Lus10011038 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10011038 pacid=23148691 polypeptide=Lus10011038 locus=Lus10011038.g ID=Lus10011038.BGIv1.0 annot-version=v1.0
ATGGCGGATTCGGTGGAGCTCCCCAGTCGGTTGGCGATCCTGCCTTTCAGGAACAAGGTCCTCTTGCCCGGCGCCATCATCCGCATCCGCTGCACTTCTC
CCAGCAGTGTGAAATTGGTGGAGCAAGAGCTATGGCAGAGAGAGGAGAAGGGTATGATTGGCATTTTGCCTGTTCAGGACACTGCTGAGGCGTCCGTCGG
GTCAATGCTGTCTCAAGGAGTTGGAAGCGAGTCAGGGGGTGATAGAAGCTCGAAAGCTCAAGTGGGAGCCACCTCGGAGAGTCTCAAAGTAATTGATGGT
AAAAACCAGCAGGAAGTTATTCGGTGGCACAATAGGGGAGTTGCTGCTCGTGCTTTGCATTTGTCGAGAGGGGTGGAGAAGCCGAGTGGGAGAGTGACTT
ACATTGTTGTGCTTGAAGGTTTATGCAGATTCAGTGTTCAGGAACTTAGTAAGCGAGGGATGTATTACACTGCCCGGATATCTCCGGCTGAGATGACAAA
GGCTGAGATGGAACAAGTGGAGCATGATCCTGACTTCGTCACACTGTCTCGACAGTTCAAAGCAACTGCCATGGAACTTATTTCAATTCTTGAGCAGAAA
CAGAAACCCGGTGGCCGGACCAAAGCTCTTTTGGAGACGGTTCCTGTTCACAAGTTGGCAGATATATTTGTTGCCAGCTTTGAGATAAGTTTTGAAGAGC
AGTTGTGTATGTTGGATTCTGTTGATCTGAAAGTAAGGCTTTCGAAAGCTACTGAGCTGGTTGACAGGCATTTACAGTCGATTCGTGTAGCAGAAAAAAT
TACCCAAAAGGTTGAGGGACAGTTATCAAAGTCGCAGAAAGAGTTTCTTCTACGTCAGCAGATGCGTGCTATAAAAGAGGAGCTCGGTGACAATGATGAT
GATGAGGATGATGTCGCTGCACTAGAAAGAAAGATGCAAAGTGCAGGAATGCCTTCAAATGTCTGGAAACATGCACAGAGGGAATTGCGGCGGCTCAAGA
AGATGCAACCACAACAACCCGGATATAACAGTTCACGAGTGTATTTGGAGCTGCTTGCTGAACTTCCATGGCAGAATGCTAGTGAAGTAGCTGAACTGGA
TATAAAAGCTGCGAAAGAACGTCTGGATAATGACCACTATGGTCTTGTGAAGGTCAAGCAACGGATTATAGAATATCTAGCAGTGCGCAAGCTTAAACCG
GATGCACGAGGTCCAGTATTGTGTTTTGTCGGTCCACCAGGCGTCGGGAAGACATCTTTAGCATCATCTATTGCTGCTGCCTTAGGCCGGAAATTCATTC
GTATCTCTCTTGGGGGTGTAAAAGACGAGGCTGATATTAGAGGGCACAGAAGAACTTACATTGGGAGCATGCCAGGGCGTCTTATTGATGGATTAAAGAG
AGTCGGTGTTTGCAATCCGGTCATGCTTCTGGACGAGATTGACAAGACTGGTTCTGATGTGCGTGGCGATCCTGCATCAGCTCTGCTCGAAGTTCTGGAT
CCTGAACAGAACGGGACATTTAATGATCACTATTTGAATGTTCCTTTCGACCTGTCTAAGGTGATTTTCGTGGCAACAGCAAATCGGGCGCAACCTATTC
CAGCTCCACTGCTGGACCGTATGGAAGTTATTGAGCTTCCGGGTTACACTCCTGAAGAAAAGCTCAGAATCGCCATGCGACATCTGATACCTCGAGTGTT
GGACCAACATGGTTTGACCTCCGATTTCTTACAAATTCCAGAGGATTCAGTAAAAATTGTAATCCAGAGGTACACCAGGGAGGCCGGTGTTCGGAACCTA
GAGAGACACCTGGCTGCCTTGGCTCGTGCAGCAGCTGTCAGACTTGCAGAGCAAGACCAAACAGTGCCGCTAAGCAAGGATGTTCAACGGCTTGCCACAC
CATTGCTGGAGAACAGACTTTCCGATGGAGCTGAAATGGAAATGGAAGTAATACCAATTGGCGAAAACGGTCATGAGATCTCCAACGGTTTCAGAGTTGC
CTCATTGGTTGTGGATGAGGCCATGCTCGAGAAAGTTCTTGGGCCTCCAAGATTCGACGACAAAGAAGCTTCAGAACGTGTCGCAAGTCCAGGTATCACT
GTTGGGCTTGTTTGGACCGCATTCGGTGGGGAAGTCCAGTTCGTGGAGGCCACATCAATGGCTGGAAAGGGTGAGTTGCATCTCACCGGACAACTTGGAG
ACGTCATCAAGGAATCTGCACAGATAGCATTGACTTGGGTAAGAGCCAGGGCAACTAATCTCAAGTCGACGGATGGTCATGAAATTAGTCTTCTTGAAGG
CAAGGATATCCACATACATTTCCCTGCGGGTGCCGTACCTAAGGACGGGCCTTCAGCGGGTGTGACACTCGTGACCACTCTTGTTTCGTTGTTCAGTCAG
CAGCGAGTACGAGCTGATACAGCAATGACCGGAGAAATGACTTTGAGAGGCCTCGTGCTTCCTGTTGGTGGGATTAAGGATAAGATATTGGCTGCTCATC
GTTACGGAATCAAGCGAGTGATTCTACCGGAGCGAAACCTCAAGGACTTGGTGGAAGTACCGGCTGCTGTCCTTGCCAATATAGAGTCGTACTCTCCATC
TCTCAGAGCTATCGCCGTCTCTTCCTTCTCCGCCGTCCAGCTTTCGCCGCCGACGAAGCAACTCCGAATCATGTTGAAGCATGCATCGCGTCTGGTATCC
AGATCCGTCGCTAGGCCGAAGCTCGTTCGAGCATTCTCCAGTGATGTCCCAGCTACGCCTACAGAGGACTCGGCATTCACCGCTTCGTGGAAGAAAGTGA
TCCCGACGATCGATCCGCCGAAGACTCCTTCAGCTTTCATCAAGCCTCGCCCGACGACGCCGTCGACCATCCCCGCTAAGCTCACTGTCAATTTCGTCCT
TCCTTATTCCACCCAGCTCGCGGAAAAAGAGGTCGATATGATAATCGTCCCAGCAACAACAGGGCAGATGGGTATCCTACCGGGCCATGTCCCAACCATA
GCTGAGCTGAAACCGGGAGTGATGTCTGTGCACGAAGGAAGCGAGGTCTCCAAGTACTTTGTCAGCAGCGGGTTTGCCTTCATCCACGGGAACTCAGTTG
CTGATGTGATTGCAGTCGAGGCTGTGCCTGTCGATCAAATCGATGCAAACTTGGTCCAGAAAGGGATAACCGAGTTCAACCAGAAGCTCGGTTCCGCCTC
AACTGATCTCGAAAAGGCCGAGGCTCAGATTGGCGTGGACGTTTACAGTGCTCTCAATGCTGCTCTCACAGGCTAA
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10011038 pacid=23148691 polypeptide=Lus10011038 locus=Lus10011038.g ID=Lus10011038.BGIv1.0 annot-version=v1.0
MADSVELPSRLAILPFRNKVLLPGAIIRIRCTSPSSVKLVEQELWQREEKGMIGILPVQDTAEASVGSMLSQGVGSESGGDRSSKAQVGATSESLKVIDG
KNQQEVIRWHNRGVAARALHLSRGVEKPSGRVTYIVVLEGLCRFSVQELSKRGMYYTARISPAEMTKAEMEQVEHDPDFVTLSRQFKATAMELISILEQK
QKPGGRTKALLETVPVHKLADIFVASFEISFEEQLCMLDSVDLKVRLSKATELVDRHLQSIRVAEKITQKVEGQLSKSQKEFLLRQQMRAIKEELGDNDD
DEDDVAALERKMQSAGMPSNVWKHAQRELRRLKKMQPQQPGYNSSRVYLELLAELPWQNASEVAELDIKAAKERLDNDHYGLVKVKQRIIEYLAVRKLKP
DARGPVLCFVGPPGVGKTSLASSIAAALGRKFIRISLGGVKDEADIRGHRRTYIGSMPGRLIDGLKRVGVCNPVMLLDEIDKTGSDVRGDPASALLEVLD
PEQNGTFNDHYLNVPFDLSKVIFVATANRAQPIPAPLLDRMEVIELPGYTPEEKLRIAMRHLIPRVLDQHGLTSDFLQIPEDSVKIVIQRYTREAGVRNL
ERHLAALARAAAVRLAEQDQTVPLSKDVQRLATPLLENRLSDGAEMEMEVIPIGENGHEISNGFRVASLVVDEAMLEKVLGPPRFDDKEASERVASPGIT
VGLVWTAFGGEVQFVEATSMAGKGELHLTGQLGDVIKESAQIALTWVRARATNLKSTDGHEISLLEGKDIHIHFPAGAVPKDGPSAGVTLVTTLVSLFSQ
QRVRADTAMTGEMTLRGLVLPVGGIKDKILAAHRYGIKRVILPERNLKDLVEVPAAVLANIESYSPSLRAIAVSSFSAVQLSPPTKQLRIMLKHASRLVS
RSVARPKLVRAFSSDVPATPTEDSAFTASWKKVIPTIDPPKTPSAFIKPRPTTPSTIPAKLTVNFVLPYSTQLAEKEVDMIIVPATTGQMGILPGHVPTI
AELKPGVMSVHEGSEVSKYFVSSGFAFIHGNSVADVIAVEAVPVDQIDANLVQKGITEFNQKLGSASTDLEKAEAQIGVDVYSALNAALTG
|
DESeq2's median of ratios [FLAX]
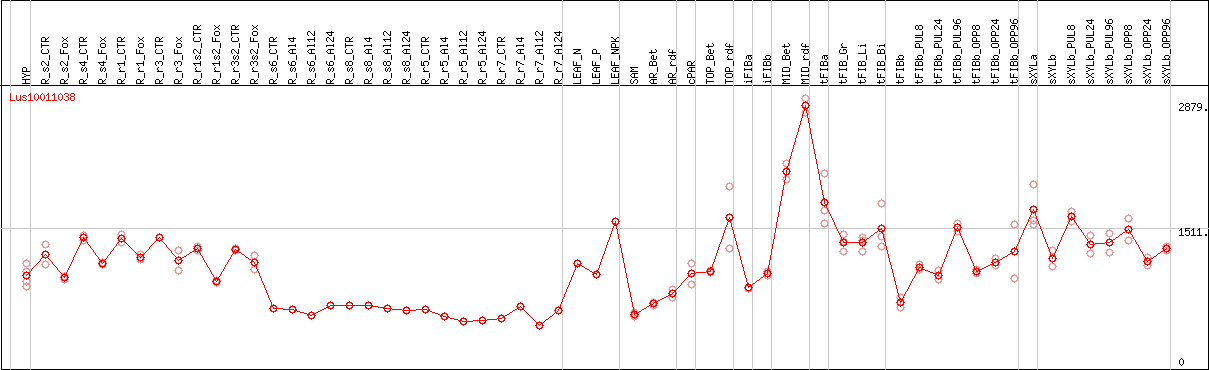
Coexpressed genes
Lus10011038 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
