External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G13840 207 / 8e-65
GRAS
GRAS family transcription factor (.1)
AT3G49950 73 / 3e-15
GRAS
GRAS family transcription factor (.1)
AT5G17490 59 / 2e-10
GRAS
AtRGL3, RGL3
RGA-like protein 3 (.1)
AT3G03450 56 / 2e-09
GRAS
RGL2
RGA-like 2 (.1)
AT4G17230 54 / 1e-08
GRAS
SCL13
SCARECROW-like 13 (.1)
AT4G37650 50 / 3e-07
GRAS
SGR7, SHR
SHORT ROOT, SHOOT GRAVITROPISM 7, GRAS family transcription factor (.1)
AT5G66770 47 / 4e-06
GRAS
GRAS family transcription factor (.1)
AT2G01570 44 / 5e-05
GRAS
RGA1
REPRESSOR OF GA1-3 1, REPRESSOR OF GA, GRAS family transcription factor family protein (.1)
AT1G66350 44 / 5e-05
GRAS
RGL1
RGA-like 1 (.1)
AT1G14920 43 / 8e-05
GRAS
RGA2, GAI
RESTORATION ON GROWTH ON AMMONIA 2, GIBBERELLIC ACID INSENSITIVE, GRAS family transcription factor family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10022097
367 / 1e-130
AT3G13840 264 / 5e-86
GRAS family transcription factor (.1)
Lus10019281
61 / 5e-11
AT4G37650 509 / 2e-177
SHORT ROOT, SHOOT GRAVITROPISM 7, GRAS family transcription factor (.1)
Lus10018498
61 / 7e-11
AT3G49950 347 / 3e-116
GRAS family transcription factor (.1)
Lus10039709
61 / 8e-11
AT3G49950 350 / 3e-117
GRAS family transcription factor (.1)
Lus10032882
58 / 6e-10
AT1G55580 294 / 2e-95
SCARECROW-LIKE 18, Lateral Suppressor, GRAS family transcription factor (.1)
Lus10033583
56 / 3e-09
AT2G01570 681 / 0.0
REPRESSOR OF GA1-3 1, REPRESSOR OF GA, GRAS family transcription factor family protein (.1)
Lus10025200
56 / 5e-09
AT4G37650 484 / 8e-169
SHORT ROOT, SHOOT GRAVITROPISM 7, GRAS family transcription factor (.1)
Lus10017626
55 / 8e-09
AT2G01570 679 / 0.0
REPRESSOR OF GA1-3 1, REPRESSOR OF GA, GRAS family transcription factor family protein (.1)
Lus10012089
54 / 1e-08
AT4G37650 483 / 2e-167
SHORT ROOT, SHOOT GRAVITROPISM 7, GRAS family transcription factor (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF03514
GRAS
GRAS domain family
Representative CDS sequence
>Lus10000532 pacid=23140357 polypeptide=Lus10000532 locus=Lus10000532.g ID=Lus10000532.BGIv1.0 annot-version=v1.0
ATGAATATAAACCTGCAGATCAATCGCCTCGACAATCGATCCTTACAGAATCTGAACACCCAGGTAATCAACAAGGACAGGGAAGAAACTCTAGTTGTCT
GCGCCCAATTCAGGCTACACCACCTGAATCACAACAACCCGGATGAGAGGAGCGAGTTTTTAAGGGTTCTGAGGAGTCTGGAGCCCAAGGGAGTAATCCT
GAGCGAGAACAACATGGATTGCAGCTGCAACGGGTGCGTGGATTATTCGAGCGGGTTCACGAGGAGGGTGGAGTATCTGTGGAGGTTTCTGGATTCTACG
AGCTCAGCGTTCAAAGGGAGGGAGAGCGAAGAGAGAAGGATGATGGAAGGGGAAGCAGCAAAGGCGTTGACAAACAGTGGGGAGATGAACGAGAGGAAAG
AGCAGTGGTGCGAGAGGATGAGAGGGGTTGGGTTTTCAGGGGAAGGGTTTTGCGAGGACGCCATTGATGAAGCTCGAGCTCTGTTGAGGAAGCATGATAA
CAATTGGGAAATGAGGGTTGAAGAGAAAGATGGGTTCGTTGGGTTGTGGTGGAAAGGACAGCCCGTTTCGTTTTGCTCCTTGTGGAAATTGCAAGCAGAA
TGA
AA sequence
>Lus10000532 pacid=23140357 polypeptide=Lus10000532 locus=Lus10000532.g ID=Lus10000532.BGIv1.0 annot-version=v1.0
MNINLQINRLDNRSLQNLNTQVINKDREETLVVCAQFRLHHLNHNNPDERSEFLRVLRSLEPKGVILSENNMDCSCNGCVDYSSGFTRRVEYLWRFLDST
SSAFKGRESEERRMMEGEAAKALTNSGEMNERKEQWCERMRGVGFSGEGFCEDAIDEARALLRKHDNNWEMRVEEKDGFVGLWWKGQPVSFCSLWKLQAE
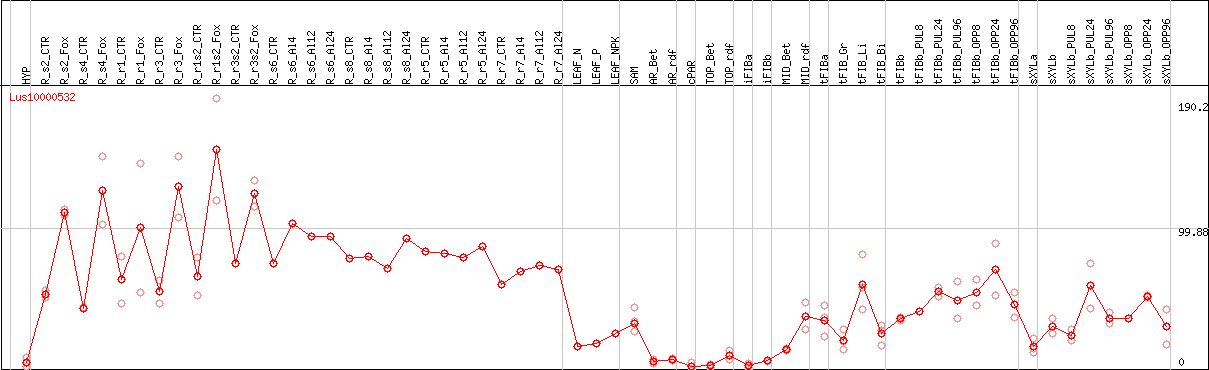
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10000532 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.