Potri.019G103600 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.019G103600.3 pacid=42773177 polypeptide=Potri.019G103600.3.p locus=Potri.019G103600 ID=Potri.019G103600.3.v4.1 annot-version=v4.1
ATGGCTGGAAAGAAGGCATCCGGGCGTGATGATGAGAACGCGGCTGTGGGTGGTGGAGGGAAGTCAAGGAAGAAGGGGTTGGTGATTGGTAATGATGATC
AGTATTCGATTGGTACTGAGTTGTCGGAGAAGAAGGGGAATTATGAGGGAGAGGAAGGGGAGGATGAGCAGGTGCCGGAAGTTGTGCTTGTTTCTGGGAA
AAAGAAGTTGAAAGGGAGGAAGAGTGGTGCTGGTGGGAATACTGGTTTTAATTGGTCGAATTTTGGGTTATTGGGAGGTGGTGATGAGGATGATAATGAT
GGTGATGATGAGGAGATATCCAAGTTGACTGGTGAGAAAGATAGTGATCAGGAGGAGGAGGATGAGCCGGTTGTGAGTTTTAAAGGGAAGAAAAAGGGTA
AGGGCAATAAGAAGAGTGGTGGTGGTGGTGGTAGTGTGTTTAGTGCTTCGGGATTTGATGCTACCGATGATGGTGAGAATGATGGAGAGGTGATGGATAA
AGATGGGGATGACGACGATGACGTGCCTGTCATTGAATTTACGGGGCAGAAGAAGAAGTCGTCAAAGGGTGGTAAGAAGGTTGCTGGGAGATTTTCATCC
AAGTCTAACAAAAAGAGTGGTAATAATAAGTTTAGTGCAGCATTGCTTGATGAGAAGAGTGAAGATCAGGTGGAAGAAACTTTGAAGCATGAGAAGAAGA
AGAAGAATAAGAGTGCAAGGGATGCTCAAGAAGAGGATGATTTGAATAAGATTCTTGCGGAGCTTGGTGACGCGCCACCTACTTCAAAATCCTTGGCACC
ACCTCCACAAGAGGAGAAACCCAATCTTCAGCCTGAACCTGTTTCAGTTCCAGATTCTGTGGCTGAGAAGGAGGGTGAAGATGAAAAAGAAGAGTCTGCT
GGTGCAAAGAAGAGGAAGAAGAAGAAGAAAGAGAAGGAGAAAGAGAAGAAGGCAGCAGCAAAGGAAGAAAGACTGGAAGAAGAAAAAGCTGAAACAAGTG
AGCCAAAAAAGACTGATGCAAAGGGTAAAGGTTCTGAGAAGAAGCTTCCAAAACATGTTAGAGAGATGCAAGAGGCACTTGCTAGGCGAAAAGAAATGGA
GGAGAGAAAGGCGAGGGAGGAAGAGGATAAGAGGAGGAAGGAAGAAGAGGAAAGGCTAAGGCAAGAAGAACTCGAGAGGCAAGCAGAAGAGGCTAGGCGT
AGGAAGAAGGAAAGGGAAAAGGAGAGACTCTTGAAGAAGAAACAGGAAGGCAAGCTTCTAACAGGGAAGCAGAAGGAAGAACAACGGCGACTAGAGTCTA
TGAGGTATCAAATACTTGCTAATGCAACTATTACCGTCCCTAATGCAGAAAGAGATGGTGGACCTACACGACGTCCTAGGTACCAGACTAGGAGGTCAAG
ACCAGTGCACGATCAAGCAAATGGTATCAGGATAGAGGAACCTGTAGAAGCTAAGGAAAAAGAGCAGGAGGAGGAGGAAGTAGTCCACGAGGTCGAATCC
TTAGAATTAGAGAATTTTGAGCCAGTGGAGGAAGAGAAAACAGAAGTTGCTAATGTACTTGTGGAGGATGGAATGGAGGAAGATGATGATGATGAGGAGT
GGGATGCAAAGAGCTGGGATGATGTAAATCTCAATGTTAAGGGTGCATATGATGATGATGAGGATTCTGAGCCTGAACCTGTCTTGAAGAAGGAAACAAA
AGCCCCTCTTGCAGCTGCACAGCCTGCAATTGCTGTGAGAAAACCTGTTACATCACAGCCAATGGACTCTCACGATGTTGAAGATAAGAAAAGTCAGGCT
GGGGTCGAAGTTTCTGACAAAAACAGGAAGAAAGATGCTGCTGCGAAAAGTAAAGGAGCAGTTTCAGATGCCATCCCTCATAAAGGTGAAGAGACCCTGC
GCTCACCAATTTGCTGTGTTATGGGCCATGTCGATACTGGCAAAACCAAGCTGCTGGATTATATTCGAGGTACCCATGTTCAGGAAGGGGAGGCTCGGGG
TATCACACAACAGATTGGTGCAACATTCTTTCCTGTTGAGAACATACGAGACAGGACTAAGGAATTGAAAGCTGATGCAAGATTAAATGTTCCTGGTCTG
CTAGTTATTGATACCCCTGGCCATGAAGCTTTCACCAATCTACGGTCACACGGTTCAGGTTTATGCGACATTGCCATTCTGGTTGTTAACATAATGCATG
GCTTGGAGCCGCAAACCATAGAATCACTCAATCTTTTGAGGAAGAGGAACACGGAGTTCATTGTTGCATTAAATAAGATCGACAGACTCTATGGATGGAA
AACACAACCTAATGCACCTATTAGGAAGGCATTGAAGCAGCAGTCAAAAGATGTGCAAAATGAATTCGACAGGAGGCTCATAGAGGTCATAACGCAGTTG
AAGGAGCAAGGATTAAACACTGAATTGTACTATAAAAACAAGGACATGGGAGAAACTTTTAACATCGTACCTACAAGTGCGATTAGTGGAGAAGGCATTC
CTGATTTGTTATTACTACTGATCCAATGGTCTCAGAAAACTATGGTTGAGAAACTCACGTTCAGAAACGAAGTGCAGTGTACGGTATTGGAGGTCAAGGT
TACTAAAGGCCATGGAACAACAATTGATGTTGTGCTGGTTAATGGTGTGCTCCATGAAGGCGATCAAATTGTGTTTTGCAGCTTACAGGGTCCCATTGTT
ACTACAATTCGAGCTCTGCTGACGCCTCACCCAATGCAGGAACTCCGGGTTAAGGGGACATATCTGCACCATAAGGAAATCAAGGCTGCACAAGGTATCA
AGATCACTGGACAGGGCCTTGAACATGCTATTGCTGGAACTAGTCTATATGTGGTCCGGCGTGATGATGATTTGGAAGAGGTCAAGGAATCTGCAATGCA
AGACATGAAGTCAGTCACGAGTCGGATTGATAGAAGTGGCGAGGGAGTTTGTGTACAAGCATCTTCCCTTGGCTCACTGGAAGCGTTGCTTGACTTCTTG
AAGTCCCTGGAACCAAGCATTCCTGTTAGTGGTATAGGCATTGGCCCAGTACATAAAAAGGATGTCATCAAGGCCAGTGTAATGCTTGAGAAGAAAAAGG
AGTATGCCAACATATTGGCATTTGGTGTTGAAGTGACACCTGAGGCTCGGGAACTTGCAGATAAACTAGGAGTGAAGATTTTCAAAGAGGATACCATTTA
TTGCTTTTCTAAAGAATTCAAGGCTTACATTCAGAATCTGAAGGAGGAAAGGAAGAGGGAAGCTGCTGGTGAAGCTATTTTCCCTTGTGTTCTTGAGATA
ATACCAGAGTGCATTTTCAACAAGAAAGCTCCAATCGTCTTGGGGGTTGATGTTCTTGAGGGCATACTCAAGGTTGGGACTCCAGTATGTGTTCTACAAA
AAGACTTCACTGACATTGGTCGGATTGCTTCCATTCGGTTTAACGAGAAAGCAGTGGATCATGCAAGGAAAGGCCAGAAGGTGACCATAAAGATCGTAGG
CACCAATCCGGAGGAGCAGCAAAAAATGCATGGCAGGCATTTCGACAATGATGATCAATTAGTCAGCCACATTACAAGAAGGTCAATCGATGTTCTTAAA
GCTTATTATCGGGATGATCTGTCTATGGAAGATTGGAAGCTGGTTCTTAAATTGAAGACCCTTTTCAAGATACAGTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.019G103600.3 pacid=42773177 polypeptide=Potri.019G103600.3.p locus=Potri.019G103600 ID=Potri.019G103600.3.v4.1 annot-version=v4.1
MAGKKASGRDDENAAVGGGGKSRKKGLVIGNDDQYSIGTELSEKKGNYEGEEGEDEQVPEVVLVSGKKKLKGRKSGAGGNTGFNWSNFGLLGGGDEDDND
GDDEEISKLTGEKDSDQEEEDEPVVSFKGKKKGKGNKKSGGGGGSVFSASGFDATDDGENDGEVMDKDGDDDDDVPVIEFTGQKKKSSKGGKKVAGRFSS
KSNKKSGNNKFSAALLDEKSEDQVEETLKHEKKKKNKSARDAQEEDDLNKILAELGDAPPTSKSLAPPPQEEKPNLQPEPVSVPDSVAEKEGEDEKEESA
GAKKRKKKKKEKEKEKKAAAKEERLEEEKAETSEPKKTDAKGKGSEKKLPKHVREMQEALARRKEMEERKAREEEDKRRKEEEERLRQEELERQAEEARR
RKKEREKERLLKKKQEGKLLTGKQKEEQRRLESMRYQILANATITVPNAERDGGPTRRPRYQTRRSRPVHDQANGIRIEEPVEAKEKEQEEEEVVHEVES
LELENFEPVEEEKTEVANVLVEDGMEEDDDDEEWDAKSWDDVNLNVKGAYDDDEDSEPEPVLKKETKAPLAAAQPAIAVRKPVTSQPMDSHDVEDKKSQA
GVEVSDKNRKKDAAAKSKGAVSDAIPHKGEETLRSPICCVMGHVDTGKTKLLDYIRGTHVQEGEARGITQQIGATFFPVENIRDRTKELKADARLNVPGL
LVIDTPGHEAFTNLRSHGSGLCDIAILVVNIMHGLEPQTIESLNLLRKRNTEFIVALNKIDRLYGWKTQPNAPIRKALKQQSKDVQNEFDRRLIEVITQL
KEQGLNTELYYKNKDMGETFNIVPTSAISGEGIPDLLLLLIQWSQKTMVEKLTFRNEVQCTVLEVKVTKGHGTTIDVVLVNGVLHEGDQIVFCSLQGPIV
TTIRALLTPHPMQELRVKGTYLHHKEIKAAQGIKITGQGLEHAIAGTSLYVVRRDDDLEEVKESAMQDMKSVTSRIDRSGEGVCVQASSLGSLEALLDFL
KSLEPSIPVSGIGIGPVHKKDVIKASVMLEKKKEYANILAFGVEVTPEARELADKLGVKIFKEDTIYCFSKEFKAYIQNLKEERKREAAGEAIFPCVLEI
IPECIFNKKAPIVLGVDVLEGILKVGTPVCVLQKDFTDIGRIASIRFNEKAVDHARKGQKVTIKIVGTNPEEQQKMHGRHFDNDDQLVSHITRRSIDVLK
AYYRDDLSMEDWKLVLKLKTLFKIQ
|
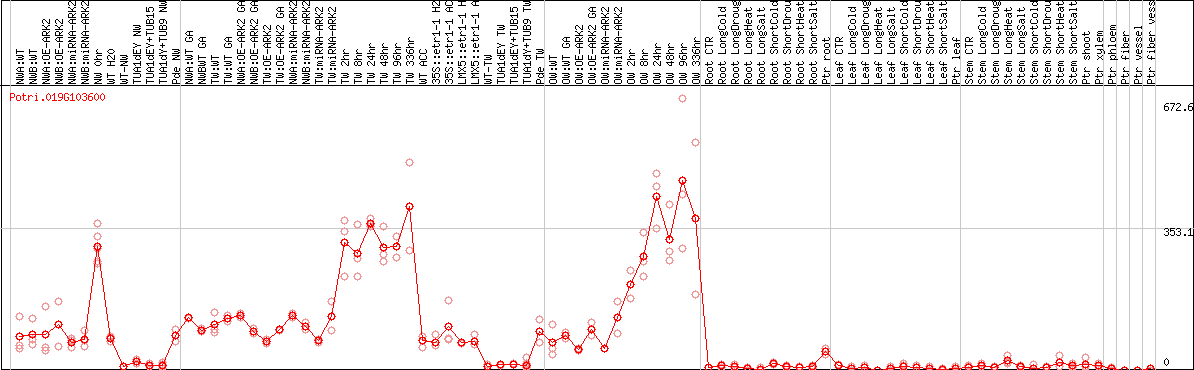
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G103600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
