External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G10360 281 / 2e-95
TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.1), TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.2)
AT4G19645 212 / 3e-68
TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.1), TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.2)
AT1G31300 212 / 3e-68
TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.1), TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G056600
217 / 4e-70
AT4G19645 352 / 4e-123
TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.1), TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.2)
Potri.005G206000
215 / 2e-69
AT1G31300 348 / 2e-121
TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.1), TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.2)
Potri.015G113200
209 / 4e-67
AT1G31300 406 / 3e-144
TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.1), TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.2)
Potri.012G118100
204 / 7e-65
AT1G31300 349 / 7e-122
TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.1), TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10034433
305 / 7e-105
AT4G10360 321 / 3e-111
TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.1), TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.2)
Lus10019130
305 / 7e-105
AT4G10360 315 / 7e-109
TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.1), TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.2)
Lus10038335
214 / 5e-69
AT1G31300 375 / 3e-132
TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.1), TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.2)
Lus10036199
213 / 2e-68
AT4G19645 379 / 6e-134
TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.1), TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.2)
Lus10018287
209 / 1e-66
AT1G31300 375 / 5e-132
TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.1), TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.2)
Lus10040622
174 / 6e-53
AT1G31300 336 / 4e-116
TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.1), TRAM, LAG1 and CLN8 (TLC) lipid-sensing domain containing protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF03798
TRAM_LAG1_CLN8
TLC domain
Representative CDS sequence
>Potri.019G133500.1 pacid=42774491 polypeptide=Potri.019G133500.1.p locus=Potri.019G133500 ID=Potri.019G133500.1.v4.1 annot-version=v4.1
ATGGCGGCGGCCTTGTCTTTTCCTTGTGCACTTGACACTGCAACCAAACAATTTCATCTGCTGTCTTCCGTCTTGTCTGGCATCATTGCCTGCGGAATTG
TTTACAAATTAACAGCTTTTGTTAGCCGTTTGTACTTCAAAGGTTATGGGAAACTCACTGATGCGCAGAAAGTTGAGTGGAATAACCGAGGGTTCTCGAC
ATTTCATGCTCTCTTTGTTGCATCAGCTTCTCTATACCTCCTGCTGTTATCGGGTCTTTTTTATGAGGATTCTAGAGATGAATTAGTTGTTAATAGAACC
TCTACTTTATCAAACTCCACATTGGGGATATCCATTGGCTATTTTCTGTCAGATCTGGCAATGATTCTATTTCATTTTCCAGCATTAGGTGGTATGGAGT
ATCTTTTGCACCATGGACTATCCATGTTCTCCATAATCCTCGCTCTCCTAAGTGGCCAAGCACAGATTTACATACTAATGGTTCTCTTTTCTGAGATCAC
AACTCCCTTTGTAAATTTAAGATGGTATTTGGACGTTGCTGGTCAGAAGAGCTCTAAATTGTACATTTGGAATGGAGTTTTATTGTTCATGGGGTGGCTG
GTTGCAAGGATTCTTTTGTTCATCTTCTTCTTCAGCCACATGTTTATCCATTTTGATCAGGTCAAGCAAATATTTCCATTGGGCTTCTATAGCATTCTTG
TGATACCTGGGACATTGGCAGTGATGAATGTGTTATGGTTTTGGAAGATAGTGAAAGGTTTGATGAAAACCTTGTCTAAAGCTAGACATGGTCAGTGA
AA sequence
>Potri.019G133500.1 pacid=42774491 polypeptide=Potri.019G133500.1.p locus=Potri.019G133500 ID=Potri.019G133500.1.v4.1 annot-version=v4.1
MAAALSFPCALDTATKQFHLLSSVLSGIIACGIVYKLTAFVSRLYFKGYGKLTDAQKVEWNNRGFSTFHALFVASASLYLLLLSGLFYEDSRDELVVNRT
STLSNSTLGISIGYFLSDLAMILFHFPALGGMEYLLHHGLSMFSIILALLSGQAQIYILMVLFSEITTPFVNLRWYLDVAGQKSSKLYIWNGVLLFMGWL
VARILLFIFFFSHMFIHFDQVKQIFPLGFYSILVIPGTLAVMNVLWFWKIVKGLMKTLSKARHGQ
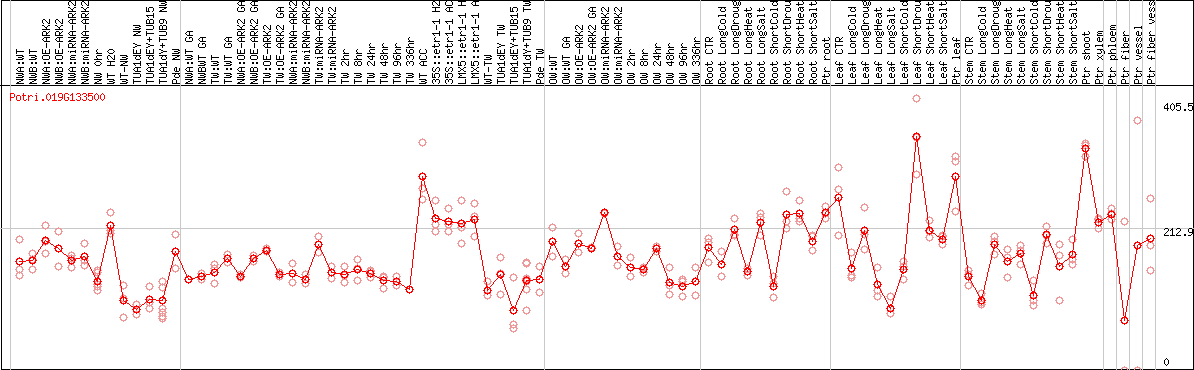
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G133500 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.