Lus10000423 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10000423 pacid=23142582 polypeptide=Lus10000423 locus=Lus10000423.g ID=Lus10000423.BGIv1.0 annot-version=v1.0
ATGGTGCTTCGGTGGTCTGGACACGATGGGTCACAATTAGTGTCACTTAGCGAAGCTGTTACTGCAGTCCGAGTCAGGGTGGAATGTGGACTTCAACCCG
AAGCTTTCATGTATGAGAGAAATTTATGGACCAAAGTTAAGGAGAAGAACTCAAAGGGAGGACAGCCAAAGGACGACTTGAGTGGATGGGGGGAATGGTT
GGAGACTTTGGTGACAGAAATATGCTGTCTTTGTATCAAAAGAAACTTGGTGGATCGGACGATCGAATTGCCCTGGGATTCTGCTGAGGAGAAGTATCTT
TCTAAGTGTCTTCTTGACTCTGCGGTTCACGATCCTTCCTCGACGACTGGAAGTCTTCTGGCAGTATACTATTTCAGGGGAAATTCCAAACAGCTGCTAT
CGCCTATCGAACAGAGTCTACACAAGGACGAATCGTATGGTTTTGATGATCCCATGGACCTAGGATGGAGAATGCCCATAGAAGACCTAACCAAGTTGAA
GAAGTTAACATTGTCTGTCCTTCTACTTTTACGAGAGATTGTGGAACTCGGCACACTGAAGTCACTAAAAAAGTTAGAAGTGAGGGGTTGCATTTCGTTG
GAAAGAATGCCCGTCCAGGGTCTAGTGTCGGATCTACGGGAAAGAATGCCCGTCATCGATAGTATGGACTACTTAGATGTGTCATCTATAAAGTTGGACT
TGAGAGGATGCACAAAGTTCACAAACCTGGATTCTGATCTTTCTCCCCTCGCCACCGAAGGAGCAGATCTAAGAGGAGTTGAGTTAATCATCAACTGGCC
AGACGAACCAGGAACAGATATTCCTCCCGCCGCTTTCCTTTGGAGGGACATCATGTTGTCAGAGGATGCAGAGGACTCTAACTCTGAAATACTCTCCATA
TTCTCCAAACAGCATAGCTCACAAACAGCAAAATCCGACCGACAAGATCACCAGGAACCAACGATTCAGTCGGAGGGTGCAGTAGAAATGTATTTCTTAG
TAGGAGCTGCGATAGTTGCTCTGTTTATTGCTCTCTTTAAGCTTTTTCTGGGAAGGAGGAGGAAGACGGTAAATTCTGATGATTCGAATAATGATACTCT
TCAACAATCGGATTCCGCTTCTACTTTTGGGGATTCTGGTTCGGTGCCCCTTCCCATCGGGGAGTACGAGGTGTTTTTGAACTTTAGAGGTCCGGATACA
CGCGATAACATTACGAATATTCTTTACCGTTTTCTCGTTCTCTCGAAAATCCGCACGTTCAGAGACGATGATGAACTCCGTAAGGGAGAAGGAATATGGC
CCAGCCTTGTCAAAGCCATCGGTCAGTCGAAGATTTCAGTTCCTATCTTCTCGCCTCGGTATGCCGAAAGTAAGTGGTGTCTGAAAGAACTGGCTGCGGT
TGTAGAGCATCAAAAGCGAGAGAAAGGTCATGTCATACTTCCCATTTTTTATAAGATTGATCCTAGAGACGTACGCCATCAAACTGGACCTTACAAAACG
GCTTTTCAACAACATAAACGGTCGAATAAATTTGATGAGGATACTATACAAAGTTGGAAAGTAGCTTTGAATGAGGTTGGATCCTTGAAAGGGTGGCACA
TCAAAACCGAAAAAGAGGCTGTAGATGTTGCCGATGTTATTTCTGGGGTTGTATGGTCACATCTAATCAAGAATAACAGTACATTAGAGATCGATGAGTT
GGTTGGAATCGATGATCATATGAAACAAGTAGTAGATATGCTAGATCTAGGCGCTAAAGGTGTGAATCTAATTGGTATTCATGGTATGGGTGGTATTGGG
AAAACCACTCTTGCAACAGCGGTCTATAACAAAGTATCTACCTGCTTTGATCGCTATAGTTTTGTGAAAGATATACGAGAAACACAAAAACAAACAAAAG
ATGTTCTCGTCTTACAAAAGAAGTTACTTTCTGATATCTTAAGTATGGAATCAGTTGGACCAAAAGCAAAGGAAATCATACGGGAGAGGGTTACTCAGTT
CAAGATTCTTGTGGTTCTCGATGATGTAGACGAAAAGTTCAACTTCGAAGAAGTCTTGGGAAATTCGAAAAGTTTTGCTCCAGGAAGTTGTTTTATCATC
ACGTCGAGAGATATAAAGGTCTTGGAAAGGTTAACCGGAGATAAAAGCAAGTTGTACGAAGTTAAAAAAATGGATGATAGTCATTCGCTTCAATTATTTT
ACAAGCATGCGTTCAGAGAGGATCCCGTGTCGTCAGATTTAGATGATTTATCAAAAAGCATTGTATCTAAAGCAGGAGGACTTCCATTGGCGCTTAAGGT
TATAGGGTCTCTTCTTTGGCGAGAAGAAAAGGTCTTTTGGAAAGCCAAATTAGAGCAGTTACAAGAAATGCCAGATGAAGACGTAATGGACAGGTTAATG
ATTAGCTACACAGCTTTAACCGAGGAAGCACAACAAATATTTTTAGATATTGCTTGCTTTTACATTGGAGAGAATAAAGAAAAGGCGTCTTACATGTGGA
GCGATTGCAAATATCATCCTGCCATTAACGTCAAGATTCTCATTCAAAGATCTATGCTCAAAATTGGGGATCGTAATGAGTTTCAAATGCACGATCAATT
TAGAGATATGGGGAGAGAAATTGTCCGTAAAGAAGATACTGACCATCCAGAAAGGAGAAGTAGAATATGGGTAGAGAATGAAGGCAAAGAAATTTTACTC
GACAACAAGGGAACAAATCAAGTCAGAGCACTCAGAGGCGGAAGAGCACTCAGAATCATATTTTACCCCAGGAGCAAATCTGTTGAAAGTGGATGTTTTA
GGAAGATGACAGAATTAATGAAGTGTAAGGCGACCAACTTCAATATGAAAAACATGGTCATATTTGAAGCCAATACTTATGGAATTGACGATATCCCAAC
AGTAAAGGAGGCAAAGAAGCTCAAATTTCTCAAGCTTGGCAATTGTAGAAAGATGAGTAAACTTCCAGAATTGCCAGAGAGCCTAGAGATGCTACATCTT
AGTAAGTTCGACAATAAAAAGGAAGATATGGAGCTCGGGAATCTGCACAACTTAAAAGTAGTAAGTTTAAAATCTTGTACGCTGGGGAGGTTCAACGGCG
GCACATTTGGGATGATGAAGGAGCTTCGAGAGCTCAAGCTCACTCACATTAAATGTGATTTTGATAGTTTCAGAAGAGTGATCGCCAATGTCGGACAACT
CTCATCTCTCGAAATCTTGGAGTTCAACAGTCCCAAGCTGAAGGATGTTTTGGAAGGGATTAAGCTACCGAAAAGCTTAAAGGTGCTAGAAACTTCCTCT
TGTTTCGATAATCTTCCGGAGCTCTTGGATTTAGAGGAATTCTCTATCATCAAGAATGATGCGAGGACAAGTATGGTTGAAGGTCAGAACACTGTGCTAC
CATCATCCCTCACTACCGTAGTCATTTTCCATCTACAATCTCATACATTCCTGAATCTGACCAATCTGAGCAACTTGACTCAATTAAAACTTTGGGGTAG
GCCTTATCTTCAAGAGATCCAAGGACTTGGAGGGCTGAAATCATTACAGATTCTTTCGATTGGTAAAATGGAAAAGTTAGCTCACATTGAGGGCCTTGGG
ATTCTCATGTCTTCCTCCTACTGCAAGTTGACACATGTGGAGATAATCGAATGTCCTCTTCTTCGTGCATTGCAGACATTCGAACAACATGATGATGAAG
ATGATACTAATGATGCAGTACAAATCGAATCTTTAATCAAACTGGAGATAGTTAAATGTCCATTAATGGATTGTAGATCAATACCCAAATTGTCCGAGTT
TCCAAAACTCAATGTGTTGACAATTCGGGACATCGGTTCGAATGTCAATGATGAATCAACCAGCCAGCAGGATCAGCTACTGGAAGGACTTGACAATCTT
GGAGAGTTGGTTCAATTGCAAGTGCGCCATTTAGCAAAAGTACAAAGACTACCTTCTCTATCAAAGCTAAGCAAGTTGACGAAGTTAACATTGTATGACC
TTCCATGTTTGCGAGAGATTGTGGGACTCGGAAATTTGAAGTCTTTAAAAGAGTTAGAAGTGAGTGTTTGCATTTCGTTGGAAAGAATGCTCGTCGAGGA
TCTGACCAAGTTGAAGAAGCTAAGTTTGTCTGACCTTCCACGTTTGAGAGAGATTGTGGGACTCGGAGATTTAAAGTCGTTAAAAGAGTTAGAAGTGAGT
GATTGCATTTCGTTGGAAAGAATGCTCGTCGAGGATCTGACCAAGTTGACGGTGTTAATAGTGTCTGGCCTTCCATGTTTGCAAGAGATTGTGGGGCTCG
GAGCATTGAAGTCGTTACAAGGGTTAGCAGTGAGACGTTGCATTTCGTTGGAAAGAATGCCCGTCGAGGGACTCTCCTACTCTACTATGTTTTATATAGA
TTTGGATTTGACAGGATGTACAAAGTTCACAAACGCTCATTCTGATCTCTCCCCCCTCACCACCCAAAGGGCCTTCTATCCACGTCGGCTAGAAGTAAAC
ATCATCTGGCCAGAAGAACATTTGGGGCCTTTTCAGAGAGGGTCGAGGCCACCGAGTGAGGAATCTGACGATGATACGTGGGAGGAATCTGACGACAATA
TGGGTGAGGAATCTGACGACGATTTATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10000423 pacid=23142582 polypeptide=Lus10000423 locus=Lus10000423.g ID=Lus10000423.BGIv1.0 annot-version=v1.0
MVLRWSGHDGSQLVSLSEAVTAVRVRVECGLQPEAFMYERNLWTKVKEKNSKGGQPKDDLSGWGEWLETLVTEICCLCIKRNLVDRTIELPWDSAEEKYL
SKCLLDSAVHDPSSTTGSLLAVYYFRGNSKQLLSPIEQSLHKDESYGFDDPMDLGWRMPIEDLTKLKKLTLSVLLLLREIVELGTLKSLKKLEVRGCISL
ERMPVQGLVSDLRERMPVIDSMDYLDVSSIKLDLRGCTKFTNLDSDLSPLATEGADLRGVELIINWPDEPGTDIPPAAFLWRDIMLSEDAEDSNSEILSI
FSKQHSSQTAKSDRQDHQEPTIQSEGAVEMYFLVGAAIVALFIALFKLFLGRRRKTVNSDDSNNDTLQQSDSASTFGDSGSVPLPIGEYEVFLNFRGPDT
RDNITNILYRFLVLSKIRTFRDDDELRKGEGIWPSLVKAIGQSKISVPIFSPRYAESKWCLKELAAVVEHQKREKGHVILPIFYKIDPRDVRHQTGPYKT
AFQQHKRSNKFDEDTIQSWKVALNEVGSLKGWHIKTEKEAVDVADVISGVVWSHLIKNNSTLEIDELVGIDDHMKQVVDMLDLGAKGVNLIGIHGMGGIG
KTTLATAVYNKVSTCFDRYSFVKDIRETQKQTKDVLVLQKKLLSDILSMESVGPKAKEIIRERVTQFKILVVLDDVDEKFNFEEVLGNSKSFAPGSCFII
TSRDIKVLERLTGDKSKLYEVKKMDDSHSLQLFYKHAFREDPVSSDLDDLSKSIVSKAGGLPLALKVIGSLLWREEKVFWKAKLEQLQEMPDEDVMDRLM
ISYTALTEEAQQIFLDIACFYIGENKEKASYMWSDCKYHPAINVKILIQRSMLKIGDRNEFQMHDQFRDMGREIVRKEDTDHPERRSRIWVENEGKEILL
DNKGTNQVRALRGGRALRIIFYPRSKSVESGCFRKMTELMKCKATNFNMKNMVIFEANTYGIDDIPTVKEAKKLKFLKLGNCRKMSKLPELPESLEMLHL
SKFDNKKEDMELGNLHNLKVVSLKSCTLGRFNGGTFGMMKELRELKLTHIKCDFDSFRRVIANVGQLSSLEILEFNSPKLKDVLEGIKLPKSLKVLETSS
CFDNLPELLDLEEFSIIKNDARTSMVEGQNTVLPSSLTTVVIFHLQSHTFLNLTNLSNLTQLKLWGRPYLQEIQGLGGLKSLQILSIGKMEKLAHIEGLG
ILMSSSYCKLTHVEIIECPLLRALQTFEQHDDEDDTNDAVQIESLIKLEIVKCPLMDCRSIPKLSEFPKLNVLTIRDIGSNVNDESTSQQDQLLEGLDNL
GELVQLQVRHLAKVQRLPSLSKLSKLTKLTLYDLPCLREIVGLGNLKSLKELEVSVCISLERMLVEDLTKLKKLSLSDLPRLREIVGLGDLKSLKELEVS
DCISLERMLVEDLTKLTVLIVSGLPCLQEIVGLGALKSLQGLAVRRCISLERMPVEGLSYSTMFYIDLDLTGCTKFTNAHSDLSPLTTQRAFYPRRLEVN
IIWPEEHLGPFQRGSRPPSEESDDDTWEESDDNMGEESDDDL
|
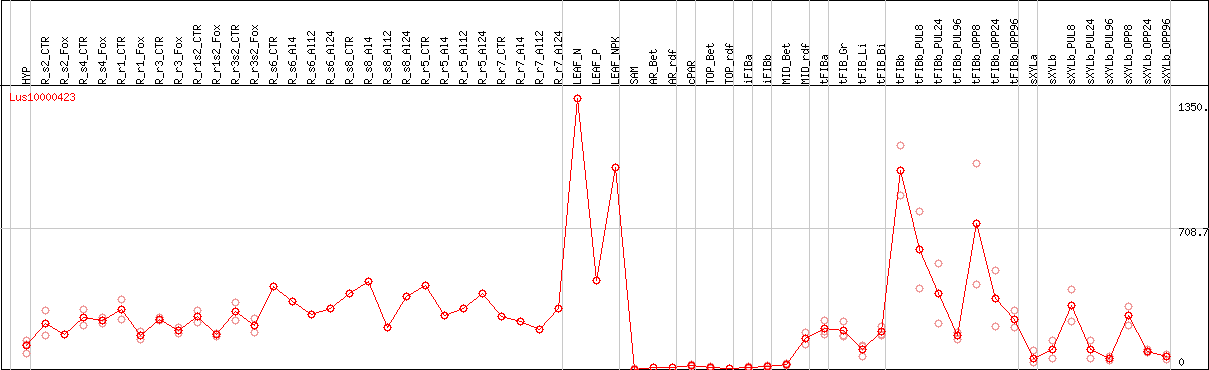
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10000423 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
