Lus10007261 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10007261 pacid=23165710 polypeptide=Lus10007261 locus=Lus10007261.g ID=Lus10007261.BGIv1.0 annot-version=v1.0
ATGGATGTAGCTTTAATCGGAGGATCATTCCTCTCTGCGTTTTTCCAGGTCTTGTTTGACAGGATGGCTTCTCCTGAGATAGTTAATTTCATCAGGGGCC
ACAAACTCAATCACGTAATGATGAACAGGTTAAACCTGATGTTGAATTCTATCAGCGGAGTGCTTGATGATGCAGAAGAGAAACAGATAAGTAAGCCATC
CGTGAAGAGGTGGCTTGATGAACTGAAAGATGCTACTCTTGAAGCTGATGACTTGTTGGATGAGATATGTTACGAATTCAAGCGGTTGGAGCTGGGACAT
TGGGGAAAAGTGCGCAACTTCATATCTTACAGCTCATTCAGAGATGGTACCAATAAAAAGTTGAAAGAGATTCTTGACAGGCTGGAGGATTTAGTAAAAC
AGAAGGATGTTCTTGGTCTGATTGAAGGCATTGCACGGAACCCATCATTTCAAAACAGAGTGACCACCTCTCTCCCAGATGGGTCAGCTGTTTACGGCAG
GAAGAATGATGTAGAAGCCATAATATCTTTGCTGCAATCAAATTTCACTGCGAGTAGTAGCAACTCTATAGGTTTAATCCCACTTCTGGGTATGGGTGGT
ATTGGTAAAACCACCATTGCTCAACTTGTCTACAAGGACATCAGAGTCCAACAATGGTTTGAGCTTAGGGCATGGATTTATGTCTCGGCCGAATTTGATG
TCTATAAGGTGTCAAAAGATATTCTTGAAGAAGTCTGTGGTAGAAGTTGTGGCAGCACGACTCTCAATCAACTTCAACTTGGGTTGGAGAAGGCATTTGA
AGGGAAAAGGTGTTTGCTAGTCTTTGATGACGTTTGGCATGACAAGTCTGCTTCTGACTGGGTCAACTTGCTTAAACCCTTGCAGTCCATGGCACCAGGA
AGTATGATAATTGCTACGACCCGGAATCATAGTGTAGTGTCAGCTCTGCGCACTGTGCCAACCATTCGAATGTTTCCTCCTTACAATCTACAGATTCTAG
ATGATGAAAACTGTTGGTCTTTGTTCGCTAAGACTGCTTTCGGTGATGGATATTCTAGTGCATGGGAGAAGTTACACGGCATTGGCAGAGAAATAATGAA
GAAATGTAATGGCCTTCCATTAGCTGCAAAAACTCTGGGATGTCTTCTCCGTGCTGATTCGGATGTAGAAAAATGGCAGAATATATTGAAGAGCAACATA
TGGGATTCATCAAATGACGACATTCTTCCAGCATTAATCTTAAGTTATCACTATCTTCCATCGCATCTAAAACCGTGTTTTGCGTACTGTGCAATGTTCC
CCAAAGGATATGAATTTGAGAAGGAAGAAGTAGTCCGTTTATGGATGGCAGAGGGTTTTGTGATTCAACCAAAAGGAAGCAGGGTAATGAAAGACACTTG
TGAGGACTACTTCAATGACCTTGTATCAAGGTCATTTTTACAACTCACAAGTGAACATCAGTTCTGCTTTACGATGCATGACCTCATACATGACCTGGCC
AAATACGTTGCTGGAGAATTTTGCTATAGGATGGAGGATGCCAATTCATTCAAGGTTGCAGGAAGAACTCGTCATCTATCATTTGCAAGGGCAGAACTTG
ATACCAGTATCATATGGGAGGGTATGTATAACGCTAAACGTTTGCGGACCTTATTGCCGGTGGATGGATCAAAATGGTGGAGTTATGATCCTGAGTGCAT
AGATGACAAGGTACTCCAAAACATATTTTTGTCACTTACACGATTACGGGTGCTATCTTTGTCTGGGTTTAAAGTGAGCAAGTTGCCTGACGCAATTGGT
GCCTTGAAGCATTTAAGATATCTGAATCTCTCCCTAGTTCTGTTACATAAGTTGCCTGATACCCTGTGCAACTTGTATAATTTGGAAATCTTGGTCTTGC
ACAAATGCAAAAATATTCTGGCTTTACCTGATTTGATTGGCAATTTAAAGCAACTGCAGATTATGGATATCAGTGGAACACAAATTACAAGGTTTCCAGA
TTCTTTTGTCAGTTTAATCAACTTATGTAGCCTAGATATTACAGAAACTACGTTGCAGGAGATGCCACTACAGATGAGTAACTTGTCAAGTCTTCGGAAT
TTAACTGACTTTGTTATTGGAGCTCAGAGTGGGTCTCGTATTAAGGAGATGGGGGGGCTTCAGCATCTCTTAGGAAGTCTTCGTATATGGAATCTTGAAA
ATGTTTTGGATTGTCAAGATGCCTTAGATGCTAGCTTCAAGATTAAGAAGCACATCAGTAGATTGGAGTTAAGATGGAATGGCGATACTAATGATTCATT
GCATGAAAGAGGTGTATTGGAGAACTTGCAGCCTCATATGAACTTGCAGCATCTTGTCATAGAAGGTTATGGAGGAACAAGCTTTCCAGGTTGGATTGGA
CATTCATCCTTCTCTTATTTACTATCTATCGAACTTACTGGATGCAAGTACTGCTCCTCCTTACCACCACTTGGGCAGTTAATGTCTCTTGAAAAGCTTT
CTATCAAAGTATTTGAGGGAGTTGTCACTGTGGGTCCTGAGTTCTATGGCAGCAGCACATTCACAAACAAGCCATTTCGGTCTCTTAAAAGTTTGAAATT
TGAACAGTTGCCAGAATGGACTGACTGGATTGCTTATGTCGGACGAGATGGAAGTGAAGCTTTCCCAGTCCTCGAGGACCTTTGCATTAAACATTGTCCT
AACTTGAGGAAAGCACTACCTTCCCACCTTCCGTCTCTAGAAAGGCTCGAGATTAGAGGATGCCAACAGCTTAGATTGGATTCAGTTTCGTGGGCTTCAA
ACATAATTTCAATGAGATTGAAGGACGATACTCGTGATGTATGGCTGGAGATGTTGACCAAGGGGCTGCACCGTCTCCGAGTTCAAGTTGCTCAGGATTT
GCTAGATGACCTGGAACTGGTACTTGATCTTTCCAACACTTTACAGGAAATTATGGTCAAGGACTGTAATTCATTTAAGCACCTGCAACTAGAGTTGTTC
CCGAAATTGAAAGCTGTTTCCATAAATGGATGTCAAAGTTTAGCATCCCTCTGCAGACCTGACGCGACCTTGGACACTCTGGAGTATCTTAATTTCTTGG
AGATATCAAAGTGCCCAAATCTCCTCTCATTTCCTGCAGGACGATTACATGCCCCAGAAATGACCAGGCTCATATTGAGTGATTGTACCAACTTGAAGTC
TCTCCCTGCAGGTATGCACTCTCTTCTCCCATCACTAGTGGAGTTGTTCTTGGATAACTGTCCGGAGGTGGAGTCGTTCCCATCAGGTGGGTTACCCCTA
AAATTGGAGACACTAATTGTCAATGGATGCGACAAACTCATAGCTGGTTGCAGGAAATGGAATTTACAGTTACTGCCTTGTCTTTTGAGCCTTAGCATTG
GGCAGAACAAGGAACTGGAATGTTTACCATTGGAGGAAATGTTGCCATGCACGCTGAACTCTCTCAAAATCAGGAACTTTCAGAATTTGAAGTCTCTAGG
TTTCAATCGGATTCAGGGAGGTAAACACATGTCCATAAAAAAGTTGGAATTCAAAGACTGTCCCCTTTTAGAGTTATTCGGGATAGGCAGTATCTTCACC
GGTTTAGCATCACTTGAGATCAAAGGTTGCGACAGGCTCATAGCTGGGCGAAGCCAGTGGAATCTGGTAGCAATGTCATCTCTGGAGGAGTTATATTTAA
GTACCAACAACAACTTGGAATCCTTTCCGGAAGATATGCATCTGCCTCCAACTCTAACGTCTCTTTACTTGTGGAATCTTCCGAATCTGACGTGCCTAAA
CTGCAACAGACTTGAACACCTCACATCTTTTAAAGAACTGAAGATCTCAAACTGTCCGAAGCTGCAGTGTATCCCTGTGGATGGGCTTCCATCATCCCTT
TCGTCTGTATACATCAACTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10007261 pacid=23165710 polypeptide=Lus10007261 locus=Lus10007261.g ID=Lus10007261.BGIv1.0 annot-version=v1.0
MDVALIGGSFLSAFFQVLFDRMASPEIVNFIRGHKLNHVMMNRLNLMLNSISGVLDDAEEKQISKPSVKRWLDELKDATLEADDLLDEICYEFKRLELGH
WGKVRNFISYSSFRDGTNKKLKEILDRLEDLVKQKDVLGLIEGIARNPSFQNRVTTSLPDGSAVYGRKNDVEAIISLLQSNFTASSSNSIGLIPLLGMGG
IGKTTIAQLVYKDIRVQQWFELRAWIYVSAEFDVYKVSKDILEEVCGRSCGSTTLNQLQLGLEKAFEGKRCLLVFDDVWHDKSASDWVNLLKPLQSMAPG
SMIIATTRNHSVVSALRTVPTIRMFPPYNLQILDDENCWSLFAKTAFGDGYSSAWEKLHGIGREIMKKCNGLPLAAKTLGCLLRADSDVEKWQNILKSNI
WDSSNDDILPALILSYHYLPSHLKPCFAYCAMFPKGYEFEKEEVVRLWMAEGFVIQPKGSRVMKDTCEDYFNDLVSRSFLQLTSEHQFCFTMHDLIHDLA
KYVAGEFCYRMEDANSFKVAGRTRHLSFARAELDTSIIWEGMYNAKRLRTLLPVDGSKWWSYDPECIDDKVLQNIFLSLTRLRVLSLSGFKVSKLPDAIG
ALKHLRYLNLSLVLLHKLPDTLCNLYNLEILVLHKCKNILALPDLIGNLKQLQIMDISGTQITRFPDSFVSLINLCSLDITETTLQEMPLQMSNLSSLRN
LTDFVIGAQSGSRIKEMGGLQHLLGSLRIWNLENVLDCQDALDASFKIKKHISRLELRWNGDTNDSLHERGVLENLQPHMNLQHLVIEGYGGTSFPGWIG
HSSFSYLLSIELTGCKYCSSLPPLGQLMSLEKLSIKVFEGVVTVGPEFYGSSTFTNKPFRSLKSLKFEQLPEWTDWIAYVGRDGSEAFPVLEDLCIKHCP
NLRKALPSHLPSLERLEIRGCQQLRLDSVSWASNIISMRLKDDTRDVWLEMLTKGLHRLRVQVAQDLLDDLELVLDLSNTLQEIMVKDCNSFKHLQLELF
PKLKAVSINGCQSLASLCRPDATLDTLEYLNFLEISKCPNLLSFPAGRLHAPEMTRLILSDCTNLKSLPAGMHSLLPSLVELFLDNCPEVESFPSGGLPL
KLETLIVNGCDKLIAGCRKWNLQLLPCLLSLSIGQNKELECLPLEEMLPCTLNSLKIRNFQNLKSLGFNRIQGGKHMSIKKLEFKDCPLLELFGIGSIFT
GLASLEIKGCDRLIAGRSQWNLVAMSSLEELYLSTNNNLESFPEDMHLPPTLTSLYLWNLPNLTCLNCNRLEHLTSFKELKISNCPKLQCIPVDGLPSSL
SSVYIN
|
DESeq2's median of ratios [FLAX]
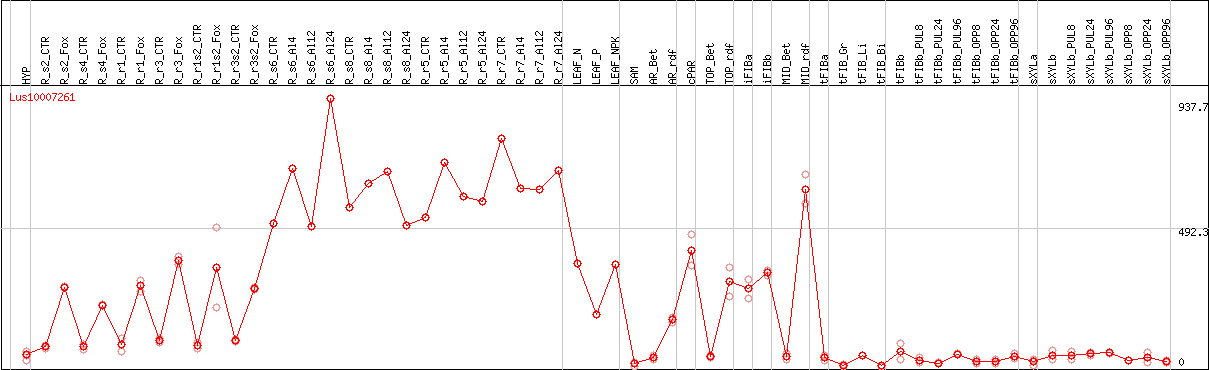
Coexpressed genes
Lus10007261 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
