External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G49330 142 / 4e-43
MYB
PFG3, ATMYB111
PRODUCTION OF FLAVONOL GLYCOSIDES 3, ARABIDOPSIS MYB DOMAIN PROTEIN 111, myb domain protein 111 (.1)
AT2G16720 140 / 7e-43
MYB
AtY49, AtMYB7
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 7, myb domain protein 7 (.1)
AT1G22640 138 / 2e-42
MYB
AtMYB3
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 3, myb domain protein 3 (.1)
AT2G47460 140 / 4e-42
MYB
PFG1, ATMYB12
PRODUCTION OF FLAVONOL GLYCOSIDES 1, myb domain protein 12 (.1)
AT4G21440 138 / 2e-41
MYB
ATMYB102, ATM4
A. THALIANA MYB 4, MYB-like 102 (.1)
AT3G62610 137 / 4e-41
MYB
PFG2, ATMYB11
PRODUCTION OF FLAVONOL GLYCOSIDES 2, myb domain protein 11 (.1)
AT3G23250 136 / 4e-41
MYB
ATMYB15, ATY19
myb domain protein 15 (.1.2)
AT4G09460 134 / 4e-41
MYB
ATMYB6, ATMYB8
myb domain protein 6 (.1)
AT3G02940 135 / 1e-40
MYB
ATMYB107
myb domain protein 107 (.1)
AT4G28110 135 / 1e-40
MYB
ATMYB41
myb domain protein 41 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10033438
187 / 7e-61
AT1G22640 194 / 2e-61
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 3, myb domain protein 3 (.1)
Lus10000470
180 / 2e-58
AT2G16720 196 / 2e-61
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 7, myb domain protein 7 (.1)
Lus10033889
141 / 6e-43
AT3G23250 239 / 1e-78
myb domain protein 15 (.1.2)
Lus10030378
137 / 3e-41
AT3G61250 341 / 8e-118
LATE MERISTEM IDENTITY2, myb domain protein 17 (.1)
Lus10009448
136 / 3e-41
AT4G09460 290 / 2e-99
myb domain protein 6 (.1)
Lus10036453
135 / 8e-41
AT5G49330 237 / 1e-76
PRODUCTION OF FLAVONOL GLYCOSIDES 3, ARABIDOPSIS MYB DOMAIN PROTEIN 111, myb domain protein 111 (.1)
Lus10015713
137 / 9e-41
AT5G60890 162 / 5e-47
ALTERED TRYPTOPHAN REGULATION 1, myb domain protein 34 (.1)
Lus10003557
135 / 1e-40
AT3G23250 239 / 4e-78
myb domain protein 15 (.1.2)
Lus10001548
134 / 1e-40
AT4G09460 291 / 2e-99
myb domain protein 6 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G221800
162 / 2e-51
AT4G38620 197 / 5e-62
myb domain protein 4 (.1)
Potri.018G049401
154 / 2e-48
AT2G16720 190 / 4e-59
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 7, myb domain protein 7 (.1)
Potri.018G049600
154 / 3e-48
AT2G16720 192 / 4e-60
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 7, myb domain protein 7 (.1)
Potri.014G100800
152 / 7e-48
AT2G16720 207 / 3e-66
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 7, myb domain protein 7 (.1)
Potri.001G086700
150 / 8e-47
AT5G49330 203 / 6e-64
PRODUCTION OF FLAVONOL GLYCOSIDES 3, ARABIDOPSIS MYB DOMAIN PROTEIN 111, myb domain protein 111 (.1)
Potri.003G144200
150 / 1e-46
AT5G49330 205 / 2e-64
PRODUCTION OF FLAVONOL GLYCOSIDES 3, ARABIDOPSIS MYB DOMAIN PROTEIN 111, myb domain protein 111 (.1)
Potri.002G173900
150 / 2e-46
AT3G13540 207 / 4e-66
myb domain protein 5 (.1)
Potri.018G049200
153 / 4e-46
AT2G16720 189 / 8e-57
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 7, myb domain protein 7 (.1)
Potri.003G144300
147 / 2e-45
AT1G22640 197 / 1e-62
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 3, myb domain protein 3 (.1)
Potri.006G275900
143 / 1e-43
AT1G22640 186 / 1e-57
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 3, myb domain protein 3 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0123
HTH
PF00249
Myb_DNA-binding
Myb-like DNA-binding domain
Representative CDS sequence
>Lus10000008 pacid=23170262 polypeptide=Lus10000008 locus=Lus10000008.g ID=Lus10000008.BGIv1.0 annot-version=v1.0
ATGGGGAGGAGCCCTTGTTGTTCAAAGGAAGGAATGAACAGAGGCGCATGGACAGCTGAGGAGGACCAGCTCCTGACCCGCTACATCCAATCCCACGGAA
TTGGAAAATGGCGTAACCTACCGAAGAAAGCTGGCCTGAACCGGTGCGGCAAGAGCTGCCGCCTCCGGTGGCTCAATTACTTGCGCCCTGACATCAAGCG
CGGCAACATTTCCCCCGACGAGGAAGACCTTATCATCCGCCTCCACAAACTCTTAGGCAACCGGTAG
AA sequence
>Lus10000008 pacid=23170262 polypeptide=Lus10000008 locus=Lus10000008.g ID=Lus10000008.BGIv1.0 annot-version=v1.0
MGRSPCCSKEGMNRGAWTAEEDQLLTRYIQSHGIGKWRNLPKKAGLNRCGKSCRLRWLNYLRPDIKRGNISPDEEDLIIRLHKLLGNR
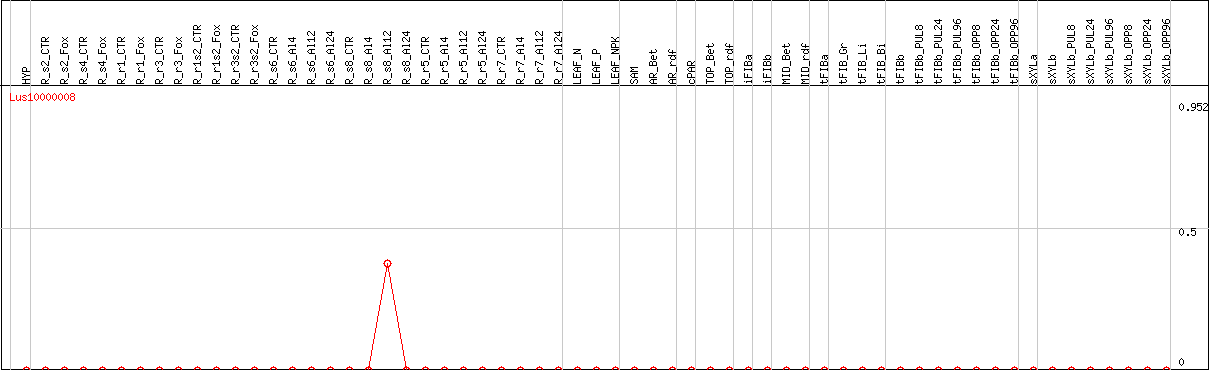
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10000008 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.