External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G24050 464 / 3e-165
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT1G64590 407 / 6e-143
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT5G50130 310 / 2e-104
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT2G37540 262 / 4e-86
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT5G02540 251 / 1e-81
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT4G23420 246 / 8e-80
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2), NAD(P)-binding Rossmann-fold superfamily protein (.3)
AT4G23430 244 / 3e-79
AtTic32-IVa
translocon at the inner envelope membrane of chloroplasts 32-IVa, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2), NAD(P)-binding Rossmann-fold superfamily protein (.3)
AT4G11410 243 / 1e-78
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT4G09750 95 / 6e-22
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT5G53100 93 / 4e-21
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10017308
578 / 0
AT4G24050 495 / 1e-177
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10042088
317 / 7e-107
AT5G50130 460 / 3e-163
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10018092
259 / 3e-84
AT5G50130 403 / 1e-140
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10024415
254 / 1e-82
AT5G02540 471 / 4e-168
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10023762
253 / 3e-82
AT2G37540 429 / 8e-152
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10035480
243 / 1e-78
AT4G11410 444 / 5e-158
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10028781
243 / 1e-78
AT4G11410 470 / 3e-168
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10035479
243 / 3e-78
AT4G11410 444 / 2e-157
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10035481
240 / 2e-77
AT4G23430 479 / 6e-172
translocon at the inner envelope membrane of chloroplasts 32-IVa, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2), NAD(P)-binding Rossmann-fold superfamily protein (.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G086900
465 / 2e-165
AT4G24050 441 / 2e-156
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.003G144100
428 / 7e-151
AT4G24050 397 / 9e-139
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.012G087800
315 / 1e-106
AT5G50130 426 / 4e-150
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.015G081102
315 / 1e-106
AT5G50130 454 / 2e-161
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.T124508
315 / 1e-106
AT5G50130 454 / 2e-161
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.006G083900
263 / 2e-86
AT5G02540 452 / 3e-161
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.003G128800
252 / 4e-82
AT4G23420 442 / 5e-157
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2), NAD(P)-binding Rossmann-fold superfamily protein (.3)
Potri.001G103000
242 / 2e-78
AT4G11410 446 / 4e-159
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.015G146600
237 / 3e-76
AT4G23420 444 / 6e-158
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2), NAD(P)-binding Rossmann-fold superfamily protein (.3)
Potri.003G128700
233 / 8e-75
AT4G11410 497 / 4e-179
NAD(P)-binding Rossmann-fold superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF00106
adh_short
short chain dehydrogenase
Representative CDS sequence
>Lus10003204 pacid=23155743 polypeptide=Lus10003204 locus=Lus10003204.g ID=Lus10003204.BGIv1.0 annot-version=v1.0
ATGCTGCAAACAGTCAAATACTTGATCGGCGCACGCGGAGCCAGCGGCTACGGCTCCAAATCCACGGCAGAGGAAGTCACGGAAAACTGCAGCGATCTCC
GCTCCGTCACCGCCATCATCACCGGAGCGACGTCGGGGATCGGTGCGGAGACCGCCAGAGTGCTGGCGAAGAGAGGCGCGACGGTGGTGATTCCGGTGAG
GAGCATGAAAGCGGCAGAGGATACGAAGGCGCGGATATTGTCAGAGTGTCCTGATGCTGCGATCATCGTTATGGCGCTCGATCTCAGCTCGCTCCGATCC
GTCAGAAGCTTCGTCTCCGAGTTCGAATCGCTCGATTTGCCCCTCAATCTCCTCATAAATAACGCCGGGAAATTCGCGCACCAGCATGCGATATCGGAGG
ATGGAATCGAGATGTCGTTCGCCACTAACTACCTAGGCCATTTTCTGCTGACGAATCTGTTACTGGAGAAGATGAAGGAAACAGCGGAGGAAACGGGAGT
GCAGGGGAGAATAGTGAACGTGTCATCGGCGATTCACGGGTGGTTCGCTGGCGATATGCTGCAGTACCTGGAAGAGATTTCTGGGCAGAAATGTCCTCAT
TACGATGCTACACGTGCATACGCTCTCTCCAAGCTTGCTAACGTCCTCCATACCAAACACCTTGCTCTGAAGCTCAAGGAAACGGATGCCAACGTAACAG
TCAACTGCGTACACCCCGGAATCGTCCGGACGAGGCTGACGAGAGAAAGAGAAGGGTTCGCAACAGACGCAGCCTTCTTCTTAGCATCAAAACTCTTAAA
AACAATCCCGCAGGCAGCCGCGACGACTCTCTATGTAGCGACCCATCCAAGGGTGGGGAATGCATCGGGGAAGTACTTTGCGGATTGCAACGAAGCAGGA
GTGTCCAAATTGGGATCCAGCTCAAGAGAAGCCGCTCGGCTATGGTCCGCATCCGAAACCATGGTTTCTTCTGCGTCGTCCACGAACTCAAAAGCATCAG
CTTTTCATGCACTTTTGCTTTAG
AA sequence
>Lus10003204 pacid=23155743 polypeptide=Lus10003204 locus=Lus10003204.g ID=Lus10003204.BGIv1.0 annot-version=v1.0
MLQTVKYLIGARGASGYGSKSTAEEVTENCSDLRSVTAIITGATSGIGAETARVLAKRGATVVIPVRSMKAAEDTKARILSECPDAAIIVMALDLSSLRS
VRSFVSEFESLDLPLNLLINNAGKFAHQHAISEDGIEMSFATNYLGHFLLTNLLLEKMKETAEETGVQGRIVNVSSAIHGWFAGDMLQYLEEISGQKCPH
YDATRAYALSKLANVLHTKHLALKLKETDANVTVNCVHPGIVRTRLTREREGFATDAAFFLASKLLKTIPQAAATTLYVATHPRVGNASGKYFADCNEAG
VSKLGSSSREAARLWSASETMVSSASSTNSKASAFHALLL
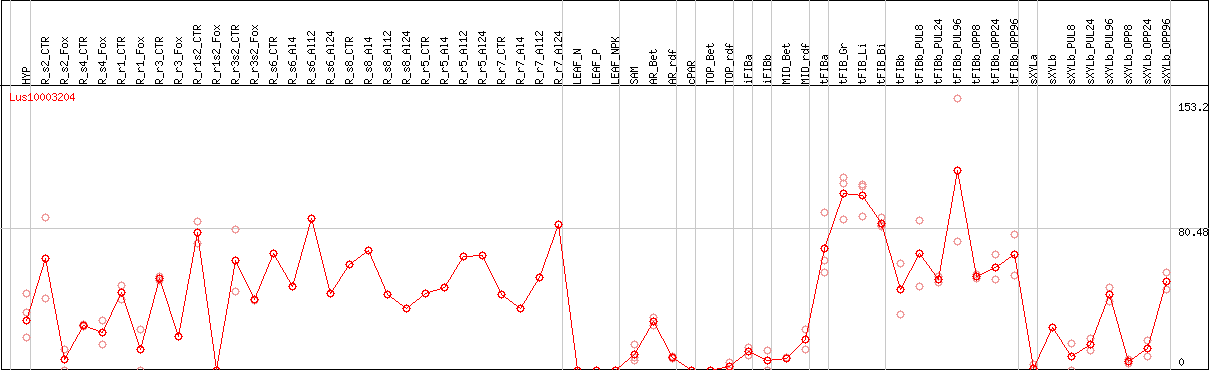
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10003204 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.