Potri.019G129100 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.019G129100.1 pacid=42773087 polypeptide=Potri.019G129100.1.p locus=Potri.019G129100 ID=Potri.019G129100.1.v4.1 annot-version=v4.1
ATGACGTCCCTGCTACACAAACCATTCTTCTCAATCTTTCTCCACGTCCTGTTTTTGCTCCTTCACATGTTCTACTACTCTTCTTTCTTTGTTATTGCTG
ATCACACTTCCTCTAAAACTTCAATATTTGGTACGGCTATAAGTGCTGCCAATTCCAAAGTTGCTGGAGGTAATAATACAGAAGCTGAGGCTCTTCTTAA
ATGGAAGGCAAGTCTTGACAACCAAAGCCAATCTCTTCTTTCTTCTTGGTTCGGCATCAGCCCTTGCATTAACTGGACTGGAATCACTTGCGACAGTTCT
GGAAGCGTCACCAACTTGAGCCTTCCACACTTTGGCTTGAGAGGTACGCTTTATGATCTCAACTTCTCATCCTTCCCCAATCTTTTCTCTCTCAATCTCC
AGAGGAATTCAATTCATGGAACCGTTCCATCAGGCATTGATAATCTACCCAAGATTACTGAGCTCAATTTATGTGATAATAATCTCACGGGAAGCATCCC
ATCTAAGATAGGCTTAATGAAAAGTCTCAACATCTTGTATCTGTGTGGAAATATTCTTAGTGGCTCCATCCCATGTGAAATTGGAAAGTTGACGTCTCTT
TCCCTTCTTTCATTATCTGCAAATAATCTCACTGGTGTGATCCCCTTCTCTATTGGAAACTTGACAAACTTATCCTTGCTTCACCTCTTTCAAAACCAGC
TCTCTGGTCCCATCCCTTCTTCTATCGGGAACATGAGCTTTCTCATTGATCTACAACTACAACAAAATAATCTCACTGGTTTTATCCCCTCCTCTGTTGG
AAATTTGAGGAGTCTATCGATACTCTATCTTTGGGGTAATAAACTCTCTGGTTCCATTCCTGGAGAAATTGGGTTGCTAGAATCACTCAATGATCTTGAC
TTTTCAAGTAATAATTTAACCGGTGCAATCCCTAATTCAATAGGAAACTTAACGAACTTATCATTTTTCCACCTTTTTCAAAATCAACTCTCTGGTCCCA
TTCCTACCTCTATTGGGAATATGATTATGCTCATTGATGTAGAGCTCGGACAAAATAATCTCATTGGTTCAATCCCCTCCTCTGTTGGAAATTTGAGGAA
CCTATCAATATTCTATCTTTGGCGTAACAAACTCTCTGGTTCCATTCCTAAAGACATTGGATTGCTAGAATCTCTCTATGATCTTGACTTTTCAAGTAAT
ATTTTAACCGGTGAAATCCCTAATTCAATTAAAAACTTGACAAATCTATCTTTTCTCTCCCTTTATCAAAACCAACTATCTGGTCCTATACCTTCTTTTA
TTGGGAACATGAGCATGCTCGTTGATATAGAACTAGATGAAAACAATCTCAATGGTTTGATCCCCTCCTCTATTGGAAATTTAAAAAACCTATCCTTTCT
CTATCTAGGTGAAAATAATTTATATGGATATGTTCCTTCAGAAATAGGAAAGCTTAAATCTCTAGAGAAATTGACCTTCGGTGAGAACAAACTTCGCGGA
TCATTGCCCCTAAAGATGAACAATCTTACCCATTTGAAGTTTTTGGATTTGTCTTACAATGAATTCACGGGTCATTTGCCACAAGAGCTATGCCATGGAG
AAGTACTTGAAAGATTTATTGCTTGTAACAACTATTTCTCCGGTTCCATCCCGAAAAGCTTGAAAAACTGTACCGGTTTACACCGACTAAGACTTGATAG
GAATCAATTGACAGGAAATATTTCTGAGGATTTTGGCATCTATCCACATTTGAATTATGTTGATTTGAGTTACAATAATTTTTATGGCGAGCTTTCTTTG
AAATGGGGAGATTATCGTAACATAACGAGCTTGAAAATCTCAAACAATAATGTTTCTGGGGAGATACCAACAGAGCTCGGGAAGGCTACTCAATTACAAT
TGATTGATCTGTCCTCAAATCACTTAGAAGGAACCATCCCAAAAGAATTAGGGGGGTTGAAGCTGTTGTACAACCTTACTCTTAGCAACAACCATCTTTC
AGGTGCCATTCCCTCAGACATCAAGATGCTTTCTTCCCTCAAAATCCTTGATTTAGCATCTAATAATCTAAGTGGATCGATTCCAAAACAACTTGGGGAG
TGCTCAAATTTATTGTTGCTGAACCTCAGTAATAATAAATTTACAAATAGCATTCCACAGGAGATGGGCTTTCTGCGTTCTCTTCAAGATCTTGATCTTA
GTTGCAATTTCTTAGCTCAAGAGATACCGTGGCAACTTGGGCAACTGCAAATGTTGGAAACTCTGAATGTCTCTCACAACATGCTTTCTGGATTGATTCC
GCGCACTTTCAAAGATCTATTAAGCTTGACAGTTGTGGATATATCATACAATGAGCTACATGGCCCTATTCCCGATACCAAAGCCTTCCACAACGCTTCA
TTTGAGGCATTAAGGGATAATATGGGTATATGTGGCAACGCCAGTGGTCTAAAGCCATGCAATCTTCCAAAAAGCAGCAGAACTGTTAAGAGAAAGAGCA
ACAAACTTGTTATTTTGATTGTACTCCCTCTTTTAGGAAGCTTACTATTAGTGCTTGTTGTGATTGGAGCTTTATTCATTCTCCGCCAAAGAGCTAGGAA
GAGAAAAGCTGAGCCAGGGAATATTGAACAAGATCGAAACTTATTCACGATATTGGGCCATGATGGGAAGTTGTTGTATGAGAATATCATTGCAGCTACC
GAGGAATTCAATTCCAACTACTGCATTGGTGAAGGAGGATATGGAACTGTTTATAAAGCAGTGATGCCAGCAGAACAGGTGGTTGCTGTGAAGAAACTTC
ACCGGTCACAAACAGACAAGTTGTCCGATTTTAAAGCTTTTGAAACGGAAGTTTGTGTTTTGGCAAATATTCGACATCGAAATATCGTGAAGCTGTATGG
ATTTTGCTCACATGCAAAGCACTCTTTTTTGGTTTATGAGTTCATAGAAAGAGGAAGCTTGAGAAAGATCATAACCAGTGAGGAACAAGCAATTGAGCTG
GATTGGATGAAAAGGCTAAATGTTGTGAAAGGGATGGCTGGAGCTTTATCCTATCTACACCATTCTTGCTCTCCTCCGATCATTCATCGAGACATTACCA
GCAATAATGTCCTTCTGGATTTGGAATATGAGGCACATGTCTCTGACTTTGGCACAGCTAGACTGTTGATGCCTGACTCATCCAATTGGACCTCATTTGC
AGGTACTTTTGGATATACTGCTCCAGAGCTAGCTTACACGATGAAAGTGACTGAAAAATGTGATGTCTATAGCTTCGGAGTGGTAACAATGGAAGTGATG
ATGGGAAGGCACCCAGGCGATCTCATCTCCACAATCTCATCACAAGCATCTTCCTCATCATCTTCGAAGCCACCGATTTCTCAACAGACATTGTTGAAGG
ATGTGCTTGACCAACGCATCTCGCTTCCTAAAAAAGGAGCAGTAGAGGGTGTTGTCCATATCATGAAGATAGCACTTGCATGCTTGCATCCCAATCCGCA
ATCCCGACCTACAATGGGAAGAATTTCTTCGGAGCTTGTGACTCAGTGGCCTTCTCTGCCAAAGGAATTTTACACAATCAGTTTGGAAGATCTGTTCTTA
CATACAGTTAGTGTGGTCGACTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.019G129100.1 pacid=42773087 polypeptide=Potri.019G129100.1.p locus=Potri.019G129100 ID=Potri.019G129100.1.v4.1 annot-version=v4.1
MTSLLHKPFFSIFLHVLFLLLHMFYYSSFFVIADHTSSKTSIFGTAISAANSKVAGGNNTEAEALLKWKASLDNQSQSLLSSWFGISPCINWTGITCDSS
GSVTNLSLPHFGLRGTLYDLNFSSFPNLFSLNLQRNSIHGTVPSGIDNLPKITELNLCDNNLTGSIPSKIGLMKSLNILYLCGNILSGSIPCEIGKLTSL
SLLSLSANNLTGVIPFSIGNLTNLSLLHLFQNQLSGPIPSSIGNMSFLIDLQLQQNNLTGFIPSSVGNLRSLSILYLWGNKLSGSIPGEIGLLESLNDLD
FSSNNLTGAIPNSIGNLTNLSFFHLFQNQLSGPIPTSIGNMIMLIDVELGQNNLIGSIPSSVGNLRNLSIFYLWRNKLSGSIPKDIGLLESLYDLDFSSN
ILTGEIPNSIKNLTNLSFLSLYQNQLSGPIPSFIGNMSMLVDIELDENNLNGLIPSSIGNLKNLSFLYLGENNLYGYVPSEIGKLKSLEKLTFGENKLRG
SLPLKMNNLTHLKFLDLSYNEFTGHLPQELCHGEVLERFIACNNYFSGSIPKSLKNCTGLHRLRLDRNQLTGNISEDFGIYPHLNYVDLSYNNFYGELSL
KWGDYRNITSLKISNNNVSGEIPTELGKATQLQLIDLSSNHLEGTIPKELGGLKLLYNLTLSNNHLSGAIPSDIKMLSSLKILDLASNNLSGSIPKQLGE
CSNLLLLNLSNNKFTNSIPQEMGFLRSLQDLDLSCNFLAQEIPWQLGQLQMLETLNVSHNMLSGLIPRTFKDLLSLTVVDISYNELHGPIPDTKAFHNAS
FEALRDNMGICGNASGLKPCNLPKSSRTVKRKSNKLVILIVLPLLGSLLLVLVVIGALFILRQRARKRKAEPGNIEQDRNLFTILGHDGKLLYENIIAAT
EEFNSNYCIGEGGYGTVYKAVMPAEQVVAVKKLHRSQTDKLSDFKAFETEVCVLANIRHRNIVKLYGFCSHAKHSFLVYEFIERGSLRKIITSEEQAIEL
DWMKRLNVVKGMAGALSYLHHSCSPPIIHRDITSNNVLLDLEYEAHVSDFGTARLLMPDSSNWTSFAGTFGYTAPELAYTMKVTEKCDVYSFGVVTMEVM
MGRHPGDLISTISSQASSSSSSKPPISQQTLLKDVLDQRISLPKKGAVEGVVHIMKIALACLHPNPQSRPTMGRISSELVTQWPSLPKEFYTISLEDLFL
HTVSVVD
|
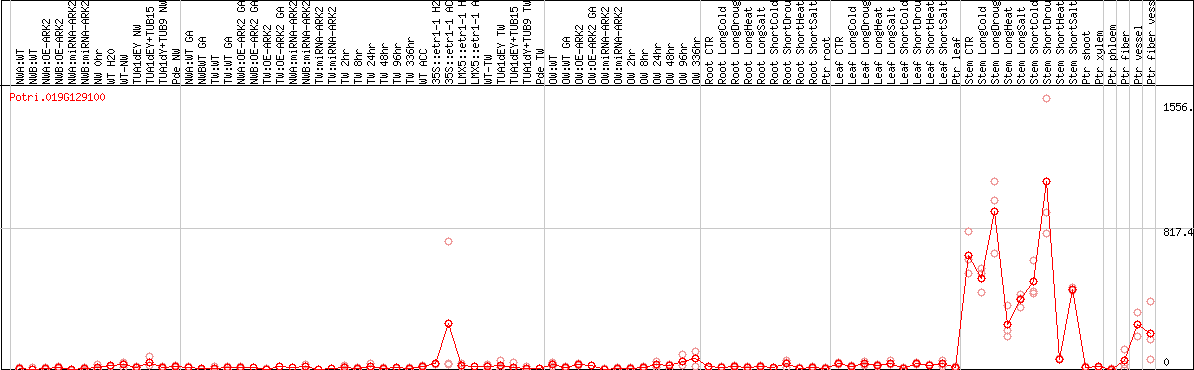
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G129100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
