External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G78340 172 / 1e-55
ATGSTU22
glutathione S-transferase TAU 22 (.1)
AT1G17170 157 / 7e-50
ATGSTU24
Arabidopsis thaliana Glutathione S-transferase \(class tau\) 24, glutathione S-transferase TAU 24 (.1)
AT1G17180 156 / 3e-49
ATGSTU25
glutathione S-transferase TAU 25 (.1)
AT1G78320 155 / 3e-49
ATGSTU23
glutathione S-transferase TAU 23 (.1)
AT1G78370 155 / 5e-49
ATGSTU20
glutathione S-transferase TAU 20 (.1)
AT1G78380 155 / 8e-49
GST8, ATGSTU19
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
AT1G17190 152 / 9e-48
ATGSTU26
glutathione S-transferase tau 26 (.1)
AT1G78360 145 / 3e-45
ATGSTU21
glutathione S-transferase TAU 21 (.1)
AT1G53680 135 / 3e-41
ATGSTU28
glutathione S-transferase TAU 28 (.1)
AT3G43800 133 / 3e-40
ATGSTU27
glutathione S-transferase tau 27 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.019G130433
211 / 6e-71
AT1G17180 292 / 3e-101
glutathione S-transferase TAU 25 (.1)
Potri.001G437400
170 / 6e-55
AT1G78380 316 / 2e-110
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
Potri.011G140800
169 / 1e-54
AT1G78380 318 / 1e-111
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
Potri.001G436800
167 / 1e-53
AT1G17180 308 / 2e-107
glutathione S-transferase TAU 25 (.1)
Potri.001G436600
166 / 2e-53
AT1G78380 318 / 2e-111
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
Potri.001G437200
165 / 6e-53
AT1G78380 308 / 2e-107
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
Potri.011G140700
164 / 9e-53
AT1G78380 287 / 3e-99
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
Potri.001G437500
163 / 1e-52
AT1G17180 232 / 2e-78
glutathione S-transferase TAU 25 (.1)
Potri.005G037801
161 / 2e-52
AT1G78340 197 / 7e-65
glutathione S-transferase TAU 22 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10042469
154 / 1e-48
AT1G78380 284 / 5e-98
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
Lus10042468
154 / 1e-48
AT1G78380 306 / 6e-107
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
Lus10026199
160 / 3e-47
AT1G17160 496 / 8e-173
pfkB-like carbohydrate kinase family protein (.1.2)
Lus10026198
150 / 7e-47
AT1G78380 295 / 3e-102
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
Lus10042470
145 / 4e-45
AT1G78380 291 / 1e-100
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
Lus10005592
142 / 5e-44
AT1G17180 271 / 9e-93
glutathione S-transferase TAU 25 (.1)
Lus10019480
134 / 2e-40
AT1G17180 231 / 6e-77
glutathione S-transferase TAU 25 (.1)
Lus10021102
125 / 4e-37
AT3G09270 245 / 2e-82
glutathione S-transferase TAU 8 (.1)
Lus10030020
124 / 7e-37
AT1G17180 238 / 1e-79
glutathione S-transferase TAU 25 (.1)
Lus10021103
123 / 4e-36
AT3G09270 250 / 1e-84
glutathione S-transferase TAU 8 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0172
Thioredoxin
PF02798
GST_N
Glutathione S-transferase, N-terminal domain
Representative CDS sequence
>Potri.019G130566.1 pacid=42773601 polypeptide=Potri.019G130566.1.p locus=Potri.019G130566 ID=Potri.019G130566.1.v4.1 annot-version=v4.1
ATGGAGGACAGAGTGACTCTATTGGATTTCTGGCCAAGTTCATGGGCAATGAGAGTGAAGGTTGCCTTGGCAGAGAAGGGGATAGAATACGAGTCCAGGG
AACAGAACTTGATAGACAAGAGCTCTCTGCTTCTTGAAATGAACCCGGTTCATAAGATGATTCCTGTTCTTATTCACAATGGGAAGCCCATTTGTGAGTC
ACATAATATTGTCCAATATATTGATGAGGTGTGGAAGGACAAGTCTCCATTGCTACCCTCGGATCCTTACCAAAGATCCCAAGCCAGATTCTGGGCTGAT
TACATTGACAAGAAGGCGAGTATTTCAGTACTTGTTTAA
AA sequence
>Potri.019G130566.1 pacid=42773601 polypeptide=Potri.019G130566.1.p locus=Potri.019G130566 ID=Potri.019G130566.1.v4.1 annot-version=v4.1
MEDRVTLLDFWPSSWAMRVKVALAEKGIEYESREQNLIDKSSLLLEMNPVHKMIPVLIHNGKPICESHNIVQYIDEVWKDKSPLLPSDPYQRSQARFWAD
YIDKKASISVLV
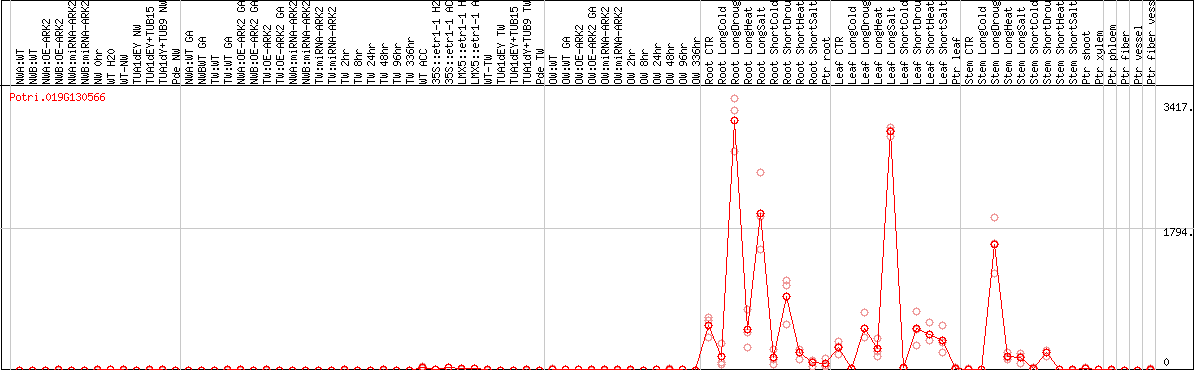
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G130566 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.