Potri.019G087000 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.019G087000.4 pacid=42774217 polypeptide=Potri.019G087000.4.p locus=Potri.019G087000 ID=Potri.019G087000.4.v4.1 annot-version=v4.1
ATGTGTAGCATGTGTCATTATTGTGGTGCTGATCGGACTAAGTTGAAGGATGAAAAGAAGAAACTAGAGAATGGAGGTTGTTTAAAACTAAATGGTGAAG
AGCCCATTTGGTCTTGCCGACTTTGTCAGGAAAAGCAGGAACCAGATTTGATGAACCGTGATGGTTCGAGTCATTCTATATTACCACTGATCAGTCCAGC
TACTACGTTGCCTAGCAGTGATAGATTTATGTCTAGCTGCAGCGATCTTTATGTCGATGTAAACTCGCATGACTGGGCTCATCAAGAGGAAGAAGCAGCA
CGCAGTGCTCAGAAGGATCTCAGTTATGGGATGAATGACCAACTGCACAACTCAAGATTGGAAGCTCCTTTAAATAGAGTGGATGGATTACTTAAAGCGA
CAGAAAACAACTTGAAGGATAGTCATAACGGAACTGATAGAGAGACAGTGAGAGATGTAGAAATTGTGGAACTTCTTCACGGTCAAGAAGCAAAAGATAA
TGCTTTTGAGAAATGTGTTGGATCTTCAAACGAAGGAAGTGATGTTTCTCAAATCAGTGATGATGAAGTAGATGCGCAGGTTTGGGAGCCTCCTGAAGCA
GAAGATCCAGAGGATGATTTGGATGGCAGTGTGGCTTTTATCGATGACGACGACGATGAGTGTGGTGACGGAACAGAATGGGGTAAACCTAGTTCTTTGA
GTTACTCGAGGGATGAAGGCAGTCGGAGCTTCAAGTTCAAAGAAGAAAAACAAAAGGCAATGGACGAAGTAGTTAACGTGAAATTCAAGGCTGTCGTGAG
TCAACTTCTTAAAACTGCAGGTGTTGCCTCTCTGATGAGAGATGGTGAAAGTTGGGTGGATATTGTTACTTATTTATCATGGGAAGCTGCTTCGTTTTTG
AAGCCTGAAGCAATAGATCGGAAAGCAATGGATCCAGATGGCTATGTGAAAGTGAAATGCATTGCAACTGGTTCTCGTAGTGAAAGTGAAGTAGTGAAAG
GCTTGGTCTTTAAAAAGCGTGCTGCTCACAAGCACATGCCCACTAAGTACAAGAACCCCAGGTTGTTGCTGATCCAAGGTGTGCTTGGTCAGTCTTCAAG
TGGATTGTCATCGTTTAAATCAATGGAGCAGGAAAAGGATAATCTGAGGGCTCTCATTGAGACCATAGAAATGTGCCATCCAAATGTAGTTTTAGTAGAA
AAATCTGTTTCTCGTGATGTCCAAGAGTGCATCCTAGCAAAAGGAATGACACTGGTCTATGACATGAAGCTCCATCGACTAGAAAGAATTGCTCGCTGTA
CTGGTTCGCCAATTTTATTATCTGATGCTTTAATGAATCAGAAGTTAAAGCAATGTGATTCCTTTCATATTGAGAGATTTGTTGAGGAACATGTTGCGGT
ATGCGAAGGAGGAAAGAAGCCAAGGAAGACATTGATGTTCATCGAGGGCTGTCCAACATGTCTGGGATGTACAATTTTGTTGAAAGGAAGTCATAGTGAT
GAGTTGAAGAGGGTTAAATATGTGGTCCAATTTGCTGTAATCATGGCATATCACTTGATTCTTGAGACATCGTTTCTTGTTGATTGGAAGGCAATGTTCT
CTTCAGAAATATTTGGTGGAGTAGTTAATACCTCATCAATAGATCAACACTCTTCTGCTTTAGAAACTAGGATTCCTTGTGTTGAGGAGTCTACAACTGA
AAGTGGTTCATCTATCATTGATATTCCCATTTCTAATGGATTTCATGAAGAAGGTTCCCACAATATAAACATAGGATTAGAGGGATATGATCCTGCAGTT
TTTTCTGGATTTTCCTCTCTTTCAGCCTCTCTGAAGAAAGTTATGGGGGACAGCTTTCCTCTTGTCTCCTCTCCTCCTTATCGGTCCTTGTCTAATTACT
TTGGCTTCAATGGACAAGAAACTAATGGTCAAATCATGGAAGAGGTTCCAGTTTTAAAAACTTTGGAAGCTTTTGACCCTAGTGATATGGAAGGTAAAAA
GGATTCTGATGAAGAGAAGTCTGCCAATGATGGACAACCTCAATCTTTGTCACCTTACTCTGTAGCCACTCTGGACAGCGGAAATGATGTTGGAAATAAA
GAAGATCAAATCCAAAGTAAGGGTGATGCCAATGCAGTGTTGGATTCTCAGAGTATTTTGGTTTTGATGTCTAGACGGAATGCCTTGAGAGGGATTATAT
GTGAGCAAAGCCATTTTTCTCATATAATGTTCTACAGGAATTTTGATGTTCCCCTTGGAAAGTTTCTTCGGGATAATTTACTCAATCAGAGAAGCCAGTG
CAACACTTGCGGTGAGTTGCCTGAGGCTCACTTTTACTATTATGCTCATCATAATAAGCAGTTGACAATACGAGTTAAAAGGCTTTTCAAGACTTTGCCT
GGAGAAGGTGAAGGAAAACTTTGGATGTGGATTCGCTGTGGTAAATGTAAACATGAAAGCAGATTACCAAAATCTACTAAACGAGTCTTGATTTCAACTG
CTGCGCGTAGTCTGTCTTTCGGAAAATTCTTGGAGATAAGCTTTTCTCACCAATTTTCATCTGGTAGCTTGTTCAGCTGTGGCCATTCGCTTGAGAGAGA
CTTCCTCTACTTCTTTGGATTGGGTCCAATGGCAGCAATGTTTAAATACTCTCCAGTGACAACCTATAATGTGTCTCTCCCACCCCAGAAGCTGGAGTTC
TATCATTCAATCAGACTTGATGGCCTCAAGAAAGAATTCCATGCTGTTTATTCAAAAGGGATGCTGATCTTCAATGGGGTTGGTGAGGCGTTGAAGAATT
TAAGATCTCGATTTGCAGGGTCAGTTCTGAACCTTCAAGGTTCATTGAAAGAATTCTCTGATATTGAAGATATGTTGAAGCAGGAGAGTTCTGAGTTTGA
GTTAAACATTCAGAATGCAGTTGCTAAGAATGGTGATGAGGCTGTTTACAAGCTTCTTAGTTTGAATCAGTTATCATGGGAGCTTCTTCTTGAATCATGC
ATTTGGGAAAGACGCCTGCATTCATTGCTCTTGCCTGATACTTTGATGTTAGTAACCGGTGCCAGTAAGAAAGAATTGCAGGAACAGTTTGAGTCGCAGA
TGACTGACACTGCTGATGGAAAAATCCAGTGGAATGATAATACTTTGGGAAGCAGTGATGAAGTTTCTGACAACAGTGGCAATTTAAGGGACATGCTCAG
TACAACTGTCGAGGCAAGTGAATTTTCAATCAAAGAAATTCCAGTTGATGATCATGTTCATGAATTTAGGAAGCAGGATAATCTTTACACCTCATCTGCT
GTGGCTGAGGATATTGAGAGGTCAAGAGTGAGTGGTCTGAGCCAGAACAGATTTATCAACCAAGAACTATCTGTGAAACCTAGTGGCTCTGCCCACCAGC
ATTCTGATGATGGAAATTGCCAAGCAGACTATTTATCTGATGTACAAGTTGAAAGAACCATTCCAATTACTACAAGTATAGGGAGCAGTGATTCTTTTGT
TGATTCTGATTCAATTAAAAAGGGTACGTCTGCACGTTCTTTGGCATCCAGCTTAGAGAACTCAAATGGATGGTTCTGGATGCCATTTCCAGAAATCCGA
CAGATCTACATGAAGGATCTTCAGAGAGGTTTCATGCCAAAATTTCAGTCTATCAGTAGCAATATTCAGGAACACATGTCTGCAGCCCATCAGCTAATCA
CCAAGGAATGCCAGAGGCTGCACATCCCTCTTGGAACTGATAATTATATGGTGAAAGATTATGATGATGAACTCTCTAGCATAATTGCCTGTGCACTGGC
ATTTTTGAAAGGTCAACCTATTTCAACAGAGCTTTACAATGAGGATGATAGGAAAGAGGGTGGCATGTCTTTTAAATCAACTGATAGTTTAGACATACTG
ACTCGAATTCCTACAATGATCTCCCCACATTGGTCCTCTAATGGTTCTGATTCAGATTCTGTCCATTCTATGCTAAACATTTCTTCGGACGAGTCGCGCT
TGTCCAGCTTTGATGGGCTGAATTTGTTGGAATCACTTGTTCGTCCAGAAAATCTTAGCCCTGAAGTCGCTTTTGGGCGGTCAAAATCACTTGGAAAGGG
TAAATACTCAGTAATTTGTCTATACGCTAAGCAGTTCCATGATCTTCGGAATCGATGCTGTCCATCTGAACTTGATTATATCGCTTCCTTAAGTCGTTGT
AAGAATTGGGATGCAAAAGGTGGAAAGAGCAAATCCTTTTTTGCAAAGACACTTGATGACAGATTTATTATAAAGGAAATCAAGAAGACAGAATTTGAGT
CATTTGTGAAGTTTGCTCCACATTATTTCAAATACATGAACGAGTCATTTGAGTTAGGAAACCAAACTTGCCTTGCAAAAGTTCTGGGGATTTATCAGGT
GATCCTTAGACAGACAAAAAGCGGGAAAGAGATCAAGCATGATCTAATGGTCATGGAGAATCTTACCTTTGGTCGAAACATTACTCGCCAGTATGATCTT
AAGGGAGCTTTACATGCTAGATACAATTCAGCAGCTGATGGTTCAGGAGATGTTCTTTTGGATCAAAACTTTGTTGATGACATGAATTCCTCTCCACTGT
ATGTCAGTAATACAGCAAAGCGTCTCCTGGAACGGGCTGTTTGGAATGACACAACTTTTCTTAACTCAATCAATGTAATGGATTATTCTTTACTGGTCGG
AGTAGACACCCAGCGACGAGTGCTTGTGTGTGGGATTATTGATTATCTCAGGCAGTATACCTGGGACAAGCAGCTGGAAACATGGGTGAAGTCTTCCCTT
GTTCCTAAAAATCTGTTGCCAACTGTTATATCTCCTATAGAGTACAAAAAGAGGTTCAGAAAGTTCATGACTGCGCACTTCTTGAGTGTCCCGGATAACT
GGTGCTCACAGAGTTCCTCTAACCCTTGTGAACTTTGTGGCACCAGAGATGATGATCCATCCCAATCCACGTCCCGAAAACAAGGAGGGCAGAATGGGCT
TACTCATTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.019G087000.4 pacid=42774217 polypeptide=Potri.019G087000.4.p locus=Potri.019G087000 ID=Potri.019G087000.4.v4.1 annot-version=v4.1
MCSMCHYCGADRTKLKDEKKKLENGGCLKLNGEEPIWSCRLCQEKQEPDLMNRDGSSHSILPLISPATTLPSSDRFMSSCSDLYVDVNSHDWAHQEEEAA
RSAQKDLSYGMNDQLHNSRLEAPLNRVDGLLKATENNLKDSHNGTDRETVRDVEIVELLHGQEAKDNAFEKCVGSSNEGSDVSQISDDEVDAQVWEPPEA
EDPEDDLDGSVAFIDDDDDECGDGTEWGKPSSLSYSRDEGSRSFKFKEEKQKAMDEVVNVKFKAVVSQLLKTAGVASLMRDGESWVDIVTYLSWEAASFL
KPEAIDRKAMDPDGYVKVKCIATGSRSESEVVKGLVFKKRAAHKHMPTKYKNPRLLLIQGVLGQSSSGLSSFKSMEQEKDNLRALIETIEMCHPNVVLVE
KSVSRDVQECILAKGMTLVYDMKLHRLERIARCTGSPILLSDALMNQKLKQCDSFHIERFVEEHVAVCEGGKKPRKTLMFIEGCPTCLGCTILLKGSHSD
ELKRVKYVVQFAVIMAYHLILETSFLVDWKAMFSSEIFGGVVNTSSIDQHSSALETRIPCVEESTTESGSSIIDIPISNGFHEEGSHNINIGLEGYDPAV
FSGFSSLSASLKKVMGDSFPLVSSPPYRSLSNYFGFNGQETNGQIMEEVPVLKTLEAFDPSDMEGKKDSDEEKSANDGQPQSLSPYSVATLDSGNDVGNK
EDQIQSKGDANAVLDSQSILVLMSRRNALRGIICEQSHFSHIMFYRNFDVPLGKFLRDNLLNQRSQCNTCGELPEAHFYYYAHHNKQLTIRVKRLFKTLP
GEGEGKLWMWIRCGKCKHESRLPKSTKRVLISTAARSLSFGKFLEISFSHQFSSGSLFSCGHSLERDFLYFFGLGPMAAMFKYSPVTTYNVSLPPQKLEF
YHSIRLDGLKKEFHAVYSKGMLIFNGVGEALKNLRSRFAGSVLNLQGSLKEFSDIEDMLKQESSEFELNIQNAVAKNGDEAVYKLLSLNQLSWELLLESC
IWERRLHSLLLPDTLMLVTGASKKELQEQFESQMTDTADGKIQWNDNTLGSSDEVSDNSGNLRDMLSTTVEASEFSIKEIPVDDHVHEFRKQDNLYTSSA
VAEDIERSRVSGLSQNRFINQELSVKPSGSAHQHSDDGNCQADYLSDVQVERTIPITTSIGSSDSFVDSDSIKKGTSARSLASSLENSNGWFWMPFPEIR
QIYMKDLQRGFMPKFQSISSNIQEHMSAAHQLITKECQRLHIPLGTDNYMVKDYDDELSSIIACALAFLKGQPISTELYNEDDRKEGGMSFKSTDSLDIL
TRIPTMISPHWSSNGSDSDSVHSMLNISSDESRLSSFDGLNLLESLVRPENLSPEVAFGRSKSLGKGKYSVICLYAKQFHDLRNRCCPSELDYIASLSRC
KNWDAKGGKSKSFFAKTLDDRFIIKEIKKTEFESFVKFAPHYFKYMNESFELGNQTCLAKVLGIYQVILRQTKSGKEIKHDLMVMENLTFGRNITRQYDL
KGALHARYNSAADGSGDVLLDQNFVDDMNSSPLYVSNTAKRLLERAVWNDTTFLNSINVMDYSLLVGVDTQRRVLVCGIIDYLRQYTWDKQLETWVKSSL
VPKNLLPTVISPIEYKKRFRKFMTAHFLSVPDNWCSQSSSNPCELCGTRDDDPSQSTSRKQGGQNGLTH
|
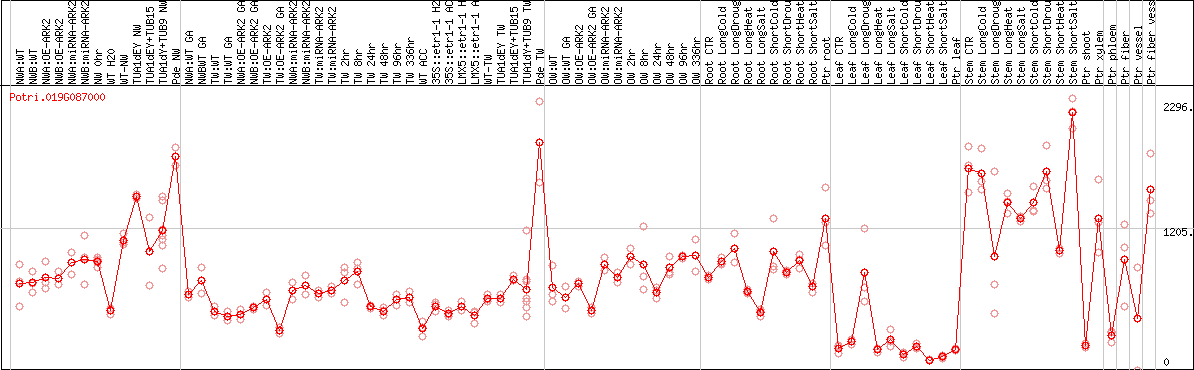
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G087000 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
