External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G14920 285 / 6e-95
GRAS
RGA2, GAI
RESTORATION ON GROWTH ON AMMONIA 2, GIBBERELLIC ACID INSENSITIVE, GRAS family transcription factor family protein (.1)
AT2G01570 281 / 6e-93
GRAS
RGA1
REPRESSOR OF GA1-3 1, REPRESSOR OF GA, GRAS family transcription factor family protein (.1)
AT1G66350 278 / 2e-92
GRAS
RGL1
RGA-like 1 (.1)
AT3G03450 278 / 6e-92
GRAS
RGL2
RGA-like 2 (.1)
AT5G17490 265 / 4e-87
GRAS
AtRGL3, RGL3
RGA-like protein 3 (.1)
AT5G48150 146 / 9e-42
GRAS
PAT1
phytochrome a signal transduction 1, GRAS family transcription factor (.1.2)
AT2G04890 140 / 6e-40
GRAS
SCL21
SCARECROW-like 21 (.1)
AT1G55580 129 / 8e-36
GRAS
SCL18, LAS
SCARECROW-LIKE 18, Lateral Suppressor, GRAS family transcription factor (.1)
AT1G50600 127 / 3e-34
GRAS
SCL5
scarecrow-like 5 (.1)
AT1G50420 125 / 5e-34
GRAS
SCL-3, SCL3
scarecrow-like 3 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.017G125200
331 / 1e-111
AT2G01570 701 / 0.0
REPRESSOR OF GA1-3 1, REPRESSOR OF GA, GRAS family transcription factor family protein (.1)
Potri.004G089800
326 / 7e-110
AT2G01570 709 / 0.0
REPRESSOR OF GA1-3 1, REPRESSOR OF GA, GRAS family transcription factor family protein (.1)
Potri.008G131700
303 / 5e-101
AT2G01570 781 / 0.0
REPRESSOR OF GA1-3 1, REPRESSOR OF GA, GRAS family transcription factor family protein (.1)
Potri.010G110700
301 / 3e-100
AT2G01570 764 / 0.0
REPRESSOR OF GA1-3 1, REPRESSOR OF GA, GRAS family transcription factor family protein (.1)
Potri.001G326000
167 / 1e-48
AT1G66350 296 / 2e-92
RGA-like 1 (.1)
Potri.005G095100
164 / 3e-48
AT1G66350 252 / 3e-77
RGA-like 1 (.1)
Potri.015G091200
159 / 2e-46
AT1G66350 256 / 4e-79
RGA-like 1 (.1)
Potri.012G093900
158 / 3e-46
AT1G66350 257 / 2e-79
RGA-like 1 (.1)
Potri.016G027800
149 / 1e-43
AT1G14920 244 / 1e-75
RESTORATION ON GROWTH ON AMMONIA 2, GIBBERELLIC ACID INSENSITIVE, GRAS family transcription factor family protein (.1)
Potri.001G361700
148 / 2e-42
AT1G50600 637 / 0.0
scarecrow-like 5 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF03514
GRAS
GRAS domain family
Representative CDS sequence
>Lus10009133 pacid=23150064 polypeptide=Lus10009133 locus=Lus10009133.g ID=Lus10009133.BGIv1.0 annot-version=v1.0
ATGCTAGACCTACGCCCCGCCGACGTGGAGGCGGTGGCGGTGAACTCTGTGTTCGAGCTACACCGTCTTCTCAGCCAATCGGCAGGGATAGATAAAGTAC
TGTCGTCGATCAAGGCGATGAAGCCAAAGATCGTGACGGTTGTGGAGCAAGAAGCCAACCACAACGGGCCGGTATTCGTGGACCGGTTTACGGAGGCTCT
GCATTACTACTCCAGCCTGTTCGACTCTCTGGAAGGGTCGTCCCCGAGCCAGGATCTGGTGATGTCGGAGCTCTACCTCGGGAGGCAGATTTGCAACGTG
GTGGCGTGCGAAGGAGGCGACCGAGTTGAACGGCACGAGACGTTGACTCAGTGGAGGGGCCGGATGGAATCGGCCGGATTTGACCCGGTTCACCTGGGGT
CGAATGCGTACAAGCAAGCTAGTACGTTACTTGCGCTGTTCGCCGGCGGGGATGGGTACAAGGTGGAGGAGAACGACGGTTCTCTGATGCTTGGGTGGCA
CACTAGACCTCTCATAGCCACGTCGGCATGGCAGCTCGCCAAGTGA
AA sequence
>Lus10009133 pacid=23150064 polypeptide=Lus10009133 locus=Lus10009133.g ID=Lus10009133.BGIv1.0 annot-version=v1.0
MLDLRPADVEAVAVNSVFELHRLLSQSAGIDKVLSSIKAMKPKIVTVVEQEANHNGPVFVDRFTEALHYYSSLFDSLEGSSPSQDLVMSELYLGRQICNV
VACEGGDRVERHETLTQWRGRMESAGFDPVHLGSNAYKQASTLLALFAGGDGYKVEENDGSLMLGWHTRPLIATSAWQLAK
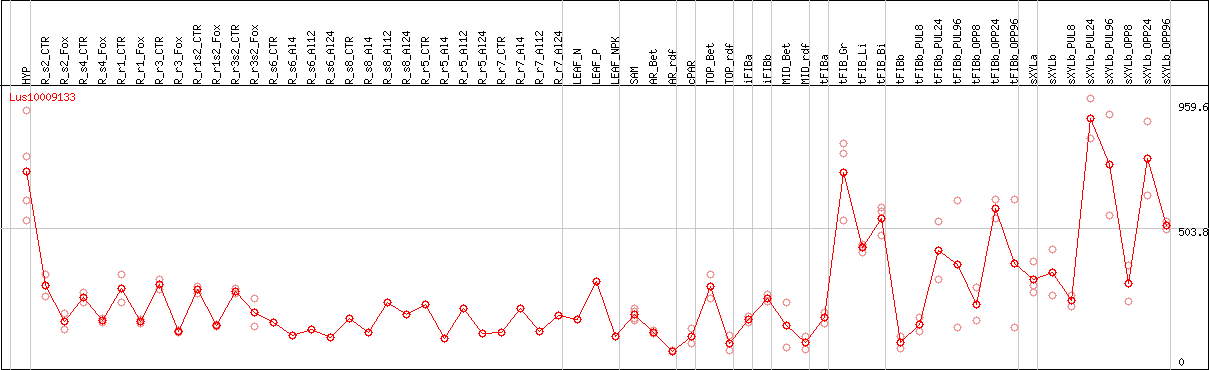
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10009133 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.