External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G27170 62 / 5e-13
transmembrane receptors;ATP binding (.1.2)
AT1G27180 54 / 4e-10
disease resistance protein (TIR-NBS-LRR class), putative (.1)
AT5G36930 50 / 4e-09
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
AT5G17890 46 / 1e-07
CHS3, DAR4
CHILLING SENSITIVE 3, DA1-related protein 4 (.1)
AT1G17600 45 / 3e-07
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT5G40910 45 / 4e-07
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT4G12010 44 / 9e-07
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT5G48770 44 / 1e-06
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT5G44870 44 / 2e-06
TTR1, LAZ5
tolerance to Tobacco ringspot virus 1, LAZARUS 5, Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT5G17680 43 / 2e-06
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10020533
99 / 6e-26
AT1G27170 405 / 1e-122
transmembrane receptors;ATP binding (.1.2)
Lus10020534
97 / 3e-25
AT5G36930 414 / 1e-124
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10005373
94 / 3e-24
AT5G36930 404 / 8e-121
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10020524
91 / 3e-23
AT1G27170 395 / 9e-119
transmembrane receptors;ATP binding (.1.2)
Lus10026961
91 / 4e-23
AT5G36930 420 / 2e-126
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10020537
91 / 4e-23
AT1G27170 404 / 6e-121
transmembrane receptors;ATP binding (.1.2)
Lus10042714
90 / 5e-23
AT5G36930 411 / 3e-124
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10004115
90 / 6e-23
AT5G27970 689 / 0.0
ARM repeat superfamily protein (.1.2)
Lus10020536
89 / 2e-22
AT1G27170 392 / 8e-118
transmembrane receptors;ATP binding (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.019G002500
61 / 7e-13
AT5G17680 587 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.012G053200
58 / 8e-12
AT1G27170 1187 / 0.0
transmembrane receptors;ATP binding (.1.2)
Potri.019G001602
56 / 6e-11
AT5G36930 778 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.001G307300
56 / 8e-11
AT5G17680 597 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.003G084333
55 / 1e-10
AT5G36930 347 / 2e-108
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.001G028750
54 / 2e-10
AT5G36930 516 / 3e-163
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.013G097300
54 / 2e-10
AT5G17680 598 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.T011750
54 / 3e-10
AT5G36930 551 / 2e-176
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.017G104301
54 / 3e-10
AT5G17680 566 / 1e-178
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.001G066500
52 / 1e-09
AT5G36930 462 / 2e-146
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF00931
NB-ARC
NB-ARC domain
Representative CDS sequence
>Lus10008414 pacid=23163507 polypeptide=Lus10008414 locus=Lus10008414.g ID=Lus10008414.BGIv1.0 annot-version=v1.0
ATGGGAGGCCTTGGCAAAACTACTTTAGCAAAGGCTGTTTACGACAAAGTATCTACACAATTCTATCGTTGCTGTTTTTTGCCAGACGTGAGAGAAACGA
TGTCGAAAAATGATGGGATGGTTACTCTACAGAATAAGATAATCTCAAACATTCTAAGGAACAAATCTGATGCAGAAAATGCCAGTGATGGAATGTGA
AA sequence
>Lus10008414 pacid=23163507 polypeptide=Lus10008414 locus=Lus10008414.g ID=Lus10008414.BGIv1.0 annot-version=v1.0
MGGLGKTTLAKAVYDKVSTQFYRCCFLPDVRETMSKNDGMVTLQNKIISNILRNKSDAENASDGM
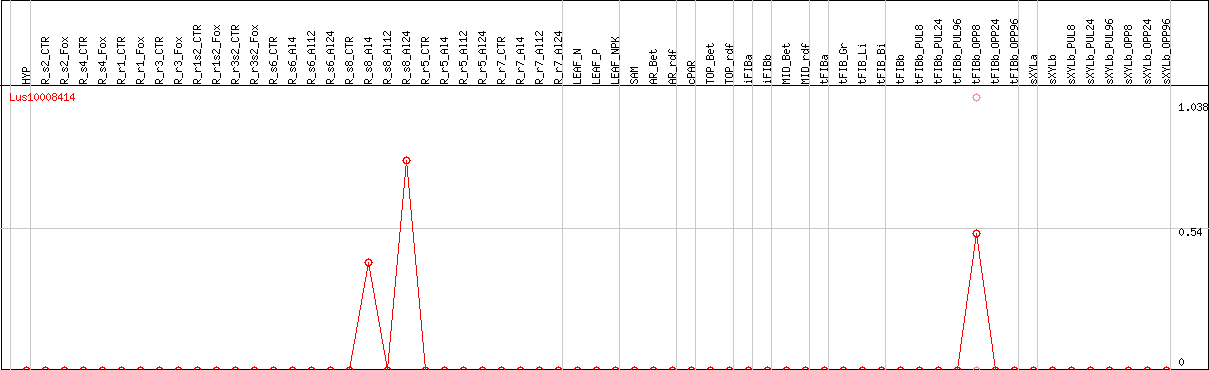
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10008414 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.