External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G42630 131 / 1e-36
GARP
KAN4, KANADI4, ATS
KANADI 4, ABERRANT TESTA SHAPE, Homeodomain-like superfamily protein (.1.2)
AT4G17695 119 / 2e-31
GARP
KAN3, KANADI3
KANADI 3, Homeodomain-like superfamily protein (.1)
AT1G32240 115 / 9e-30
GARP
KAN2, KANADI2
KANADI 2, Homeodomain-like superfamily protein (.1)
AT5G16560 111 / 4e-28
GARP
KAN1, KAN
KANADI 1, KANADI, Homeodomain-like superfamily protein (.1)
AT2G42660 81 / 6e-18
GARP
Homeodomain-like superfamily protein (.1)
AT2G38300 80 / 5e-17
GARP
myb-like HTH transcriptional regulator family protein (.1)
AT2G02060 79 / 5e-17
GARP
Homeodomain-like superfamily protein (.1)
AT1G14600 78 / 7e-17
GARP
Homeodomain-like superfamily protein (.1)
AT2G40260 79 / 1e-16
GARP
Homeodomain-like superfamily protein (.1)
AT4G28610 77 / 9e-16
GARP
ATPHR1, PHR1
phosphate starvation response 1 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10032746
296 / 4e-100
AT5G42630 194 / 3e-60
KANADI 4, ABERRANT TESTA SHAPE, Homeodomain-like superfamily protein (.1.2)
Lus10035387
112 / 3e-30
AT1G32240 222 / 1e-71
KANADI 2, Homeodomain-like superfamily protein (.1)
Lus10030989
114 / 8e-29
AT1G32240 229 / 6e-71
KANADI 2, Homeodomain-like superfamily protein (.1)
Lus10002207
82 / 9e-18
AT2G38300 171 / 9e-51
myb-like HTH transcriptional regulator family protein (.1)
Lus10012239
82 / 1e-17
AT2G38300 170 / 3e-50
myb-like HTH transcriptional regulator family protein (.1)
Lus10040945
81 / 2e-17
AT2G40260 159 / 2e-45
Homeodomain-like superfamily protein (.1)
Lus10007513
81 / 2e-17
AT2G40260 145 / 3e-40
Homeodomain-like superfamily protein (.1)
Lus10029225
78 / 3e-17
AT2G38300 110 / 5e-30
myb-like HTH transcriptional regulator family protein (.1)
Lus10008197
81 / 5e-17
AT5G29000 318 / 3e-105
PHR1-like 1, Homeodomain-like superfamily protein (.1.2.3.4)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.014G037200
139 / 4e-39
AT5G42630 209 / 6e-66
KANADI 4, ABERRANT TESTA SHAPE, Homeodomain-like superfamily protein (.1.2)
Potri.002G130200
132 / 3e-36
AT5G42630 199 / 6e-62
KANADI 4, ABERRANT TESTA SHAPE, Homeodomain-like superfamily protein (.1.2)
Potri.017G137600
117 / 8e-30
AT5G16560 256 / 8e-81
KANADI 1, KANADI, Homeodomain-like superfamily protein (.1)
Potri.003G096300
115 / 8e-30
AT1G32240 224 / 1e-69
KANADI 2, Homeodomain-like superfamily protein (.1)
Potri.004G082400
114 / 5e-29
AT5G16560 233 / 7e-72
KANADI 1, KANADI, Homeodomain-like superfamily protein (.1)
Potri.012G042100
112 / 3e-28
AT5G16560 193 / 6e-57
KANADI 1, KANADI, Homeodomain-like superfamily protein (.1)
Potri.015G031600
111 / 6e-28
AT5G16560 176 / 1e-50
KANADI 1, KANADI, Homeodomain-like superfamily protein (.1)
Potri.001G137600
107 / 1e-26
AT1G32240 216 / 6e-67
KANADI 2, Homeodomain-like superfamily protein (.1)
Potri.008G071700
87 / 2e-19
AT2G40260 166 / 2e-47
Homeodomain-like superfamily protein (.1)
Potri.009G075100
85 / 7e-19
AT2G38300 162 / 4e-47
myb-like HTH transcriptional regulator family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0123
HTH
PF00249
Myb_DNA-binding
Myb-like DNA-binding domain
Representative CDS sequence
>Lus10011660 pacid=23149664 polypeptide=Lus10011660 locus=Lus10011660.g ID=Lus10011660.BGIv1.0 annot-version=v1.0
ATGAGGTGGACTACCACTCTCCATGCTCACTTTGTTCACGCCGTTCAGCTTCTTGGCGGCCATGAAAGAGCGACGCCCAAATCGGTTCTGGAGCTGATGA
ACGTCAAAGATCTAACCCTTGCTCATGTCAAGAGCCATTTGCAGATGTATAGAACAGTGAAGAGCACTGACAAATCACCATTACCACCTGGTGGAGGAGG
GCATGGATTGAGTCATAGTAGTGATGATGACAACATTAATAATTATGGAGAGGATGCCATTATTGATTTGGACGCAGCAAAAGAGGAAGAAGAAGAAGAA
GAAGGAGATGGATACAGAAAATTATTAGCTGCTGATCATCAAACAGATTCTCCTCCTCAACCAACCATTACTCAAAATGTTCCGCGGTCATCATCCATTG
ACAGAGACAGCAACAACATTAGATTATCACATCCATTGACTCATTCTCACAATAACGGGCATGACATAACTACAAAGGAGGAGAAGGGAGGAGAGCTGAG
GTTGACAAGGACTTCAGAGATGTTGGTCAACCTAGAATTCACACTTGGAAGGCCAAGCTGGCAGAGGGATGACGTCAACTCTGTAGCTGAACCGTCCAAT
TATGACTTGACCCTCCTCAAGTACCAAGTAAGCCCAAGGCGAGTCGACTTCACCAAAGCCCATCTCAGGCCTGCTCAACCTCATCAGCAACCACACCTGT
CGCAGCCCGTTAACAAGCATGTCGAACCTAGCAATTACGCAGCAGCTTGGCAATCGAGGAAGCTTCGTGACGAGAAGATCAGAGATCGGAGGCAGTGCCT
ATGCGAGAAGATCGATTGCCAGCAATGA
AA sequence
>Lus10011660 pacid=23149664 polypeptide=Lus10011660 locus=Lus10011660.g ID=Lus10011660.BGIv1.0 annot-version=v1.0
MRWTTTLHAHFVHAVQLLGGHERATPKSVLELMNVKDLTLAHVKSHLQMYRTVKSTDKSPLPPGGGGHGLSHSSDDDNINNYGEDAIIDLDAAKEEEEEE
EGDGYRKLLAADHQTDSPPQPTITQNVPRSSSIDRDSNNIRLSHPLTHSHNNGHDITTKEEKGGELRLTRTSEMLVNLEFTLGRPSWQRDDVNSVAEPSN
YDLTLLKYQVSPRRVDFTKAHLRPAQPHQQPHLSQPVNKHVEPSNYAAAWQSRKLRDEKIRDRRQCLCEKIDCQQ
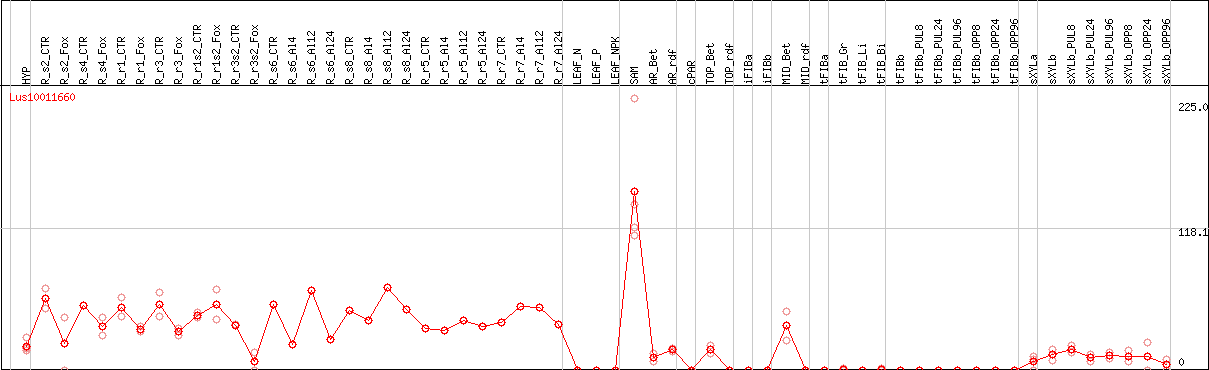
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10011660 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.