External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G25390 148 / 1e-45
AP2_ERF
SHN3, SHN2
shine3, Integrase-type DNA-binding superfamily protein (.1.2)
AT5G11190 147 / 5e-45
AP2_ERF
SHN2, SHN3
shine2, Integrase-type DNA-binding superfamily protein (.1)
AT1G15360 143 / 2e-43
AP2_ERF
WIN1, SHN1
WAX INDUCER 1, SHINE 1, Integrase-type DNA-binding superfamily protein (.1)
AT5G25190 81 / 2e-19
AP2_ERF
ESE3
ethylene and salt inducible 3, Integrase-type DNA-binding superfamily protein (.1)
AT5G65130 75 / 2e-16
AP2_ERF
Integrase-type DNA-binding superfamily protein (.1)
AT1G71450 72 / 7e-16
AP2_ERF
Integrase-type DNA-binding superfamily protein (.1)
AT4G28140 69 / 6e-14
AP2_ERF
Integrase-type DNA-binding superfamily protein (.1)
AT4G11140 68 / 7e-14
AP2_ERF
CRF1
cytokinin response factor 1 (.1)
AT1G28370 66 / 7e-14
AP2_ERF
AtERF11
ERF domain protein 11 (.1)
AT3G20310 67 / 1e-13
AP2_ERF
ATERF7, ATERF-7
ethylene response factor 7 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10002015
227 / 3e-76
AT1G15360 183 / 4e-59
WAX INDUCER 1, SHINE 1, Integrase-type DNA-binding superfamily protein (.1)
Lus10005716
154 / 2e-47
AT1G15360 236 / 3e-79
WAX INDUCER 1, SHINE 1, Integrase-type DNA-binding superfamily protein (.1)
Lus10030097
154 / 2e-47
AT1G15360 234 / 1e-78
WAX INDUCER 1, SHINE 1, Integrase-type DNA-binding superfamily protein (.1)
Lus10009480
138 / 1e-41
AT1G15360 207 / 1e-68
WAX INDUCER 1, SHINE 1, Integrase-type DNA-binding superfamily protein (.1)
Lus10019414
139 / 2e-41
AT1G15360 207 / 1e-67
WAX INDUCER 1, SHINE 1, Integrase-type DNA-binding superfamily protein (.1)
Lus10043271
94 / 6e-24
AT1G15360 160 / 2e-49
WAX INDUCER 1, SHINE 1, Integrase-type DNA-binding superfamily protein (.1)
Lus10041023
86 / 3e-21
AT5G25190 205 / 7e-68
ethylene and salt inducible 3, Integrase-type DNA-binding superfamily protein (.1)
Lus10005343
79 / 8e-19
AT5G25190 174 / 3e-56
ethylene and salt inducible 3, Integrase-type DNA-binding superfamily protein (.1)
Lus10035859
77 / 8e-18
AT5G25190 196 / 9e-65
ethylene and salt inducible 3, Integrase-type DNA-binding superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G028000
157 / 3e-49
AT5G11190 204 / 8e-68
shine2, Integrase-type DNA-binding superfamily protein (.1)
Potri.018G131400
155 / 4e-48
AT1G15360 194 / 6e-63
WAX INDUCER 1, SHINE 1, Integrase-type DNA-binding superfamily protein (.1)
Potri.006G253800
148 / 4e-45
AT5G11190 197 / 1e-64
shine2, Integrase-type DNA-binding superfamily protein (.1)
Potri.003G033000
143 / 9e-44
AT1G15360 181 / 3e-58
WAX INDUCER 1, SHINE 1, Integrase-type DNA-binding superfamily protein (.1)
Potri.006G069400
137 / 4e-41
AT1G15360 186 / 9e-60
WAX INDUCER 1, SHINE 1, Integrase-type DNA-binding superfamily protein (.1)
Potri.006G261200
83 / 5e-20
AT5G25190 169 / 9e-54
ethylene and salt inducible 3, Integrase-type DNA-binding superfamily protein (.1)
Potri.018G021900
83 / 7e-20
AT5G25190 169 / 1e-53
ethylene and salt inducible 3, Integrase-type DNA-binding superfamily protein (.1)
Potri.001G048200
81 / 3e-19
AT5G25190 162 / 5e-51
ethylene and salt inducible 3, Integrase-type DNA-binding superfamily protein (.1)
Potri.013G101100
70 / 4e-15
AT1G71450 184 / 1e-59
Integrase-type DNA-binding superfamily protein (.1)
Potri.003G054100
70 / 4e-15
AT1G71450 144 / 8e-44
Integrase-type DNA-binding superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0081
MBD-like
PF00847
AP2
AP2 domain
Representative CDS sequence
>Lus10002912 pacid=23148585 polypeptide=Lus10002912 locus=Lus10002912.g ID=Lus10002912.BGIv1.0 annot-version=v1.0
ATGGTACAATCAAAGAAATACAGAGGTGTCAGGCAGCGTCAGTGGGGTTCTTGGGTCTCCGAGATTCGCCACCCTTTACTGAAGAGGAGGGTGTGGCTGG
GGACGTTCGACACGGCCGAGGCCGCGGCAAGGGCGTACGACCGGGCCGCCGTCCACATGAGCGGCCGCAATGCGAAGACGAATTTCCCTGCCTCTTCTTC
CGATGATGATGATAATGCGTCACTTGGGGAACTTCTGAATGCGAAGCTCCGCAAGTGTTGTGACACATCGGCGTCTTCGATGACGTGTCTGAGGCTGGAC
AGTAGTGACAGCTCTCACATTGGTGTCTGGCAGAAGCGCCCGGCTGGTGGGCCGCGGTCCTCCAGCTGGGTCATGCGGGTCCGCTTGGGAGGCGGTACTG
GTGGTGGTTCTGATTCTGATCACCCCCCTCCGGCACCAGTGCTGGAGAGGGAGGAATCGTCTTCTTCTTTGTCGTCTTCGTCGGGTTCGGAGATGGTGGA
AGAGAAAGAGGAAGATAGAATTGCTATGCAGATGATTGAAGAGCTGCTTGAAGAATAG
AA sequence
>Lus10002912 pacid=23148585 polypeptide=Lus10002912 locus=Lus10002912.g ID=Lus10002912.BGIv1.0 annot-version=v1.0
MVQSKKYRGVRQRQWGSWVSEIRHPLLKRRVWLGTFDTAEAAARAYDRAAVHMSGRNAKTNFPASSSDDDDNASLGELLNAKLRKCCDTSASSMTCLRLD
SSDSSHIGVWQKRPAGGPRSSSWVMRVRLGGGTGGGSDSDHPPPAPVLEREESSSSLSSSSGSEMVEEKEEDRIAMQMIEELLEE
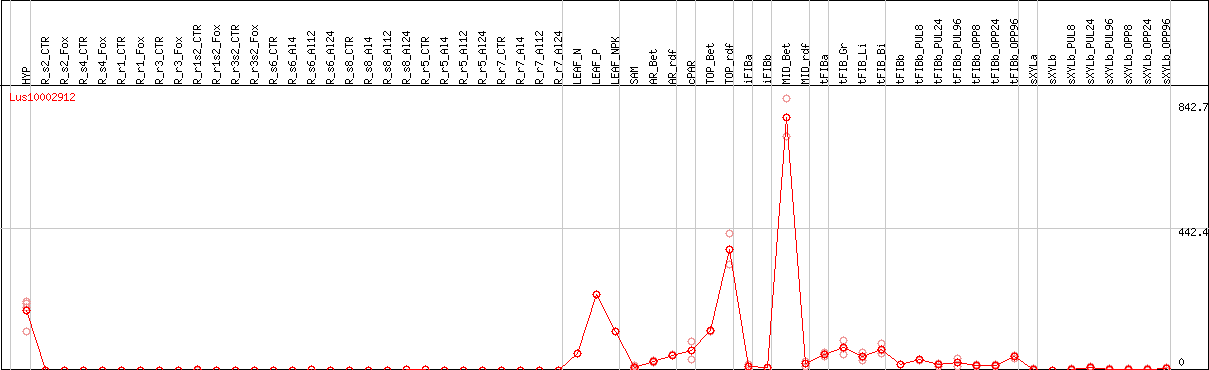
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10002912 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.