Lus10005134 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10005134 pacid=23141912 polypeptide=Lus10005134 locus=Lus10005134.g ID=Lus10005134.BGIv1.0 annot-version=v1.0
ATGGACGCTCTCTCATCCGGTGAAGACTTGCTTATCAAGACGAGAAAACCGTATACAATTACCAAGCAAAGGGAGCGATGGACCGATGAGGAGCACAATA
GGTTCCTCGAGGCTCTGAAGCTCTATGGTCGAGCTTGGCAGAAAATAGAAGAACATATAGGGACAAAGACTGCCGTGCAGATTAGAAGTCATGCACAGAA
ATTCTTCTCAAAGTTGGAGAAGGAAGCAGTAACGAAAGGTGTTCCAGTAGCACAGGCTCTTGACATTGATATTCCACCTCCACGTCCTAAGCGGAAGCCA
AGCAATCCATACCCCCGTAAAACAGGTGCCAGTTTGGTCACATCTCAGGCGGGAGCTAAGGAAGATGGAAAGCTTTCAACTTCGCTTCCCTTTCCTCCTA
GCAAAGTGCTGGACTTGGAGAAAGAACCACTTCCTGAGAAACCTAGTGGTGATGAGAAGTCAACAGGTGCCAAAGAAAACCATGATGAAAGTTGCTCAGA
GGTCTTCACCAGCAGTTCCTCCATCTTCTCATTGAAGAAAAACTCCACACTGCAACCAGGAGGCCCGAGGAATCCATGCTCCTTTAGGGAGTTTGTACCT
TGCCCGAAAGACGCGGCAAGTCAGGATGAGTCTCGTGTCACCACTGAACTTGATGGAAATAAGAAATGGCATAAACCAGAGCTCACACCAAAATTCTATG
ACAATGGCAGTAGTAAGGCCCCACAGTTGGGACACTCCTGCCATTCGCTCGAGAAGTTACATCAACACAAGAACAACGCTGATGGAGATGCAATCATGGT
CCCGGAAGACAAGATCCAAGCGATGCAGAACTGTCCTCGGCACGTACCGGTGCACATTCTAGATGGGAATGTTCCATCTAATGAATCAACGGTCCCTGTA
GAAGAAAGTCACGCACGGTCTGATGTTTACTCACTTCCGACGGCATCTGCAGCTGAATTTCCGAACAATATGGCTGCATGTTCGACGAACCAATCATTTC
CGCCTTTTCCTCATCCATTTGCACCAAGTCTCCATAATCAGGATGACTACAGGTCCTTCGTCCAGATGTCATCTGCGTTTTCGAGCCTCATTGCATCTAC
TCTGCTACAAAATCCTGCTGCCCATGCAGCAGCAAGTTTCGCCGCCTCCTTTTGGCCCTATGCAAATGTCGGTAACCCCGCAGACTCTTCAACATTCACT
TCAGGAGGATTTTCATCAAGACCAGTCCACGCTTCTGCTCCTAGTATGGCAGCCATAGCTGCAGCTACTGTCGCGGCAGCAACTGCATGGTGGGCTGCTC
ACGGTATGCTTCCCATGTGTGCCCCTCAGCATACCGCCTTCACTTGTCCTCCGGCACCGGGAATTCCAGCGGCTCATAAGGATATGCTCGGTACTGAGAA
AATTGAAACTAAAGAAGGCAAACTTGAACATCCTCCTTTAGACGATGTACGAATGGATCATGTCAAAAATTCTGAGACAGGGCAAGCACAGAACTCATCC
TCCAAGTCACCTACAATGTCTGAATCACCTAAAGTGGCATACCAGGAGATGAATCCAACGACAGCACCCGTGGCACATGATTGTAGCGAACCGAAGAACA
AGAAACAAGTCGATCGATCCTCATGCGGCTCGAATACAGCTTCCAGCAGCGAAGTGGAGACGGATGCGCTAGAGAAGAATAATGGGAACGAAGGCAATGA
CGAAACGAAAGAAGCTGATGCTGAAAATCTTCTTCCGGCCGCAATAACAGAGCCTGGCACTCGTAGAATACTTCGAAGCAATGGAGGCATGAACGATTCG
TGGAAGTCGGTCTCGCAAGTGGGGCGCCTCGCCTTCCAAGCACTATTCACCAGGGAAAAGCTCCCGCAAAGCTTCTCACCCCCTCAGAACGGAGCCCACC
GCGGAGTCGAGATGATGCAGAACGAGGACGAGAGAACCAATGGAGAGGCCACAACAATAATAGATCTCAACAACGACGCGTGGTGCCATGATCAACAAGA
GGAGGAGAACAATGCCGCCTCGTGTAATCACGAGGAGGAATGCTTGCCGAAGATAGGAATCGAAAATGGGAAACTACTTAAAGCTCGACGGACAGGATTC
AAACCTTACAAGCGATGTTCAATGGAGGCCAAGGAAAACACGTTGACAACCAGCAGCCAAGGGGACGAGAAGGAGTCCAAACGGATACGCCTCGCCCTCG
AAGGGGAAGCTTCGATCTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10005134 pacid=23141912 polypeptide=Lus10005134 locus=Lus10005134.g ID=Lus10005134.BGIv1.0 annot-version=v1.0
MDALSSGEDLLIKTRKPYTITKQRERWTDEEHNRFLEALKLYGRAWQKIEEHIGTKTAVQIRSHAQKFFSKLEKEAVTKGVPVAQALDIDIPPPRPKRKP
SNPYPRKTGASLVTSQAGAKEDGKLSTSLPFPPSKVLDLEKEPLPEKPSGDEKSTGAKENHDESCSEVFTSSSSIFSLKKNSTLQPGGPRNPCSFREFVP
CPKDAASQDESRVTTELDGNKKWHKPELTPKFYDNGSSKAPQLGHSCHSLEKLHQHKNNADGDAIMVPEDKIQAMQNCPRHVPVHILDGNVPSNESTVPV
EESHARSDVYSLPTASAAEFPNNMAACSTNQSFPPFPHPFAPSLHNQDDYRSFVQMSSAFSSLIASTLLQNPAAHAAASFAASFWPYANVGNPADSSTFT
SGGFSSRPVHASAPSMAAIAAATVAAATAWWAAHGMLPMCAPQHTAFTCPPAPGIPAAHKDMLGTEKIETKEGKLEHPPLDDVRMDHVKNSETGQAQNSS
SKSPTMSESPKVAYQEMNPTTAPVAHDCSEPKNKKQVDRSSCGSNTASSSEVETDALEKNNGNEGNDETKEADAENLLPAAITEPGTRRILRSNGGMNDS
WKSVSQVGRLAFQALFTREKLPQSFSPPQNGAHRGVEMMQNEDERTNGEATTIIDLNNDAWCHDQQEEENNAASCNHEEECLPKIGIENGKLLKARRTGF
KPYKRCSMEAKENTLTTSSQGDEKESKRIRLALEGEASI
|
DESeq2's median of ratios [FLAX]
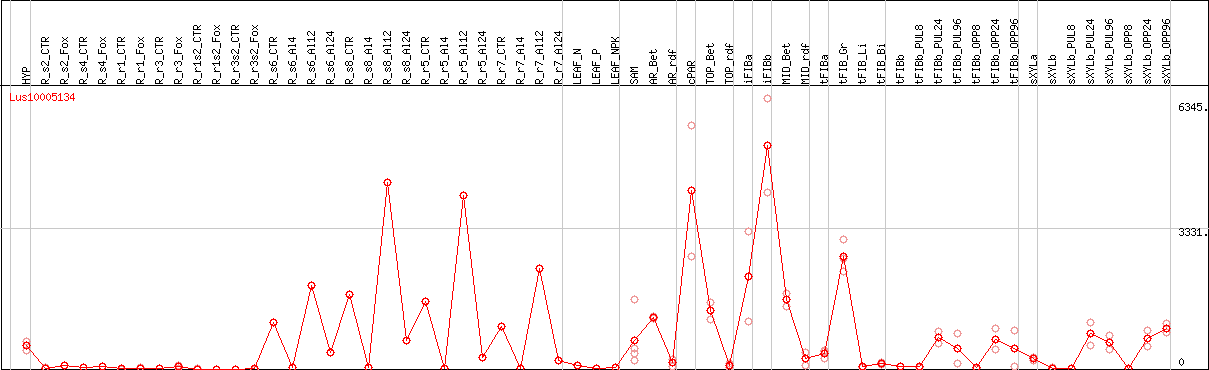
Coexpressed genes
Lus10005134 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
