Potri.019G119600 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.019G119600.3 pacid=42773339 polypeptide=Potri.019G119600.3.p locus=Potri.019G119600 ID=Potri.019G119600.3.v4.1 annot-version=v4.1
ATGGTAAGAAGAGGTCTTAACCAGAGAGATCTATGTGCAATGCATAGCATGCTATCAACCGTTTGCTTTTCTTATGCATTCTTATTATGCTCCTCCCTAT
TATGTTGCTTTGCTAGAGATACCATCACATATGCTGGTAATTTGATTAGTCATGATGGTGGAGAAACTCTTGTTTCAGCAGGAAAAAGGTTTGAACTTGG
GTTCTTTGCCCCCGAACAAAGTTCTGTTTACGGAAGCTATGTAGGAATTTGGTATTACAGGTCGCATCCACGAATAGTTGTGTGGGTTGCCAATCGAAAC
AGCCCACTTCTTGATGGTGGAGCAGTTTTGGCTGTCACAGATGATGGAAACCTCAAGATTTTGGATAAAAATGCAGATCCTTTTTGGTCTACAGCGCTTC
AATCAACATCAAAACCAGGGTATCGGTTGGCTAAGCTCTTAGATTCTGGGAACTTGGTTTTTGGTGATAGTAATACCCTTTCGACAACTATTCTATGGCA
AAGCTTTGAACATCCAACTGATACATTTCTTTCAGGTATGAAGATGAGTGGGAATCTAAAATTGACTTCATGGAAAAGTCAAGTTGATCCAAAAGAAGGT
AACTTCACATTTCAGCTAGATGGAGAAAAGAACCAGTTTGTTATCGTCAATGACTATGTAAAGCATTGGACTAGTGGGGAGTCAAGTGATTTCTTTAGCT
CAGAAAGAATGCCTGATGGAATAGTATATTTTCTATCGAACTTCACCAGATCTGTTCCAAATTCAAAAGGCAGAAGAACAACTCGATCACCCTCGGATTA
TAATAATACAAGGATTAGGTTGGATGTTAAAGGGGAGCTACAATACTGGAACTTCGATGTGTATACAAATTGGTCTTTGCAATGGTTTGAGCCGAGAGAC
AAATGCAATGTGTTTAATGCCTGTGGGAACTTTGGCAGCTGCAATCTGTACAACATGTTGGCTTGCAGGTGTTTGCCAGGATTTGAGCCTATATCACAGG
AAAATTGGAGGAATGGAGATTTCTCTGGTGGTTGCATCAGAAGCGCTGCCGGATGTGGGAAGAATGACACCTTCTTGAGCTTGAAGATGATGAGAGTTGG
GCAACCAGACACTTCATTTGTGGTTGAGGATGAAAAGCAATGCAGAGAGGAGTGTCTCAATAAGTGCCAATGCCGAGCTCATTCCTTTGTGAAGGGGGAA
GTCAATATGCGAAGGGACAGACCGCCTAGCGACAACAGTTGCTTGATCTGGATGGATGATCTTAAAGATCTTCAGGAGGAATATTCTTATGGTGGGCCCG
ATCTCTTTGTTCGTGTCACCATTGCTGACATAGAATCAAAGGCTAAAAGTTGTGAGCCTTGTGGCATAAATGTGATTCCTTATCCTCTAAGTACTGGATC
AGATTGTGGTGACCCCATGTACTTCAGTTTCTACTGCGATAATTCCACAGGAAAACTCAGCTTTAAGACACACAACGGCACCTATAATGTCACTACCATC
AATCAAGATAGAAGAACATTTGCTATCCAAGAGAAAGATGTAGCTGATTGTAATGCCAGTACAAGAGGTCAAATTGGGAAGTTCAACACGTCATCTCCAT
TTAAGATGAACGCCTCTAAACGCTGGTGTGACTCCAATGTTTCATCTCAAGGTCTGGTAGAAATAGATATTGGTTGGGAGCCACCTCCAGAGCCAGTCTG
CTCTTCATCTTCGGACTGCGACGATTGGCCAAATTCAACTTGCAATGTAACAGGAAATCGTACAGCGAGATGCCTCTGCAATTCAAATTTTTGGTGGGAT
GGCATGGCTCTAAATTGCGTTCAAGTTGTCGATGGACAAGCAGGAGGATATTCGAGAAAAAAGAAACCATTGTCTTTGATTGTAGGAGTTACAATTGCTA
GTGTCATAGTTCTATCAAGCATCTTCCTTTACACCTGCATTTTTATGAGAAAGAAGGCCAAACGACGAGAAAGCCAGCAAAATACTGAAAGAAATGCAGC
TCTTTTGTATGGCACTGAGAAACGGGTCAAGAACTTGATAGATGCAGAAGAGTTCAACGAAGAAGATAAGAAGGGCATAGATGTACCTTTATTTGATTTA
GACAGCATACTAGCGGCTACGGATTATTTTTCAGAAGCAAACAAGCTTGGAAGAGGGGGCTTCGGACCTGTTTACAAAGGAAAATTTCCAGGAGGTCAAG
AAATTGCAATAAAACGACTCTCTAGTGTTTCAGGACAAGGCCTAGAGGAGTTTAAGAATGAGGTGATTTTGATTGCTAGATTGCAGCATCGGAATCTTGT
CAGATTAGTAGGTTACTGCATAAAAGGAGATGAAAAGATTTTACTCTATGAATATATGCCCAACAAAAGCTTAGATTCCTTCATATTTGATCGAGATCTC
GGAATGTTACTGGATTGGGAGATGCGCTTAGATATCATTTTAGGAGTTGCTAGAGGTCTTCTTTATCTCCACCAAGATTCTAGACTCAGGATTATTCATC
GTGATATGAAAACGAGCAACATTTTATTAGATGCGGAAATGAATCCAAAAATTTCTGACTTTGGATTGGCAAGGATGTTCGAAGGCAAACAAACCGAAGG
GAGCACCAACAGAGTTGCTGGAACTTATGGCTACATGTCGCCGGAGTACGCATTAGATGGCTTATTCTCTGTCAAATCTGATGTTTTTAGTTTCGGCGTG
GTTGTGCTTGAGATTCTTAGTGGGAAAAGGAACACAGGATATTTTAATTCTGATGAAGCCCAAAGCCTTCTAGCTTATGCTTGGAGATTATGGAGAGAAG
ACAAGTCATTGGATTTAATGGACGAGACATCACGTGAAAGTTGCAACACCAATGAGTTTCTGAGGTGTGTAAATGCTGCACTCTTGTGCGTACAAGATGA
TCCATCCGACCGTCCGACCATGTCAAATGTTGTTGTGATGCTTAGCAGTGAAACTGCAAACTTGCCGGTTCCTAAGAACCCGGCCTTTTTTATACGAAGA
GGCCTTTCTGGCACAGCTTTTGCTTCAAGCGAACAAGAGGGAGGCCTTTCTGGCACAGCTTCATCTTCAAGTAAACAAGAAAGAAGCATCGATACAACGA
TTGCTTCAGACGAAGTTAGATAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.019G119600.3 pacid=42773339 polypeptide=Potri.019G119600.3.p locus=Potri.019G119600 ID=Potri.019G119600.3.v4.1 annot-version=v4.1
MVRRGLNQRDLCAMHSMLSTVCFSYAFLLCSSLLCCFARDTITYAGNLISHDGGETLVSAGKRFELGFFAPEQSSVYGSYVGIWYYRSHPRIVVWVANRN
SPLLDGGAVLAVTDDGNLKILDKNADPFWSTALQSTSKPGYRLAKLLDSGNLVFGDSNTLSTTILWQSFEHPTDTFLSGMKMSGNLKLTSWKSQVDPKEG
NFTFQLDGEKNQFVIVNDYVKHWTSGESSDFFSSERMPDGIVYFLSNFTRSVPNSKGRRTTRSPSDYNNTRIRLDVKGELQYWNFDVYTNWSLQWFEPRD
KCNVFNACGNFGSCNLYNMLACRCLPGFEPISQENWRNGDFSGGCIRSAAGCGKNDTFLSLKMMRVGQPDTSFVVEDEKQCREECLNKCQCRAHSFVKGE
VNMRRDRPPSDNSCLIWMDDLKDLQEEYSYGGPDLFVRVTIADIESKAKSCEPCGINVIPYPLSTGSDCGDPMYFSFYCDNSTGKLSFKTHNGTYNVTTI
NQDRRTFAIQEKDVADCNASTRGQIGKFNTSSPFKMNASKRWCDSNVSSQGLVEIDIGWEPPPEPVCSSSSDCDDWPNSTCNVTGNRTARCLCNSNFWWD
GMALNCVQVVDGQAGGYSRKKKPLSLIVGVTIASVIVLSSIFLYTCIFMRKKAKRRESQQNTERNAALLYGTEKRVKNLIDAEEFNEEDKKGIDVPLFDL
DSILAATDYFSEANKLGRGGFGPVYKGKFPGGQEIAIKRLSSVSGQGLEEFKNEVILIARLQHRNLVRLVGYCIKGDEKILLYEYMPNKSLDSFIFDRDL
GMLLDWEMRLDIILGVARGLLYLHQDSRLRIIHRDMKTSNILLDAEMNPKISDFGLARMFEGKQTEGSTNRVAGTYGYMSPEYALDGLFSVKSDVFSFGV
VVLEILSGKRNTGYFNSDEAQSLLAYAWRLWREDKSLDLMDETSRESCNTNEFLRCVNAALLCVQDDPSDRPTMSNVVVMLSSETANLPVPKNPAFFIRR
GLSGTAFASSEQEGGLSGTASSSSKQERSIDTTIASDEVR
|
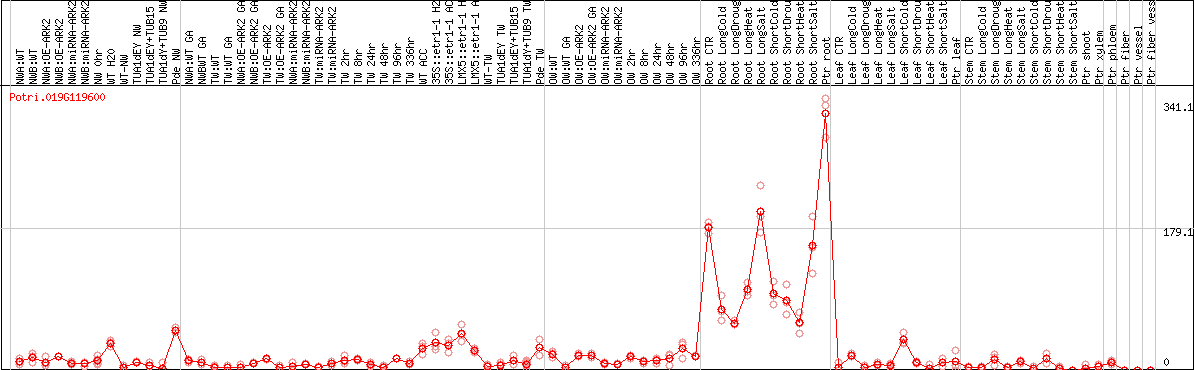
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G119600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
