATAAACACT
CTGTAGCTTATGGTAGAAAGCACAGTAAAAGATCCCATGCTAAGGGTGCCACCTACTCACTCATGGGAGATTCTGATGATGGCCGGGCACATCACAAAGA
GAAAAGTAGGAAGAATTCTGGAGAATATCCTCAGCTTGGCAGAGGTGATAATGATACTGCAATAGAAGCTTCTGGTATTGCAGTTGATAAGCATGAGATT
AATTCCTTGGCAGATGCTAATTCTGATGTATTAGAGGGTGGTTTGGAAACACATGATGCAGGTTCCATTTTGCCTTCTCAAAGTGAAACTACTTCAAAAG
AAAAACCTAAACCAATTAAAGGAGGGATATTTAAATATATTAGCAATCCTGCTGTCAGAGATGCAGAATTCAATCTTTCTGCTGCTTTAAGTTGTTACCA
AGAAGCTAGAAAGGCATTGAGTGGACTTCCTACTGGCTCAGCAGAACTTCAATCTGTAATTAAAAAGATAGGGTGGGTTTGTAACGAAATGGGTCGGAAT
AGGCTAGAAGGAAAAGAGTTGAACAAAGCTGAACTTGCATTTGCAGATGCCATAGATGCGTTCAGAGAAGTTTCTGACCATGCCAACATCATATTAATTA
ATTGTAATTTGGGCCATGGTAGACGAGCATTGGCTGAAGAGATGGTGTCCAAGATGGAAAATCTCAAATCACATCCAATCTTCCAGAATGCATACAAGGA
AGCTCTGCAAACTGCAAAACTAGAATACAGTGAATCCCTCAGATATTATGGGGCAGCAAGAGCAGAACTAAATGCTATTGTGGAAGAGGATGATTCGGTG
CCAACTGTCTTGAGAAATGAAGTTCAGACTCAGTTTGCTCATACATATTTGAGGCTTGGCATGCTTTTGGCTAAAGAAGATGTTACCACAAGAGTTTATG
AAAATGGAGCATTGGAAGATATGCCTGTTGTTACTATTAGTCCGAATGAAAAAAGAGACAGGAAAGAAGTGAGGAAGCATGAGATTTCAGCTAATGATGC
CATTAGGGAGGCATTGACTGTGTATGAATCTCTGGGTCAATTACGGAAACAAGAAGCTGCATATGCGTACTCTCAACTTGCTAGCTACCAGAGGGATTGC
TGTTTGAAATTCTTGAATTTGGATCTAAAGAATACCACTTTAAATAAAAATGGAAACAACAATCTTCAGCGGGTGAAGCAATATGCATGCCTGGCTGAGA
GGAACTGGCAAAAAGCAATGGACTTCTACAGTCCTAAGACACATCCTGCCATGCATTTGACTATTCTTATTGAGAGATCAGCTCTTTCATTGAGCCTCTC
GAGCACCTTACACTCAAATGTGATGCTAGAATCAGCTCTAGCTCGCATGCTTGAAGGACGTCACATATCAGATGCAATATCAGATTCTTTCGGGACTGAT
TATCCTGAGATAAATTCCAAATTCTGGGGTCAACTACAGATGCTCTTAAAGAAAATGCTGTCCTTGGCCCTCTCTGCAAATGCAAATAAGCCCGTCGCTT
TTGCACAGCCAATTCCATCATCGAGCAAGTGCGGAGATGCTGGAAAACTCAGGGAGCTCTACAAAATGTCCTTGAAGTCTTCTAATTTAAGTCAATTACA
TGCTATGCACACCCTATGGACATCGTAG
|
|||||||||||||||
|
AA sequence
|
>Potri.019G083100.1 pacid=42773056 polypeptide=Potri.019G083100.1.p locus=Potri.019G083100 ID=Potri.019G083100.1.v4.1 annot-version=v4.1
MEMEKQSPSPSPPSLQSESSSLRELQRVGTLEIARPKPPVGFLCGSIPVPTDKSFHAFNSALVPSSRQTCSVSAPRYRMLPTETDLNTLPVVSNLPEKVL
PISAAVQSKFKGEFPWDADAISSNLTRKCEALAVSGLVEYGDEIDVIASADILKQIFKIPYSKARLSIAVRRIGQTLVLNKGPDAEEGERLVRRHKNQSK
KCTDQSLFLNFAMHSVRMEACDCPPTHPASSTGQSNSSVLPGGDASQFVGQSDDVTRNEGFNHCSEYPHVKQDNFFWESKKNKRNKGHHPVKKSSHLGEK
PRSSMQETEKHKRVSNDGFLRVLFWQFHNFRMLLGSDLLLFSNEKYVAVSLHLWDVTRQVTPLTWLEAWLDNVMASVPELAICYHQDGVVQGYELLKTDD
IFLLKGISEDGTPAFHPHVVQQNGLSVLRFLEENCKQDPGAYWLYKSAGEDMIQLFDLCVIPKTHSSNDCDDGTSSLPSLMHRGRSDSLFSLGTLLYRIA
HRLSLSMAPNNRAKCARFFQQCLEFLDDPDHLVVRASAHEQFARLLLNHDEELELTFESLPGECEVTVPVDSSDPLSRFSESVAYENVSSVAEDRWSEEG
KAFQEVISEASVKMTLESNISTPGNLIALDDTESKDSGVLPSSSSDEMVAVCKVSPTPPHAVQTVAEPVSSKLAAVHHVSQAIKSLRWMHQLQSSDSELL
DEGSYFDGPPSSMNFSVCACGDADCIEVCDIRQWLPTSKVDEKLWKLVLLLGESYLALGQAYKEDKQLHQALKVVELACAVYGSMPQFLEDSRFISSMVT
YSSSIKCNDGDEKMISCVSNRKEVKSSSNDRFLAYEQFSSTYLFWAKAWTLVGDVYVEFHFMKGKVLSNQSETKSSARELRISTEVVKEVQRLKKKLGQH
NQNCSSCSLVNCSCQSDRASSGSSASSSSGDKHSVAYGRKHSKRSHAKGATYSLMGDSDDGRAHHKEKSRKNSGEYPQLGRGDNDTAIEASGIAVDKHEI
NSLADANSDVLEGGLETHDAGSILPSQSETTSKEKPKPIKGGIFKYISNPAVRDAEFNLSAALSCYQEARKALSGLPTGSAELQSVIKKIGWVCNEMGRN
RLEGKELNKAELAFADAIDAFREVSDHANIILINCNLGHGRRALAEEMVSKMENLKSHPIFQNAYKEALQTAKLEYSESLRYYGAARAELNAIVEEDDSV
PTVLRNEVQTQFAHTYLRLGMLLAKEDVTTRVYENGALEDMPVVTISPNEKRDRKEVRKHEISANDAIREALTVYESLGQLRKQEAAYAYSQLASYQRDC
CLKFLNLDLKNTTLNKNGNNNLQRVKQYACLAERNWQKAMDFYSPKTHPAMHLTILIERSALSLSLSSTLHSNVMLESALARMLEGRHISDAISDSFGTD
YPEINSKFWGQLQMLLKKMLSLALSANANKPVAFAQPIPSSSKCGDAGKLRELYKMSLKSSNLSQLHAMHTLWTS
|
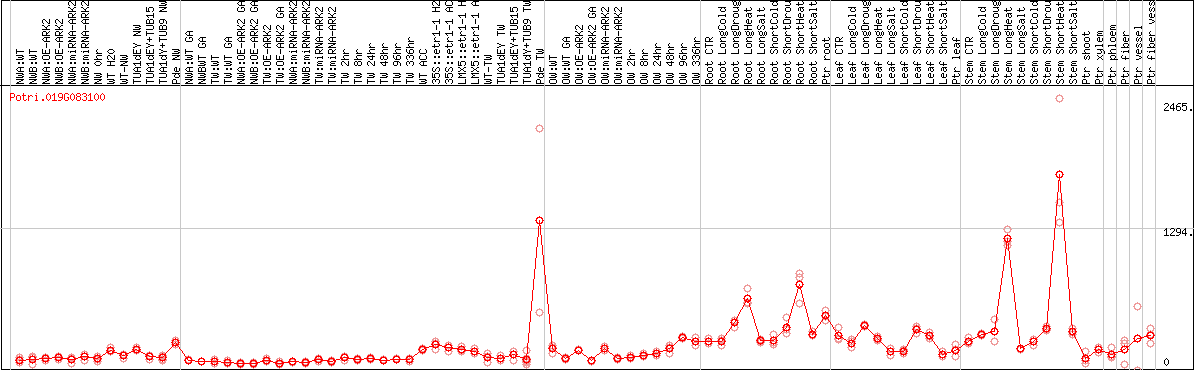
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G083100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
