External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G28470 267 / 4e-88
MYB
TDF1, ATMYB35
DEFECTIVE IN MERISTEM DEVELOPMENT AND FUNCTION 1, MYB DOMAIN PROTEIN 35, Duplicated homeodomain-like superfamily protein (.1)
AT5G56110 213 / 5e-67
MYB
MS188, ATMYB80, AtMYB103
MALE STERILE 188, ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 80, myb domain protein 103 (.1)
AT3G02940 195 / 4e-60
MYB
ATMYB107
myb domain protein 107 (.1)
AT5G15310 195 / 4e-60
MYB
ATMYB16, ATMIXTA
myb domain protein 16 (.1.2)
AT5G16770 192 / 1e-58
MYB
ATMYB9
myb domain protein 9 (.1.2)
AT5G26660 192 / 2e-58
MYB
ATMYB4, ATMYB86
myb domain protein 86 (.1)
AT4G05100 191 / 2e-58
MYB
ATMYB74
myb domain protein 74 (.1)
AT4G28110 189 / 3e-58
MYB
ATMYB41
myb domain protein 41 (.1)
AT4G01680 188 / 3e-58
MYB
ATMYB55
myb domain protein 55 (.1.2.3)
AT5G10280 189 / 1e-57
MYB
ATMYB92, AtMYB64
myb domain protein 92 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10039462
546 / 0
AT3G28470 284 / 9e-95
DEFECTIVE IN MERISTEM DEVELOPMENT AND FUNCTION 1, MYB DOMAIN PROTEIN 35, Duplicated homeodomain-like superfamily protein (.1)
Lus10033119
242 / 8e-78
AT3G28470 214 / 3e-67
DEFECTIVE IN MERISTEM DEVELOPMENT AND FUNCTION 1, MYB DOMAIN PROTEIN 35, Duplicated homeodomain-like superfamily protein (.1)
Lus10036660
241 / 3e-77
AT3G28470 213 / 2e-66
DEFECTIVE IN MERISTEM DEVELOPMENT AND FUNCTION 1, MYB DOMAIN PROTEIN 35, Duplicated homeodomain-like superfamily protein (.1)
Lus10033003
194 / 7e-59
AT5G15310 323 / 2e-109
myb domain protein 16 (.1.2)
Lus10033473
186 / 8e-58
AT2G16720 249 / 2e-83
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 7, myb domain protein 7 (.1)
Lus10018418
189 / 1e-57
AT4G21440 334 / 5e-114
A. THALIANA MYB 4, MYB-like 102 (.1)
Lus10039743
185 / 3e-57
AT4G21440 296 / 2e-100
A. THALIANA MYB 4, MYB-like 102 (.1)
Lus10002593
187 / 9e-57
AT4G21440 330 / 2e-112
A. THALIANA MYB 4, MYB-like 102 (.1)
Lus10028435
182 / 2e-56
AT2G16720 246 / 2e-82
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 7, myb domain protein 7 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.017G075000
275 / 2e-90
AT3G28470 254 / 2e-82
DEFECTIVE IN MERISTEM DEVELOPMENT AND FUNCTION 1, MYB DOMAIN PROTEIN 35, Duplicated homeodomain-like superfamily protein (.1)
Potri.015G067700
250 / 9e-81
AT3G28470 222 / 6e-70
DEFECTIVE IN MERISTEM DEVELOPMENT AND FUNCTION 1, MYB DOMAIN PROTEIN 35, Duplicated homeodomain-like superfamily protein (.1)
Potri.012G072500
249 / 1e-80
AT3G28470 228 / 3e-72
DEFECTIVE IN MERISTEM DEVELOPMENT AND FUNCTION 1, MYB DOMAIN PROTEIN 35, Duplicated homeodomain-like superfamily protein (.1)
Potri.011G167600
211 / 3e-66
AT5G56110 338 / 3e-116
MALE STERILE 188, ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 80, myb domain protein 103 (.1)
Potri.001G470500
209 / 1e-65
AT5G56110 334 / 2e-114
MALE STERILE 188, ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 80, myb domain protein 103 (.1)
Potri.004G033100
195 / 3e-59
AT4G21440 293 / 4e-97
A. THALIANA MYB 4, MYB-like 102 (.1)
Potri.010G165700
194 / 4e-59
AT3G01140 302 / 3e-100
NOECK, myb domain protein 106 (.1)
Potri.015G082700
194 / 5e-59
AT1G57560 254 / 2e-82
myb domain protein 50 (.1)
Potri.019G118200
193 / 7e-59
AT4G05100 278 / 7e-92
myb domain protein 74 (.1)
Potri.017G086300
194 / 1e-58
AT5G15310 343 / 6e-117
myb domain protein 16 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0123
HTH
PF00249
Myb_DNA-binding
Myb-like DNA-binding domain
Representative CDS sequence
>Lus10005834 pacid=23180245 polypeptide=Lus10005834 locus=Lus10005834.g ID=Lus10005834.BGIv1.0 annot-version=v1.0
ATGGGGAGACCTCCTTGCTGTGACAAGTCGAATGTGAAGAGAGGACTTTGGACCCCTGAGGAAGATGCCAAGATCCTTGCTTATGTCTCCCTCTATGGCA
TTGGCAATTGGACCTTAGTCCCCAAAAAAGCCGGACTGAATAGATGTGGAAAGAGTTGCAGGCTTAGATGGACTAATTACTTAAGACCTGATCTCAAACA
TGATAACTTCACTCCTGATGAGGAAGAGCTCATTATCAGACTTCATGAAGCCATTGGCAGCAGGTGGTCTATAATTGCAAGGCAACTGCCAGGGAGAACA
GACAATGATGTGAAGAACTACTGGAACACAAAACTAAAGAAGAAGCTTCAGAAAATAGGGATTGACCCAATCACTCACAAACCCATCTCCAAGATCATCA
ATGACTTTGGCAACATCAGTGGAATCAATCATGGAAACCAATCTGCATCCCCTTTCATGAGGAAGACACTCAACAGCAATAATCTTAACAACAACAACAA
TAATCCCATGGTCATGAACCTCCAATCAGAACCATCTTCAACTACTACTACAGCCTCATCTTGCTCCAACACCACCACATTCATGGTTGAAAATAATACT
GGTTGCTCATCTGACTACTACTTTTCTCATACCAACAACAGCTTTTTCAATCATGACTACTACTCATCCTCCTCTGCCTCATCATCTTCGACTACTCCTC
TGTCATCATCATGCCAGCAGTACCCTCAAACTGATCCACTGACGTGGAAGGAGTTCCTTGTTAGCGACCCACCAACATCATCATCAGCAGCAGCAAACTT
GTCAGAAGGAAACGGATGTTTGGATATGAATGAAAGGGGAAATATTGGGAGTAACAAAAATATTAATGCAGATGATAATAATGAGGGTGCTGCTAGCTCA
TTTGTGGAGAATATGCTTGATAAGGAGAGGGAGATGATGCTCTCTCAGTTTCCACAGCTTTTGGATCCTTCTTCTTTTGATTACTGA
AA sequence
>Lus10005834 pacid=23180245 polypeptide=Lus10005834 locus=Lus10005834.g ID=Lus10005834.BGIv1.0 annot-version=v1.0
MGRPPCCDKSNVKRGLWTPEEDAKILAYVSLYGIGNWTLVPKKAGLNRCGKSCRLRWTNYLRPDLKHDNFTPDEEELIIRLHEAIGSRWSIIARQLPGRT
DNDVKNYWNTKLKKKLQKIGIDPITHKPISKIINDFGNISGINHGNQSASPFMRKTLNSNNLNNNNNNPMVMNLQSEPSSTTTTASSCSNTTTFMVENNT
GCSSDYYFSHTNNSFFNHDYYSSSSASSSSTTPLSSSCQQYPQTDPLTWKEFLVSDPPTSSSAAANLSEGNGCLDMNERGNIGSNKNINADDNNEGAASS
FVENMLDKEREMMLSQFPQLLDPSSFDY
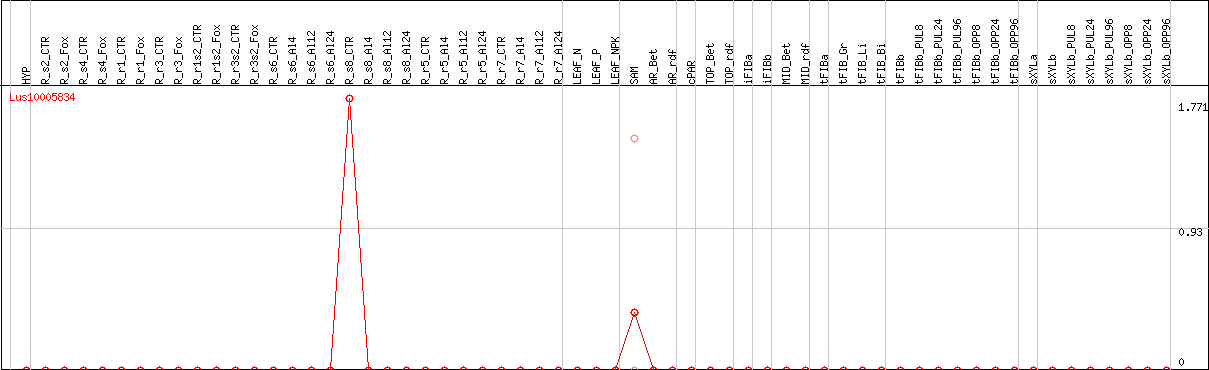
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10005834 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.