External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G66370 119 / 4e-35
MYB
ATMYB113
myb domain protein 113 (.1)
AT1G66380 107 / 1e-31
MYB
ATMYB114
myb domain protein 114 (.1)
AT1G56650 105 / 6e-30
MYB
ATMYB75, SIAA1, PAP1
SUC-INDUCED ANTHOCYANIN ACCUMULATION 1, MYELOBLASTOSIS PROTEIN 75, MYB DOMAIN PROTEIN 75, production of anthocyanin pigment 1 (.1)
AT1G66390 105 / 7e-30
MYB
PAP2, AtMYB90
PRODUCTION OF ANTHOCYANIN PIGMENT 2, myb domain protein 90 (.1)
AT1G22640 100 / 7e-28
MYB
AtMYB3
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 3, myb domain protein 3 (.1)
AT3G12720 100 / 3e-27
MYB
ATMYB67, AtY53
myb domain protein 67 (.1)
AT3G13890 100 / 4e-27
MYB
ATMYB26, MS35
MALE STERILE 35, myb domain protein 26 (.1.2)
AT5G35550 99 / 6e-27
MYB
ATTT2, TT2, AtMYB123
TRANSPARENT TESTA 2, MYB DOMAIN PROTEIN 123, Duplicated homeodomain-like superfamily protein (.1)
AT5G16770 100 / 7e-27
MYB
ATMYB9
myb domain protein 9 (.1.2)
AT5G40330 97 / 1e-26
MYB
ATMYBRTF, ATMYB23
myb domain protein 23 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10003277
153 / 3e-48
AT1G66370 160 / 2e-48
myb domain protein 113 (.1)
Lus10042522
150 / 5e-47
AT1G66370 181 / 7e-57
myb domain protein 113 (.1)
Lus10009130
131 / 4e-39
AT1G66370 202 / 3e-64
myb domain protein 113 (.1)
Lus10028513
128 / 4e-38
AT1G66370 205 / 3e-65
myb domain protein 113 (.1)
Lus10028514
120 / 3e-35
AT1G66370 179 / 1e-55
myb domain protein 113 (.1)
Lus10009129
120 / 4e-35
AT1G66370 179 / 2e-55
myb domain protein 113 (.1)
Lus10000470
108 / 3e-30
AT2G16720 196 / 2e-61
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 7, myb domain protein 7 (.1)
Lus10033438
106 / 1e-29
AT1G22640 194 / 2e-61
ARABIDOPSIS THALIANA MYB DOMAIN PROTEIN 3, myb domain protein 3 (.1)
Lus10000008
100 / 1e-29
AT5G49330 142 / 5e-43
PRODUCTION OF FLAVONOL GLYCOSIDES 3, ARABIDOPSIS MYB DOMAIN PROTEIN 111, myb domain protein 111 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0123
HTH
PF00249
Myb_DNA-binding
Myb-like DNA-binding domain
Representative CDS sequence
>Lus10006541 pacid=23149213 polypeptide=Lus10006541 locus=Lus10006541.g ID=Lus10006541.BGIv1.0 annot-version=v1.0
ATGGGAGGAGTTGCCTGGACTGAAGAAGAAGATCACTTGCTCAGAAAATGCATCGAGCAGTACGGCGAAGGCAAATGGCACCGTGTCCCCTTGCTTGCTG
GTCTAAATAGGTGCCGTAAGAGCTGCAGGCTGAGGTGGCTGAACTACCTGCGGCCGAATATAAGGAGGGGGAGTTTCGAGGATGAGGAAGTTGAACTCAT
CATTAAGCTTCATAAACTGTTGGGTAACAGGCAAGTTTAA
AA sequence
>Lus10006541 pacid=23149213 polypeptide=Lus10006541 locus=Lus10006541.g ID=Lus10006541.BGIv1.0 annot-version=v1.0
MGGVAWTEEEDHLLRKCIEQYGEGKWHRVPLLAGLNRCRKSCRLRWLNYLRPNIRRGSFEDEEVELIIKLHKLLGNRQV
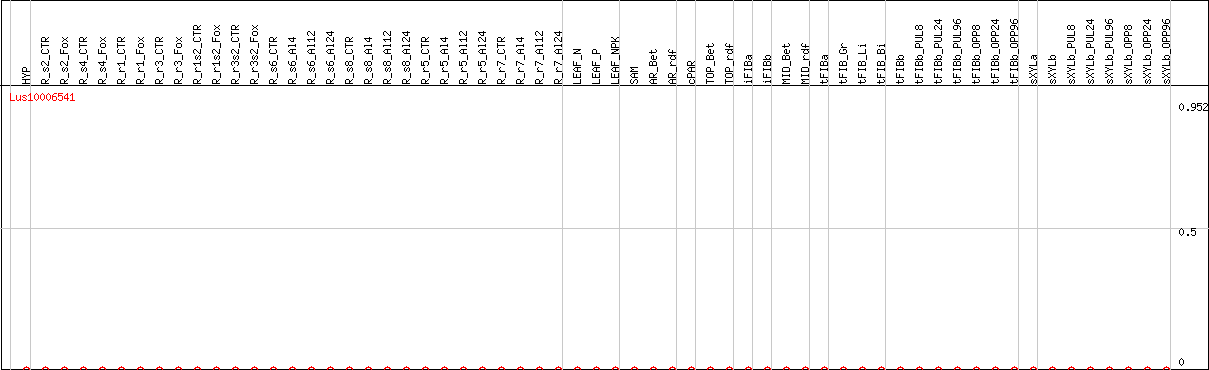
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10006541 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.