External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G11170 163 / 3e-45
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT3G44480 160 / 4e-44
COG1, RPP10, RPP1
recognition of peronospora parasitica 1, Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT1G69550 160 / 4e-44
disease resistance protein (TIR-NBS-LRR class) (.1)
AT3G44630 158 / 1e-43
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2), Disease resistance protein (TIR-NBS-LRR class) family (.3)
AT5G40910 156 / 9e-43
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT5G11250 155 / 2e-42
Disease resistance protein (TIR-NBS-LRR class) (.1)
AT5G51630 155 / 2e-42
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2), Disease resistance protein (TIR-NBS-LRR class) family (.3)
AT3G44670 154 / 3e-42
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
AT5G49140 154 / 4e-42
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT4G19510 152 / 1e-41
Disease resistance protein (TIR-NBS-LRR class) (.1), Disease resistance protein (TIR-NBS-LRR class) (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10017418
453 / 8e-153
AT5G44510 335 / 1e-96
target of AVRB operation1 (.1)
Lus10017419
437 / 1e-152
AT4G12010 272 / 9e-80
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Lus10024149
425 / 1e-149
AT4G19510 213 / 8e-61
Disease resistance protein (TIR-NBS-LRR class) (.1), Disease resistance protein (TIR-NBS-LRR class) (.2)
Lus10010221
438 / 1e-146
AT5G17680 369 / 4e-109
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10007030
367 / 2e-123
AT4G12010 316 / 3e-94
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Lus10015648
369 / 3e-119
AT4G12010 420 / 1e-126
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Lus10015650
329 / 5e-107
AT4G12010 346 / 3e-103
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Lus10042777
288 / 3e-98
AT3G44630 136 / 2e-36
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2), Disease resistance protein (TIR-NBS-LRR class) family (.3)
Lus10018021
273 / 1e-83
AT4G12010 434 / 5e-131
Disease resistance protein (TIR-NBS-LRR class) family (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0173
STIR
PF01582
TIR
TIR domain
CL0023
P-loop_NTPase
PF00931
NB-ARC
NB-ARC domain
Representative CDS sequence
>Lus10010222 pacid=23154236 polypeptide=Lus10010222 locus=Lus10010222.g ID=Lus10010222.BGIv1.0 annot-version=v1.0
ATGGAAATCAGTCCCTTCCCTGAGTTCATGGCTGCCGCTTCTTATTCTTCTTCTTCTGCATCTCATAGCTCTCTGTACGATTTCTTATGTTTTCGAGGTG
ACGACACGCGCCATGGTTTCACCAGCCACCTCATGACTGCTCTGTGTGATAGGCAAATCCAAACATTTATAGACGACAAGCTCGAGAAAACAGAGAGCAT
CGGTGAGCTTCAATCCATCCTTGAAAGAACTGCTCTTTCCGTAGTGGTTTTCTCTGAGAGATTTGCGGATTCGGTCTGGTGCTTGGATGAGGTAGCCATC
ATAGCCAGACAGATGAAGGAATCCGGGTATTGGGTTCTACCGGTTTTCTACAAAGTGGATCCATCAGACGTGACAGACATTACTGGAAGGTACACAACTA
CCATCGATCGTGAACATAAGGGCAGAAGTAGTTATGTAGAGGATAAGAAGAGATGGATGGATGCTTTGAATGTTGTGGCTGATTCTGCTGGCCATACCTC
ACAGACCTTCAGAAATGAATCTGAATTGATTAAGGCGATTGTGAAAGATGTTCACAAAAAACTAATTGACATGTCACCAGTTATCAAGAATGATAACTTG
GTTGCTATGAGTTCACGTATTTCCGAGGTTGAGCGACTATTGGCTATGGATAAAGTAGACGAGACTCGCATCGTTGGGCTATGGGGAATGGGCGGTGTTG
GGAAAACAACTCTAGCAAGAGCATGTTATGAAAGGGTAACTTCGTCCAATAAAGGAATCAAACATCTTTTCATCTCAGACGTCAATGAAAAATATGGAAA
GCATCACGACATAGAAGAAGTTGTTTGTAAATACTATTAA
AA sequence
>Lus10010222 pacid=23154236 polypeptide=Lus10010222 locus=Lus10010222.g ID=Lus10010222.BGIv1.0 annot-version=v1.0
MEISPFPEFMAAASYSSSSASHSSLYDFLCFRGDDTRHGFTSHLMTALCDRQIQTFIDDKLEKTESIGELQSILERTALSVVVFSERFADSVWCLDEVAI
IARQMKESGYWVLPVFYKVDPSDVTDITGRYTTTIDREHKGRSSYVEDKKRWMDALNVVADSAGHTSQTFRNESELIKAIVKDVHKKLIDMSPVIKNDNL
VAMSSRISEVERLLAMDKVDETRIVGLWGMGGVGKTTLARACYERVTSSNKGIKHLFISDVNEKYGKHHDIEEVVCKYY
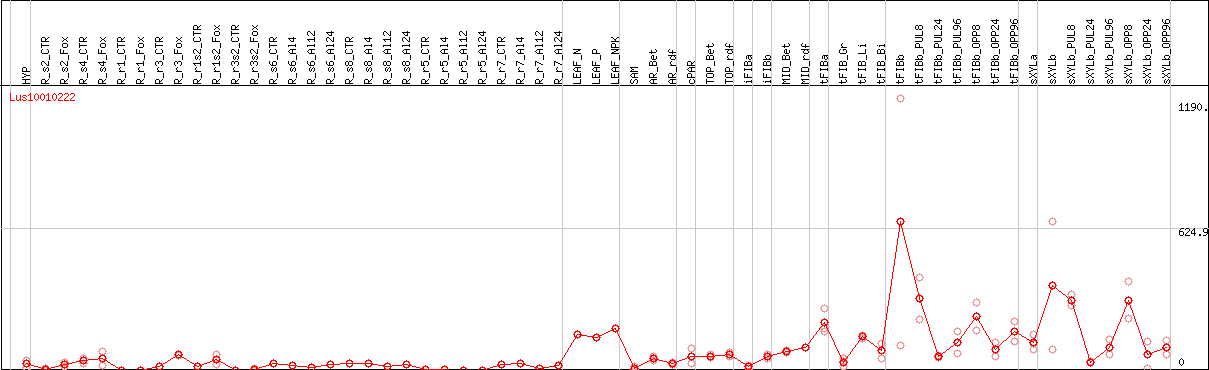
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10010222 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.