Lus10008320 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10008320 pacid=23178496 polypeptide=Lus10008320 locus=Lus10008320.g ID=Lus10008320.BGIv1.0 annot-version=v1.0
ATGGCTTCCTCATCTACTACCCCTCCGTATACAGGGCAATGGCAACACGAAGTTTTCTTGTGCTTCAGAGGCGACGACGTGAGGGACAATTTCATGAGCC
ACCTTGAACGAGCTCTGAAGTCACGGAACATCGACTACTTCATCGACACCAAGCTTAATCCAACGCAGGACAATGAGGAACTGCTCGGGATCCTGGAAAG
GACGGCAGTTTCAGTGGTGATTTTTTCAGTCAAGTTTGCTGATTCGAGCTGGTGCTTGGACGAGGTCTGCACCATCGCTGAAAGCATTGAGAAATACAGA
CACAGGGTACTTCCGGTATTCTACAACGTGGATTGGAACACAGTGATCGGCGGCGGCAACTACCATTGGTTCAATGACTGGAAATCATCCTTCCTCTCTT
GCTTTGTCCCTCAGCCCACGCCGTACGCTAAAACTATCGCTGGACTGGAGGCCACCCAAGAGCGCAAGAAGAAATGGATAGCTTCTTTGAAGACAGTAGC
TTCCAAGACTGGGTGTACCTCTGAATCAATTCCCATCGATGCCGAGTTAGTTGAGAAGGTTGTGGCAGACGTCATTGCAACACTAGCATCCATGTCACCA
AGCATCGAGTACAAAAATTTGGTTGGTATGGATACACGCGTCTCAAAGGTCGAGGAATTGTTAGCCATGGATGATGGCGACACCAGTCTCATTGTCGGAC
TATGGGGACTGGGAGGTCTAGGTAAATCGACTCTAGCCAGGACGTGTTATGAAAAATTGAGGAAACCAAATGAAAAGAAGATGAAGTTTCATTTTGTCGA
GAAAATCAATGAGACATGCGATAAATCAAGTCCGGTTAAACTATTAGTGAAGGAACTCTACTCAACATTGCTGAATGAGGAGCCTGTTAAGAGTGGAGTT
GAGATTAGAGGGGAACGTCTTCCGCGTTTAAAAGTCTTTGTTGTTCTTGACGATGTGCAGAGCATTCCACAGCTGAATGAGTTGCTGTTAGAAAAGGCGT
TGAAACTCACAGAACTCTTTGCTCCCGGAAGCAGAATAATCATAACTACGAGGAATAAGGCTGTGCTGGAATACGCAACCGCGAAGATGTATGAAGTTGA
TCATCTCAATGATGGCGAGTCACTTGAACTCTTTAAATTAAATGCTTTTGAGGACCCGGGACATATTCGACATGTTGAACTATCACGTAAAGTTGTGACG
TACTGTCAGGGGAATCCATTGGCCCTGAAAGTGTTAGGAGCCACGTTGCTACGCAAGGACGAACCATACTGGGAGAGTTTCTTGGGCAAATTAGGAGAAA
TCCAGGACCAAACAATCCATTCTGTTCTCAGACGAAGCTACGACGAGTTGGGAAATGATGATAAGAGGCTGTTCCTGGATATTTCCTGCTTCTTTTATGG
TACTGTCAGGTCCTTGTTGGTGAAGTATATGGATTCTATTTACATCACTTCATATCACAGAGTTGAGGATCTGATCCACAAGTCTCTACTAATGACTACT
GATGACAAGATACACGGAGCGAAAGTTACGGTGCATAGTTTACTGAGAGAAATGGCTTGGAACGTTGTCAACGAGGAAGAAATTATCGGAGACCGCAGCA
GACTTAAAAACCATGATGACATTCGCATCTTGTTGAGTCAAGAGGGAAAAAGAGCAACCCAAGGGATATGTTTGGACCTTTCCAAAGCTGATGTTATGTT
TCTGAAAGCTGATGCTTTCAAAGGGATGGACTGTCTTAAATGGCTCGACTTTGGCTGGCCGAATTCTGTTGAATACGCCAACCGCAAAATCCAACTATCG
GATGGCGACATTAACTACCTTCCAAATGAGTTAAGGGGCTTGAATTGGGATCACTTCCCTTGCCATTCTTTGCCGTTGGGATTTTCTCCTAGAAAACTCG
TTTACCTCGTCATACGCCATAGTCCGATTGGCTTCTGTTGGGAGCGAAATCAACCTAAGCTTGAGCACCTGATGTTGCTCTGCCTCTCCAACTGTGAGAA
TCTAGCTATTGTTCCGGACCTGTCGTTCTGTTCAAATCTGGAACACATTCTACTCCGTGGATGCAGCACCTTAGTTGCGTTACCGGCAAGCATTCAATTC
CTTGAAAAGCTGGTAACGTTGGATGCCAGAGATTGTCGAAATCTCCAGCTGGTACCATTAGAGATCAACTCCAAGTTCCTTAAACAACTTTTGCTCTCCA
ACTGTCCCAAACTCACCGGCTGTCCCAAGGTTTACTCTCTAGAATTGCAGGCTCTGGATTTAGACGGGACACCCATTGGATCCCTCCCCGATTCAATCAA
CAAGGTTAAGCAGGATGGCGTAGTAAGCCTCTACGGCCCATGCATTGCGCATTTCCCGCAAATTCCTACCGGATTATCACTCCTCCTTCTGCGCAAAACC
GCAATCCAAGAAATCAAAATCGAGCTTCAGCCTCAGCAGCAGCTGAAGTTTGACCGGTTGCATTTGATCTGGAATGCCCAGCTCACCACTTTACCTACGG
AGATCTGGGATATGGTCTCCAACGAACTCATCGTCGAGGATTGTCCATTGATCGAATGTTTACCAGTGATATCGGAGCCTCGTTCGTACAGCTTAACAAC
GCTCCACATTGCTGGCTGCAAAAAGATCGCCGAGTTCCCGGCTTCCATATGCAAACTCGAGTGTTTGGAGGTACTGGTTTTTGTAGGCACGGATATCGAG
TCTTTGCCTACAAGTATTGGGGAGCTGAAGCAGCTGTCGTATTTAGACCTCTCGTACAGTACAAGACTGAAATCGGTACCTGGTACCATCAGCGAGCTCG
GGAAGCTATCCGAGCTTGTGCTGATTGGGTGTGGGAGCAGTCTGTCTTTGCCGGAACTACTCCCACCCAATCTTAATGTTGTGAAGGAGAAAAAATATCA
CAGGACGTCCACATTACAAACACTGTTGAGAACCATCATCGTGGATCAGAAATCAAACGACCTGATGCTTGATTTCCGCGCGCTACTAGCTGGAAGTTCG
GTTGTGAGTACCAAACTTCCACTTCAGAACATGTCGAGATGTCCCTTCGAGGCCGGTGCCTCTTCATTTTGTCATCGAAGCGTAATAACGAGGGTAGCGA
AGCTTAGGCAAAAAGAGTTGCGCGCCGACGAAGCTACTGGCGGACCAATGCGTAGGCAAAAACGAGAAAGGCAGAGTCGAGATCAGAATCAGGATCGTGT
GTTTACTGTACCGACCCAACAGTTAGATGCTTATGATAGGCCATTGCCCGATCACCGTAACAGTGTCGAAGACCAATCGTTACTTTCTGCCCTGAATCCA
GAAATACACAAGGCACTTAATAGAAGTATCAAGCGAGCGTCGGTCTTGAAAAAGATGCCCATACTGACGACGCTAAGTAAAAATGGCGGTTATCAGGCGC
CTTACACCATTCTCCAAATCAAGACGGCTGGGAAAAAAGTACATCAGCCGGCGATTTTGGATTTGATAGAAAACCGTATATCATTCCTTCCTAATAACTC
AAATTGTTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10008320 pacid=23178496 polypeptide=Lus10008320 locus=Lus10008320.g ID=Lus10008320.BGIv1.0 annot-version=v1.0
MASSSTTPPYTGQWQHEVFLCFRGDDVRDNFMSHLERALKSRNIDYFIDTKLNPTQDNEELLGILERTAVSVVIFSVKFADSSWCLDEVCTIAESIEKYR
HRVLPVFYNVDWNTVIGGGNYHWFNDWKSSFLSCFVPQPTPYAKTIAGLEATQERKKKWIASLKTVASKTGCTSESIPIDAELVEKVVADVIATLASMSP
SIEYKNLVGMDTRVSKVEELLAMDDGDTSLIVGLWGLGGLGKSTLARTCYEKLRKPNEKKMKFHFVEKINETCDKSSPVKLLVKELYSTLLNEEPVKSGV
EIRGERLPRLKVFVVLDDVQSIPQLNELLLEKALKLTELFAPGSRIIITTRNKAVLEYATAKMYEVDHLNDGESLELFKLNAFEDPGHIRHVELSRKVVT
YCQGNPLALKVLGATLLRKDEPYWESFLGKLGEIQDQTIHSVLRRSYDELGNDDKRLFLDISCFFYGTVRSLLVKYMDSIYITSYHRVEDLIHKSLLMTT
DDKIHGAKVTVHSLLREMAWNVVNEEEIIGDRSRLKNHDDIRILLSQEGKRATQGICLDLSKADVMFLKADAFKGMDCLKWLDFGWPNSVEYANRKIQLS
DGDINYLPNELRGLNWDHFPCHSLPLGFSPRKLVYLVIRHSPIGFCWERNQPKLEHLMLLCLSNCENLAIVPDLSFCSNLEHILLRGCSTLVALPASIQF
LEKLVTLDARDCRNLQLVPLEINSKFLKQLLLSNCPKLTGCPKVYSLELQALDLDGTPIGSLPDSINKVKQDGVVSLYGPCIAHFPQIPTGLSLLLLRKT
AIQEIKIELQPQQQLKFDRLHLIWNAQLTTLPTEIWDMVSNELIVEDCPLIECLPVISEPRSYSLTTLHIAGCKKIAEFPASICKLECLEVLVFVGTDIE
SLPTSIGELKQLSYLDLSYSTRLKSVPGTISELGKLSELVLIGCGSSLSLPELLPPNLNVVKEKKYHRTSTLQTLLRTIIVDQKSNDLMLDFRALLAGSS
VVSTKLPLQNMSRCPFEAGASSFCHRSVITRVAKLRQKELRADEATGGPMRRQKRERQSRDQNQDRVFTVPTQQLDAYDRPLPDHRNSVEDQSLLSALNP
EIHKALNRSIKRASVLKKMPILTTLSKNGGYQAPYTILQIKTAGKKVHQPAILDLIENRISFLPNNSNC
|
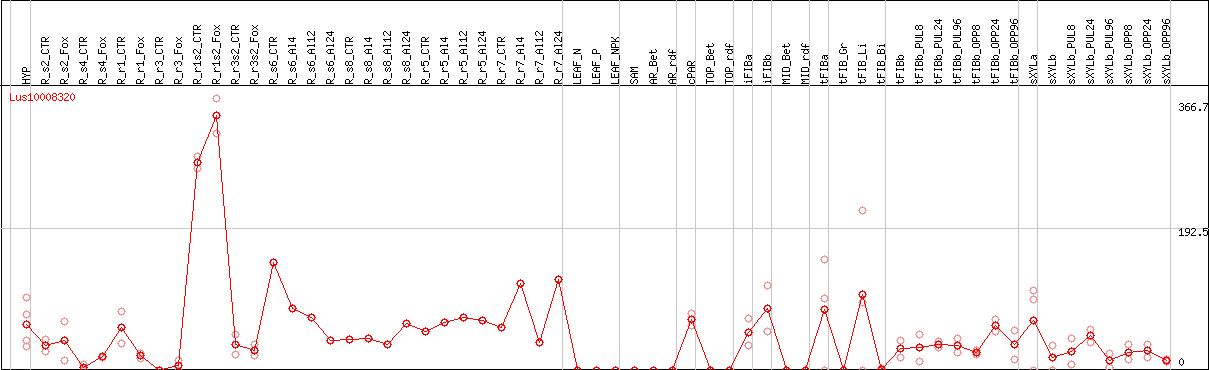
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10008320 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
