Potri.019G126000 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.019G126000.2 pacid=42774364 polypeptide=Potri.019G126000.2.p locus=Potri.019G126000 ID=Potri.019G126000.2.v4.1 annot-version=v4.1
ATGGGCAGACAGAAAGGAGAAGCAGCTGCTAACAGAAGCAAGTCTCGCGCTTCCAGTAGCAGTCTTGCGGCGTCTCTAGTGCCGTCGGGTCCGGCGGCGG
TAACGGTGGGGTTCGGTGGTTATATTGGTAGCTCTCGTTTTGATACAGATGATACTACCGCCTTTTTGGATATAGATGGCGAAGTGGCACAACATGTAAA
GCGATTAGGAAGGAAGGATCCTACTACAAAGCTTAAAGCTTTGCAAACTCTATCTGCTCTGTTCAAGGAAAAGTCTGGAAAGGAAATAGTTCTGATTATT
CCACAATGGGGTTATGAATACAAGAAGCTGTTGTTGGACTATAATAGAGAAGTTCGACGAGCCACAAATGAGACCATGACAAATCTTGTTACAGCAGTTG
GAAGAGATTTAGCTCCGTATCTGAAGTCTTTGATGGGACCGTGGTGGTTTTCTCAATTTGATACAGTTCCTGAAGTTTCTTTAGCTGCCAAAAGATCATT
GGAGGCAGCCTTCCCAGCTCAGGAAAAGAGATTAGATGCCTTAATTCTATGCACATCTGAGATATTTATGTATTTGGAAGAAAACCTGAATCACACACCA
CAAAGTATGTCATCAGATAAAGTAACTGCTTTGGATGAATTAGAAGAGATGTACCAGCAGGTGATATCTTCGTCATTACTGGCATTGGCCACTCTTCTTG
ATGTTTTGGTCTGTATGCAATCTGAGAGACCTGGTTTTGAGAACATATCCTCAGAACCAAAACATGCTTCAAAGGCTAGGGAAACTGCTATTTCTTTTGG
TGAAAAGTTGTTTTCTACTCAGAACTATTTTCTTGACTTTCTGAAGTCCAAAACCCCTGCAATCCGGTCAGCGACATATTCTGCATTAAAGAGTTTCATT
AAAAATATTCCTGATGCTTTTAATGAAGGAAATATGAAAACTCTTGCAGCTGCAATACTTGGTGCTTTTCAGGAAAAGGATCCTACTTGTCATTCATCAA
TGTGGGATGCAATATTATTATTTTCAAAAAGATTTCCTGACAGCTGGACTTCGTTCAATGTTCAGAAAACTGCAATAAACCGGTTGTGGCATTTTCTTAG
AAATGGGTGCTTTGGATCTCAGCAGGTCTCTTATCCTGCTCTAGTTATATTGCTAGACATTCTGCCTCCAAAAGCAATCTCTGGAGAAAAGTTTTTTATT
GATTTTTTCCAAAACCTATGGGATGGAAGGAACCCATCCAATGCAACAAATCCGGATCGTCTAGCATTTTTTCGTGCACTTAAAGAATGTTTTCTATGGG
GGCTATGTAATGCATCTAGGATTTGTGATGATTCAGATTCCACACATCATTTTCAAGTCAGTCTTGTGGACAATATTCTTGTGAAACTTCTGTGGCAAGA
ATACCTATTTTCTGTCAGATTGAAGAATCAGGACGGAGTAACCTCTGGAGCACCTGGGAATTCTTTGGAGCATGGAAATTTACCTTTTCACCATAAGTCA
GTAGAACCACTGAAGATCAAGTATTCTAGGAGTTATTTTCAAGAGTTGGGAAAATGTATTGTGGAAATTCTATCTGGCGTTTATTTACTGGAGCATGATC
TGCTTTCTACTTTCTCTGTGGTATTTAAGGAAAACTGCCTCCGGATGTTTCAGCCAATGGGAAACACAGAAAGTACTACTGAGAATGTAGAACAGGTCAT
TAAGTTCTTGTCACTGCTGGAAAAACATTCTGTGCGAAAAGGTGAAAGTTGGCCCTTGGTTTATGTAGTGGGACCAATGTTGGCGAAGTCTTTCCCGTTG
ATAAGATCTCATGACACACCAGATGGTGTGAGACTTTTATCAGTTGCAGTTTCATTATTTGGCCCGCAGAAGATAGTCCAAGAACTTTGCATCTCCAATG
AAGCGAATTCCTCCTATTATGTACCTGCCCACAAAGACAAAGAATTGGGACCAGAGCTTTTCATGCAAGTTTTTGAAGGAACATTTGTTCCTTGGTGTTT
GCTTGAATATAATTCCTCCCCCAATGCTCGACTTGATCTATTGCTTGCATTGCTCAATGATGAATATTTCTCTGAGCAATGGCAGATGATCCTTTCATAT
GCAATTAATCAAGAAAAATCCGAATCTGAGCCTGGGCCACAGGAAGTTCATTACTTAGATCTGTTGGCAATGCTTTTAGAAAAAGCGAGAACTGAAATTG
CAAGGAGGAAGATGAATAATGATTTCATTCACCAGTTCTGGTTCACACCAGACAAGTGGCAGCATGAGCTTCTGGAATCAGCTGCAGTTGCTGTTGCCTG
TTCACCTTCTCCCCATATGACTTCTTCTGCTCGCTTTTTGTGTGCGGTCCTTGGTGGCTCAAGCAAAGACAATTGTATCTCGTTCGCCTCCAAAAATGCA
ATGGTTCTGATATTTACGATAGTTTTCAAAAAGTTGGTTGCTTTTGGTCTTGAGTCCTCTTTCAGTGTAGTGAGAGATTCATGTGCTTTATTAGTTGCTG
GATCAACCAATTTCGCAGTGGAAAATGAAAGTTCCATAAACAAAACTGAGACAGCTCAGTTTGCTCTGAAAGTACTTGGTGGCAGCTTCTTTTGCCTAAA
GACAGTCAGCAATGAAATTGAGCTGGTTTCAGGTATTTTGACTTTGGTATTTATCATAGGCTGGGAGAATAGCTTGGATACATTGGAAGAAGATGTCCTT
AATGATGATTCAAAGGAAAAAATCAAGGGTAGGTTGAGATTTGGTGAATCTTTGAATGGTTTCTGTTCCAAGATGAATGATGAATTTTGGAAAAGCCTCG
GCATTGACAATAGAAAGAGATTGGGGAGTAATTTGGTTCGCTTTATTAGGTCTGTCATATTTAAGGAAGATAAACTTGGTGCGGATAAGATAACCACTCT
GTGCTTTTCTTGGGTTCTTGAAGTTCTTGAATGTCTCTGCCATGACCATGATGAGGAACAAAATCTGTTAGACCAGCTTCTTAGCAAGAATGATACTTGG
CCGGTTTGGATAATTCCAGATTTTAGTGCTCCAAAAGGATTGGTCAATTTAAATGCTGGAGCAGTCTCTGTTGATATCTATGCCACTGGGAATCTCAAAT
TTGTTTCTTTAGTTGACAAGCTCATCTTGAAAATTGGGATCAATAGAGTTATAACAGGCTATGTTGAAAATACTCTTTCAACCCCCCTGAAGGAAGCAGC
TAAAGAAGAGATTACTTCTCGAGCCTGGCTTGCTGCTGAAATATTATGCACATGGAAATGGCCAGGAGGTAGTGCTGTAGCTTCTTTCTTGCCTCTTCTG
AGTGCAGGTTGCCGGAGTGGAAATTATCCTTTCCAGGAAAGCCTTTTAGATTCCATCTTCAACATCTTACTTGATGGTGCTCTTGTTCACGGAGAAAGTG
GTACACAAAGTTCATTTAATCTATGGCCTGCTTTTGGTGATGAATTGGAAAAGGTTGAAGAACCATTCTTGAGAGCTCTTTTATCTTTATTAGTCAATCT
GTTCAAAGAAAATATATGGGAAGGAGACAAAGCTATAAGACTCTTTGACCTACTCATACATAAGCTTTTTATCGGTGAAGCAGTAAACCAAAATTGTTTG
AAGATTCTGCCTGTGATTGTTAGTGTTCTTGTCCATCCATTATGTCAGAGAAGCATCGAATCGGAGGAATCTAACGGTGATTCTCAAGTTGCTTCTTTGG
GTGAAAAACGGATGCAGGATACTGTTAAAGACTGGCTACGAAGGCTTTTATCCTACCCACCTTTAGTTACATGGCAGGCAGGACAAGATATGGAAGAATG
GTTCCAACTGGTTATTGCTTGCTATCCTCTTAGTGCAATGGATGATACAAAATCATTGAAGCTGGTGAGGGAAATAAGTCCTGAGGAGAGAATGCTCATA
CTTGATTTATTCAGAAAGCAGAGACATGGTGTTAGTGCATTAGTAGCTTCAAATCAACTACCACTGTTCCGGATGTTGCTATCAAAGTTAATGGTTCTTT
CGGTTGGTTATTGCTGGACTGAATTTACTGAAGAAGATTGGGAGTTTTTCTTCTCAAATTTGAGGTCCTGGATTCAGTCAGCGGTTGTGATCATGGAGGA
AGTCACGGAAAATGTGAATGATCTTATCACCAACAGCTCTACTTCTGAAAATTTGGATGTTTTTAAGAACCTTGAGAAGATTGTTTTGATCCCGGACTCT
TATCCTATAACTGTTGCTATTAATGCCTTGGCATCCTTTTCTTTATTTTGTGCTATTTTGGAACTCCAACAACCTGCTGAGGATAACCCCCTGAGAGCAG
AAAGATGGGATTCAACCAGAGATAGAATCCTAGAAGGCATTCTTCGGTTGTTCTTTTGCACAGGCATTGCGGAGTCCATAGCAAGTTCCTACAGTGTTGA
AGCTGCATCCATTGTAGCAGCAACTCGGTTTAATAATCCTTACTTCTGGGAGTTGGTGGCCTCAAACGTGGTTAAATCATCGCAACATGCTAGAGACAGA
GCAGTGAAATCAGTTGAATTTTGGGGGTTAATCAAAGGGCCGATTAGCTCTTTGTATGCTATCCTGTTCTCCTCTACTCCATTTCCGCCATTGCAGTTTG
CTACTTATGTTATTCTCTCAACTGCACCTATTTCACAGTTGGCTATTCTTGAAGAAGACACTGCATGTTCCTTGGATGGCGAGACTAGTGGTGACCGGAA
TTCAGGTGCTCTTGAAATGTCATCAGAAAGAAACATACGTTTGAAGGAGGAATTATCTCTTATGATTGAAAAGCTACCTGATGAAGTTTTTGAAGTCGAT
TTAATATCACAAGAACGGGTAAATGTGTTCCTTGCTTGGTCTTTATTGCTATCACATTTGTGGTCATTATCATCATCATCATCTGCAAAGGAGCAATTAG
TCCAGTATGTGCAAGATTCTGCTAATTCATTGATATTAGATTGCCTTTTCCAGCATATTCCTCTGGAATTGTGTCTGGCACATAATCTTAAGAAGAAAGA
CATGGAGCTTCCTGTAGACATTTCAGAGGCTGCTAGTGCAGTAAAAACTGCCATCACAACAGGGTCACTATTGTTTTCCATTGAAACACTGTGGCCTATT
GAACCCAAGAAAATGACTTCGCTTGCTGGGGCTTTGTTTGGACTGATGCTTTGCATTCTTCCTGCTTATGTTCGGGGATGGTTTACTGACTTGCGTGACC
GCACTGCATCATCTCTTATTGAATCATTTACAAGAACATGGTGTAGTCCTCCTCTTATTGTGAACGAACTGTCTCAGATAAAGAAAGCCAATTTTGCTGA
TGAGAATTTTTCAGTCAGTGTGAGCAAATCAGCAAATGAAGTCGTTGCTACATATATGAAGGATGAAACAGGAATGGATCTAGTTATTCGTCTTCCTCCA
TCTTATCCATTGCGCCCTGTGGATGTTGAATGTATGCGAAGTCTGGGAATCAGTGAGGTGAAGCAGAGGAAATGGTTGATGTCAATGATGTTGTTTGTTC
GCAATCAGAATGGAGCTTTAGCAGAAGCTATACAGACATGGAAAAGCAATTTCGACAAGGAATTTGAAGGTGTCGAGGAGTGCCCAATATGTTATAGTGT
CATTCATACGACCAACCACAGCCTTCCTCGGCTTGCTTGCAGAACTTGCAAGCACAAGTTCCATTCGGCTTGCCTCTACAAGTGGTTCTCAACCTCTCAC
AAATCATCATGTCCACTGTGCCAGTCTCCATTCTGA
|
|||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.019G126000.2 pacid=42774364 polypeptide=Potri.019G126000.2.p locus=Potri.019G126000 ID=Potri.019G126000.2.v4.1 annot-version=v4.1
MGRQKGEAAANRSKSRASSSSLAASLVPSGPAAVTVGFGGYIGSSRFDTDDTTAFLDIDGEVAQHVKRLGRKDPTTKLKALQTLSALFKEKSGKEIVLII
PQWGYEYKKLLLDYNREVRRATNETMTNLVTAVGRDLAPYLKSLMGPWWFSQFDTVPEVSLAAKRSLEAAFPAQEKRLDALILCTSEIFMYLEENLNHTP
QSMSSDKVTALDELEEMYQQVISSSLLALATLLDVLVCMQSERPGFENISSEPKHASKARETAISFGEKLFSTQNYFLDFLKSKTPAIRSATYSALKSFI
KNIPDAFNEGNMKTLAAAILGAFQEKDPTCHSSMWDAILLFSKRFPDSWTSFNVQKTAINRLWHFLRNGCFGSQQVSYPALVILLDILPPKAISGEKFFI
DFFQNLWDGRNPSNATNPDRLAFFRALKECFLWGLCNASRICDDSDSTHHFQVSLVDNILVKLLWQEYLFSVRLKNQDGVTSGAPGNSLEHGNLPFHHKS
VEPLKIKYSRSYFQELGKCIVEILSGVYLLEHDLLSTFSVVFKENCLRMFQPMGNTESTTENVEQVIKFLSLLEKHSVRKGESWPLVYVVGPMLAKSFPL
IRSHDTPDGVRLLSVAVSLFGPQKIVQELCISNEANSSYYVPAHKDKELGPELFMQVFEGTFVPWCLLEYNSSPNARLDLLLALLNDEYFSEQWQMILSY
AINQEKSESEPGPQEVHYLDLLAMLLEKARTEIARRKMNNDFIHQFWFTPDKWQHELLESAAVAVACSPSPHMTSSARFLCAVLGGSSKDNCISFASKNA
MVLIFTIVFKKLVAFGLESSFSVVRDSCALLVAGSTNFAVENESSINKTETAQFALKVLGGSFFCLKTVSNEIELVSGILTLVFIIGWENSLDTLEEDVL
NDDSKEKIKGRLRFGESLNGFCSKMNDEFWKSLGIDNRKRLGSNLVRFIRSVIFKEDKLGADKITTLCFSWVLEVLECLCHDHDEEQNLLDQLLSKNDTW
PVWIIPDFSAPKGLVNLNAGAVSVDIYATGNLKFVSLVDKLILKIGINRVITGYVENTLSTPLKEAAKEEITSRAWLAAEILCTWKWPGGSAVASFLPLL
SAGCRSGNYPFQESLLDSIFNILLDGALVHGESGTQSSFNLWPAFGDELEKVEEPFLRALLSLLVNLFKENIWEGDKAIRLFDLLIHKLFIGEAVNQNCL
KILPVIVSVLVHPLCQRSIESEESNGDSQVASLGEKRMQDTVKDWLRRLLSYPPLVTWQAGQDMEEWFQLVIACYPLSAMDDTKSLKLVREISPEERMLI
LDLFRKQRHGVSALVASNQLPLFRMLLSKLMVLSVGYCWTEFTEEDWEFFFSNLRSWIQSAVVIMEEVTENVNDLITNSSTSENLDVFKNLEKIVLIPDS
YPITVAINALASFSLFCAILELQQPAEDNPLRAERWDSTRDRILEGILRLFFCTGIAESIASSYSVEAASIVAATRFNNPYFWELVASNVVKSSQHARDR
AVKSVEFWGLIKGPISSLYAILFSSTPFPPLQFATYVILSTAPISQLAILEEDTACSLDGETSGDRNSGALEMSSERNIRLKEELSLMIEKLPDEVFEVD
LISQERVNVFLAWSLLLSHLWSLSSSSSAKEQLVQYVQDSANSLILDCLFQHIPLELCLAHNLKKKDMELPVDISEAASAVKTAITTGSLLFSIETLWPI
EPKKMTSLAGALFGLMLCILPAYVRGWFTDLRDRTASSLIESFTRTWCSPPLIVNELSQIKKANFADENFSVSVSKSANEVVATYMKDETGMDLVIRLPP
SYPLRPVDVECMRSLGISEVKQRKWLMSMMLFVRNQNGALAEAIQTWKSNFDKEFEGVEECPICYSVIHTTNHSLPRLACRTCKHKFHSACLYKWFSTSH
KSSCPLCQSPF
|
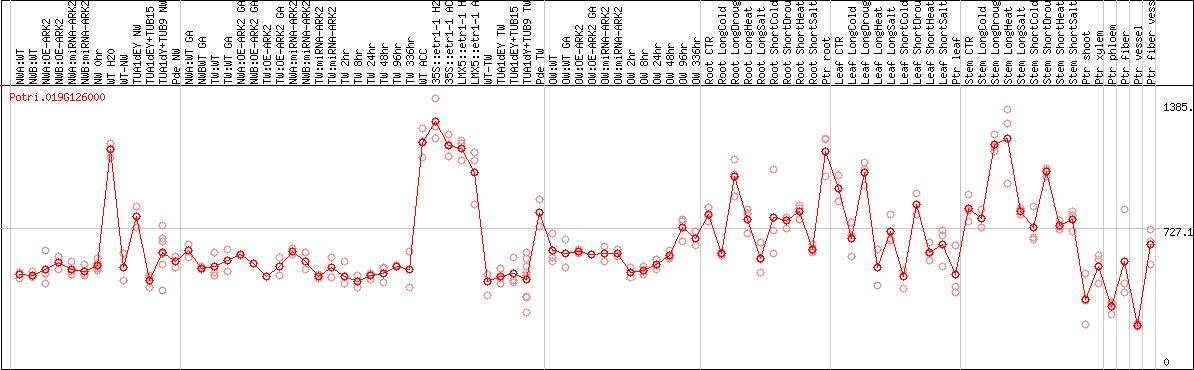
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G126000 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
