External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G63380 353 / 3e-123
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT1G62610 352 / 5e-123
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2), NAD(P)-binding Rossmann-fold superfamily protein (.3), NAD(P)-binding Rossmann-fold superfamily protein (.4)
AT3G55290 345 / 2e-120
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT3G46170 343 / 4e-119
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G17845 340 / 2e-117
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G55310 338 / 3e-117
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G01980 117 / 3e-31
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2), NAD(P)-binding Rossmann-fold superfamily protein (.3), NAD(P)-binding Rossmann-fold superfamily protein (.4)
AT1G24360 103 / 1e-25
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT5G06060 88 / 3e-20
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G03980 85 / 4e-19
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10007039
457 / 5e-164
AT3G55290 369 / 3e-129
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10006699
427 / 3e-152
AT3G46170 404 / 2e-143
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10007038
417 / 2e-148
AT1G63380 357 / 7e-125
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10006695
160 / 1e-49
AT3G55310 107 / 9e-30
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10036980
110 / 3e-28
AT1G24360 437 / 3e-155
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10015821
103 / 1e-24
AT1G24360 427 / 5e-148
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10012548
100 / 1e-24
AT3G01980 337 / 2e-117
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2), NAD(P)-binding Rossmann-fold superfamily protein (.3), NAD(P)-binding Rossmann-fold superfamily protein (.4)
Lus10041545
96 / 3e-23
AT3G01980 346 / 6e-121
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2), NAD(P)-binding Rossmann-fold superfamily protein (.3), NAD(P)-binding Rossmann-fold superfamily protein (.4)
Lus10022756
90 / 1e-20
AT3G51680 397 / 2e-139
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.016G035200
395 / 8e-140
AT3G55290 385 / 8e-136
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.010G056100
118 / 4e-31
AT1G24360 421 / 6e-149
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.008G178700
114 / 1e-29
AT1G24360 385 / 1e-134
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.004G200100
102 / 2e-25
AT3G26770 257 / 2e-85
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.017G067332
99 / 4e-24
AT3G01980 323 / 4e-112
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2), NAD(P)-binding Rossmann-fold superfamily protein (.3), NAD(P)-binding Rossmann-fold superfamily protein (.4)
Potri.005G106100
96 / 3e-23
AT2G47140 234 / 9e-77
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.004G199900
96 / 4e-23
AT2G47140 256 / 1e-85
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.006G207100
92 / 1e-21
AT4G03140 241 / 2e-78
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.016G073900
90 / 7e-21
AT3G51680 252 / 2e-83
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.006G206700
89 / 9e-21
AT3G51680 236 / 4e-77
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF00106
adh_short
short chain dehydrogenase
Representative CDS sequence
>Lus10006698 pacid=23152215 polypeptide=Lus10006698 locus=Lus10006698.g ID=Lus10006698.BGIv1.0 annot-version=v1.0
ATGGCCAGTGAGGTGTCAGCCCAGCTGGAGCCATGGTGCAACTTACCAGACAAAGTGGTCCTTGTAACCGGTGCTTCTTCGGGTTTAGGCCGAGAGTTCT
GCCTCGATCTAGCCAAGGCTGGGTGCAAGATCGTTGCAGCTGCTAGGCGCATTGACCGCCTACAATCTTTGTGCAACGAAATCAATCAGCTAACTGCTTC
ATCATCTGCGGGAGAGCCAATTGGTCCACGTGCTATGGCTGTGGAACTGGATGTGTCAGCAGACGGGCCTGCTATTGATAAGGCTGTACAGAGGTCTTGG
GAAGCCTTTGGAAGGATAGATGTGTTGATCAACAATGCTGGCATTAGCGGTAGCACGAAGAACTCGTTGATTTTATCTGAAGAGGAATGGAATCAAGTGA
TCAAGACGAACTTGACAGGAACCTGGTTGGTTTCTAAGTCAGCTGGGATACGGATGCGCGATGCAAAGCTGGGAGGTTCCATAATCAATATTTCGTCGAT
CTCTGGTCTTAGTCGTGGTTATACACCTGGAGTTGTCGCTTATGCTTCTTCGAAGACCGGTGTAAATTCCATGACAAAGGTGATGGCGCTGGAGTTGGGC
GCTTACAATATCAGAGTTAACTCTATATCGCCCGGACTGTTCAAATCCGAGATCACCAAAGGTCTCATCCAAAAGCCTTGGCTGACTACCATTGCTGAGA
AGACGATCCCTCTACGAACATTTGGAACTGTAGACCCAGCATTGACATCACTCGTTCGATACCTCATCTATGATTCAACCCGGTATGTGACGGGCAATGT
GTTCATTGTTGATGCCGGAACTACCGTACCAGGTGTCCCCATTTTCTCATCGCTCTGA
AA sequence
>Lus10006698 pacid=23152215 polypeptide=Lus10006698 locus=Lus10006698.g ID=Lus10006698.BGIv1.0 annot-version=v1.0
MASEVSAQLEPWCNLPDKVVLVTGASSGLGREFCLDLAKAGCKIVAAARRIDRLQSLCNEINQLTASSSAGEPIGPRAMAVELDVSADGPAIDKAVQRSW
EAFGRIDVLINNAGISGSTKNSLILSEEEWNQVIKTNLTGTWLVSKSAGIRMRDAKLGGSIINISSISGLSRGYTPGVVAYASSKTGVNSMTKVMALELG
AYNIRVNSISPGLFKSEITKGLIQKPWLTTIAEKTIPLRTFGTVDPALTSLVRYLIYDSTRYVTGNVFIVDAGTTVPGVPIFSSL
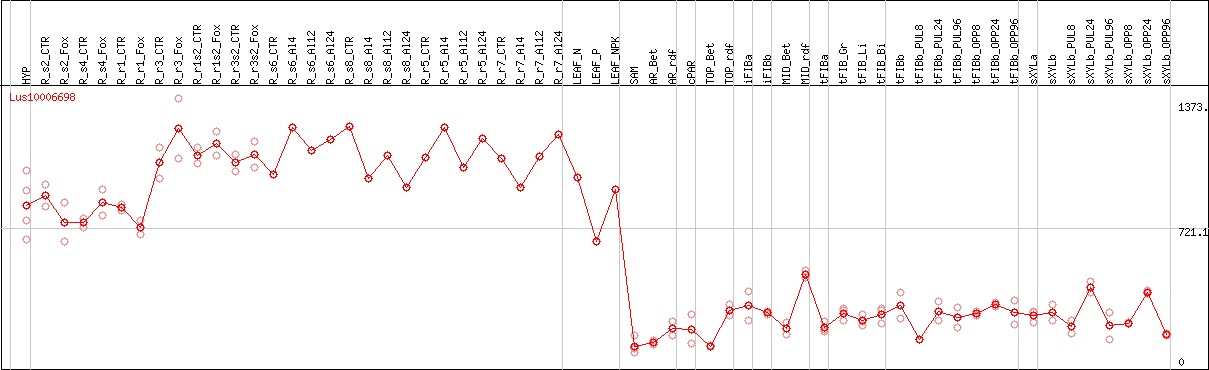
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10006698 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.