External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G47460 209 / 6e-65
MYB
PFG1, ATMYB12
PRODUCTION OF FLAVONOL GLYCOSIDES 1, myb domain protein 12 (.1)
AT3G62610 206 / 4e-64
MYB
PFG2, ATMYB11
PRODUCTION OF FLAVONOL GLYCOSIDES 2, myb domain protein 11 (.1)
AT5G49330 194 / 1e-59
MYB
PFG3, ATMYB111
PRODUCTION OF FLAVONOL GLYCOSIDES 3, ARABIDOPSIS MYB DOMAIN PROTEIN 111, myb domain protein 111 (.1)
AT4G28110 179 / 2e-54
MYB
ATMYB41
myb domain protein 41 (.1)
AT4G21440 180 / 5e-54
MYB
ATMYB102, ATM4
A. THALIANA MYB 4, MYB-like 102 (.1)
AT1G22640 176 / 9e-54
MYB
AtMYB3
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 3, myb domain protein 3 (.1)
AT1G34670 178 / 3e-53
MYB
ATMYB93
myb domain protein 93 (.1)
AT3G02940 177 / 3e-53
MYB
ATMYB107
myb domain protein 107 (.1)
AT5G16770 177 / 3e-53
MYB
ATMYB9
myb domain protein 9 (.1.2)
AT5G15310 176 / 1e-52
MYB
ATMYB16, ATMIXTA
myb domain protein 16 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10002435
486 / 2e-174
AT2G47460 206 / 9e-64
PRODUCTION OF FLAVONOL GLYCOSIDES 1, myb domain protein 12 (.1)
Lus10036336
208 / 9e-65
AT2G47460 225 / 9e-71
PRODUCTION OF FLAVONOL GLYCOSIDES 1, myb domain protein 12 (.1)
Lus10010273
207 / 2e-64
AT2G47460 221 / 3e-69
PRODUCTION OF FLAVONOL GLYCOSIDES 1, myb domain protein 12 (.1)
Lus10016855
199 / 9e-62
AT5G49330 242 / 3e-78
PRODUCTION OF FLAVONOL GLYCOSIDES 3, ARABIDOPSIS MYB DOMAIN PROTEIN 111, myb domain protein 111 (.1)
Lus10036453
192 / 5e-59
AT5G49330 237 / 1e-76
PRODUCTION OF FLAVONOL GLYCOSIDES 3, ARABIDOPSIS MYB DOMAIN PROTEIN 111, myb domain protein 111 (.1)
Lus10027459
182 / 2e-55
AT3G61250 229 / 3e-74
LATE MERISTEM IDENTITY2, myb domain protein 17 (.1)
Lus10039214
179 / 3e-54
AT3G61250 226 / 7e-73
LATE MERISTEM IDENTITY2, myb domain protein 17 (.1)
Lus10041142
179 / 9e-54
AT1G34670 315 / 2e-106
myb domain protein 93 (.1)
Lus10036472
178 / 1e-53
AT5G10280 307 / 1e-103
myb domain protein 92 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G141000
211 / 5e-66
AT5G49330 248 / 7e-80
PRODUCTION OF FLAVONOL GLYCOSIDES 3, ARABIDOPSIS MYB DOMAIN PROTEIN 111, myb domain protein 111 (.1)
Potri.002G198100
209 / 8e-65
AT5G49330 225 / 2e-70
PRODUCTION OF FLAVONOL GLYCOSIDES 3, ARABIDOPSIS MYB DOMAIN PROTEIN 111, myb domain protein 111 (.1)
Potri.014G122700
199 / 1e-60
AT5G49330 214 / 4e-66
PRODUCTION OF FLAVONOL GLYCOSIDES 3, ARABIDOPSIS MYB DOMAIN PROTEIN 111, myb domain protein 111 (.1)
Potri.004G033100
182 / 2e-54
AT4G21440 293 / 4e-97
A. THALIANA MYB 4, MYB-like 102 (.1)
Potri.007G093900
179 / 1e-53
AT1G34670 289 / 6e-96
myb domain protein 93 (.1)
Potri.019G118200
179 / 1e-53
AT4G05100 278 / 7e-92
myb domain protein 74 (.1)
Potri.005G164900
178 / 2e-53
AT1G34670 309 / 7e-104
myb domain protein 93 (.1)
Potri.015G143500
177 / 2e-53
AT3G61250 278 / 2e-93
LATE MERISTEM IDENTITY2, myb domain protein 17 (.1)
Potri.019G018200
174 / 2e-53
AT2G31180 223 / 1e-73
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 14, myb domain protein 14 (.1)
Potri.013G148600
177 / 7e-53
AT4G21440 269 / 4e-88
A. THALIANA MYB 4, MYB-like 102 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0123
HTH
PF00249
Myb_DNA-binding
Myb-like DNA-binding domain
Representative CDS sequence
>Lus10001458 pacid=23161042 polypeptide=Lus10001458 locus=Lus10001458.g ID=Lus10001458.BGIv1.0 annot-version=v1.0
ATGGGGAGGAAGCCATGCTGCGACAAGGTGGGTTTGAAGAAGGGCAGATGGTCTGCAGAGGAAGACCAGCTTCTTGCCGATTACATCGTTGCCAATGGTG
AAGGCTCCTGGAAATCATTGCCCCAAAATGCCGGACTGAAGAGGTGCGGGAAGAGTTGCAGACTAAGGTGGGTTAATTACCTGAAATCTGATATCAAGAG
GGGCAACATCTCCACCGCCGAGGCCGAAACCATACTCAACTTGCATACTTGTCTGGGCAACAGATGGTCTACAATAGCTAGTCAATTGTCGGGGAGAACA
GACAACGAGATCAAAAACTACTGGAATTCACATTTAAGCAGAAATACTCACAGCTTCATACGCCCAACTGCGACCCAACCAAGTTTGGCGACATTAGTAG
CAGACGTGTCCAAGGTTGGGACCACGGTGGTAGCCAATACAAAACGAGGAAGAAAAACAAATAACTCATCATCATCAAAGCGTACCAAGAACGGCAAAAC
CAACATATCAAAGGACACCGGTAAAAATGTTGTCAGCGAGCCATCAGTGGTAGAGAAAGCCGTCATTACAGTCAGCGAACCGGAGACAGGTCTAGCAACG
ACCACAAAACATCCGTCACCAATAAGCTATATTAGCGAAGAAACTACTTGTAGCAGAGTGATGAAAGAGCAGAAATACAATGCCTCATTAAGCTGGGCAA
CTGAAATCCCAGCTACTCCATGGGAAAAAGTCCTTCCTCTTGATGAGCAATGGGACATGGACAAAGATTGGGATGATGTCGTGAGCTACTGTACTAAGAC
TTCGACCATGGATAGTACTGTCGACCCATTGATTGGTTGGTCGTGGGAATTGTACGCTGGTGATGGTGGCCTAGAAGCAGGAAGGCATAATGGAGTGTTG
GAGTTTGATGACCATGACAATGATGACGACGACAAGCAAACTGCCTTCGTAAATTGGCTTCTTTCCTGA
AA sequence
>Lus10001458 pacid=23161042 polypeptide=Lus10001458 locus=Lus10001458.g ID=Lus10001458.BGIv1.0 annot-version=v1.0
MGRKPCCDKVGLKKGRWSAEEDQLLADYIVANGEGSWKSLPQNAGLKRCGKSCRLRWVNYLKSDIKRGNISTAEAETILNLHTCLGNRWSTIASQLSGRT
DNEIKNYWNSHLSRNTHSFIRPTATQPSLATLVADVSKVGTTVVANTKRGRKTNNSSSSKRTKNGKTNISKDTGKNVVSEPSVVEKAVITVSEPETGLAT
TTKHPSPISYISEETTCSRVMKEQKYNASLSWATEIPATPWEKVLPLDEQWDMDKDWDDVVSYCTKTSTMDSTVDPLIGWSWELYAGDGGLEAGRHNGVL
EFDDHDNDDDDKQTAFVNWLLS
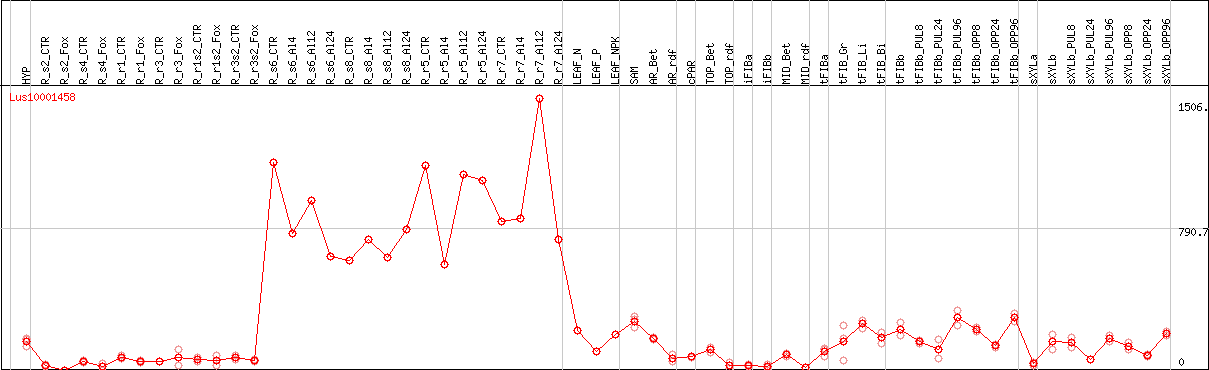
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10001458 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.