Lus10013372 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10013372 pacid=23151355 polypeptide=Lus10013372 locus=Lus10013372.g ID=Lus10013372.BGIv1.0 annot-version=v1.0
ATGAAGCCACATGATAGCGGAAATGGCAGTACTAAAAAGCCAGCCACAACTACTGCTGACAATAAACCGAAACCGCCACCTCAATTGGAGCCAGAACCTG
AACCTCCTGTGCCGGAGAGTGTAGGTAGAAGGAAAACTAGCAAGTACTTTGCAACGGATAAGCAGAAGCCGAAAGATGAAAAAGAGGCGGTGAAACCCCC
AGCCAAGAGAAAGAGTCAGAGTGAGAGTAACCCACCTGTGAAAGCAACGCCTCCTAGAAAGGTTCAGAAAGTGGAAGATGATGATGACGGTGATGACTTT
GTCATGTCAGCTAAAAAGAAGATCTCTGTTGATGCCACTCCTAGCAAGAAGTTGAAGAGTGGTTCTGGCAGGGGGATTAGTCAGAAGGGTGTCGTAAAAG
ATGAGAGTGAGGAAGAAGAAGCTGAGGAAACTGAACCACCTCCTAAGGCTGCTGGGAGAGGTCGTGGCAGTAGAGGTGCAGCTGGGGCTTCAGCTGGTGG
AAGGGGCAGAGGCGGAGGCCGTGGTGGATTCATGAACTTTGGTGAAAGGAAAGATCCTCCGCACAAAGGAGAAAAGGAAGTCCCAGAGGGTGCTCCTGAC
TGCTTAGCTGGGTTGACATTTGTAATAAGTGGCACTCTTGATAGATCCTCTATGATCGTCTTCGTTTCCTTTCTGGTTCTCATGCTTGTCTTTTATGCCG
TGAATTTTTTTCGGCCTTACAGTTTGGAAAGAGAAGAAGCAGAAGATTTGATTAAACGACATGGAGGTCGTATTACTGGATCTGTCAGCAAGAAAACGAG
CTACCTTTTATGCGATGAGGATATTGAGGGGCGAAAATCTGCCAAAGCTAAAGAGCTTGGCACTGCTTTCCTTACCGAGGATGGTCTTTTTGATAAGATT
CGCTCTTCAAAACCTGCAAAAGTGTCCCCAAAAGAAGTGCCTAAGGTCTCTGCGGAGAAAGTATCACTAGTGCCAAAAAAGAGTCCCCAAAAAGCAGAAG
TTAAGGGGACCTCTTCAGTACAAGGTGCTACAAAAAAAGGTCCAGCCAAAGCTGTCACCCCTGTTAAGAGAAAGCAATCCATTGAGCCCTCCTCATTGCC
ATGGACAGAAAAATATAGGCCTAAAGTTCCAAATGATCTTGTTGGGAATCAATCATTGGTTAACCAGCTTCATAATTGGTTAAAAAGCTGGAATGACCAG
TTCCTTGGTGGCGGGAAAAAAGGAAAGGGTAAACAGAGCGATTCTGCTGCCAAGAAAGCAGTGTTACTAAGTGGAACACCAGGGATAGGGAAAACAACAT
CTGCAAAGCTGGCGTGCCAGATGCTTGGTTTTCAAGCAGTAGAGGTCAATGCAAGTGACACACGAGGGAAGGCAGATTCCAAAATTTTCAAAGGAATTAG
TGGCAGTAATGCAAATTCCATAAAGGAACTCATCAACAACGAATCTCTGAGTTCTAAAATGGACAGGCCAAAGCATCCAAAGACAGTTCTCATCATGGAT
GAGGTAGATGGGATGTCTGCAGGAGATAGAGGTGGAGTTGCTGATCTTATAGCTAGCATAAAGATCTCAAAAATGCCTATTGTATGCATCTGTAATGATC
GATACAGTCAGAAATTAAAGAGTCTTCTCAATTACTGCCTGGTTCTGACCTTCCGCAAACCTACAAAACAACAGATGGCAAAGAGGCTAATGCAAATCGC
ATATGCTGAAGGACTCAAAGTTAATGAGATTGCACTTGAGGAACTTGCAGAAAGATCGAATGGGGACATGCGGATGGCCGTAAATCAATTGCAGTACATG
AGTCTATCCATGTCGACAATTCAGTATGATGATGTGCGGAAGCGCCTTCTAAGCAGTGCAAAGGATGAAGATATTTCACCATTTACAGCAGTCGATAAGC
TGTTTAACTTCAATTCATCGAAGCTGCGAATGGATCAGCGGGTTGACCTCAGCATGAGTGATCCAGACTTGGTCCCTCTTATAATTCAGGAAAACTATAT
TAACTATAGACCAAGTTTGGCTGGCAAGGATGAGAATGGGATAAAACGAATGAACTTGCTTTCCCGTGCTGCAGACTCTATTGCAGATGGAGATTTAGTA
AATGTACAAATCCGAAGGCATAGGCAGTGGCAGCTCTTTCAGGCTGGTTCTCTAGCATCCTGCATAATTCCTGCTGCATTGGTACATGGACAGAGAGAGA
TCATGGAACCTGTGAGTCTCTCTCTTATCGTGATGGCCTTTCATCTTTATTCTGCTATTCCATACAGGGGGGAGGGCGAAAGAAATTTTAACAGGTTTGG
TGGGTGGCTAGGAAAGAACTCAACGATGGGAAAAAATATGAGACTGTTGGAGGATCTCCATGTTCATCTTCTTGCTTCTCGAGAATCTAATATGGGGAGG
GGGACTCTTCGACTTGATTATTTTACTGTTCTTCTAAAACAATTGACAGATCCTCTGCGCACGATGTCCAAGGATGAAGCCGTGGAGAAGGTCGTGGAGT
TCATGGATGTATACTCAATAAGTCAGGAAGACATGGAGACCATTGTCGAGTTGGCTAAATTTAAGGGGCACCCAAATCCACTGGAGGGTGTACCAGCAGC
TGTCAAATCAGCTTTGACTAGAGCGTACAAAGATGCAACCAAATCACGATTGGCACGAGCTGCAGATTTAGTCCCGATTTCCGGGATGAAAAAGGCCCCT
AAGAAGCGAATAGCCGCAATGTTGGAACCTCCTAGTGAAGGTTTAGGGGAGGAAACTGTTGATCCAGTTGAAGAGAGTGAAGATGAGAACTCTTCTGACA
CAGAGGATCCGGAAAGCACTCTCGACGGGGAGAAACTGCAGTCTAGCCTACAAAGTTTGAACTCCAAAGGGATAAAAGTTGAGATGGAATTGAAGAGTAG
TACCCAAAATGCGAAGGGGAAGAAGGCACCGGGAGGTCGAGGTAGAGGCGGGGCGTCCAGTTCTGCAGAAAAGAAACCGGCAGGAAGGGGAAGAGGTGGC
AGCAGTTCTGCTGGAGCTAAGAGAAAGAGATGA
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10013372 pacid=23151355 polypeptide=Lus10013372 locus=Lus10013372.g ID=Lus10013372.BGIv1.0 annot-version=v1.0
MKPHDSGNGSTKKPATTTADNKPKPPPQLEPEPEPPVPESVGRRKTSKYFATDKQKPKDEKEAVKPPAKRKSQSESNPPVKATPPRKVQKVEDDDDGDDF
VMSAKKKISVDATPSKKLKSGSGRGISQKGVVKDESEEEEAEETEPPPKAAGRGRGSRGAAGASAGGRGRGGGRGGFMNFGERKDPPHKGEKEVPEGAPD
CLAGLTFVISGTLDRSSMIVFVSFLVLMLVFYAVNFFRPYSLEREEAEDLIKRHGGRITGSVSKKTSYLLCDEDIEGRKSAKAKELGTAFLTEDGLFDKI
RSSKPAKVSPKEVPKVSAEKVSLVPKKSPQKAEVKGTSSVQGATKKGPAKAVTPVKRKQSIEPSSLPWTEKYRPKVPNDLVGNQSLVNQLHNWLKSWNDQ
FLGGGKKGKGKQSDSAAKKAVLLSGTPGIGKTTSAKLACQMLGFQAVEVNASDTRGKADSKIFKGISGSNANSIKELINNESLSSKMDRPKHPKTVLIMD
EVDGMSAGDRGGVADLIASIKISKMPIVCICNDRYSQKLKSLLNYCLVLTFRKPTKQQMAKRLMQIAYAEGLKVNEIALEELAERSNGDMRMAVNQLQYM
SLSMSTIQYDDVRKRLLSSAKDEDISPFTAVDKLFNFNSSKLRMDQRVDLSMSDPDLVPLIIQENYINYRPSLAGKDENGIKRMNLLSRAADSIADGDLV
NVQIRRHRQWQLFQAGSLASCIIPAALVHGQREIMEPVSLSLIVMAFHLYSAIPYRGEGERNFNRFGGWLGKNSTMGKNMRLLEDLHVHLLASRESNMGR
GTLRLDYFTVLLKQLTDPLRTMSKDEAVEKVVEFMDVYSISQEDMETIVELAKFKGHPNPLEGVPAAVKSALTRAYKDATKSRLARAADLVPISGMKKAP
KKRIAAMLEPPSEGLGEETVDPVEESEDENSSDTEDPESTLDGEKLQSSLQSLNSKGIKVEMELKSSTQNAKGKKAPGGRGRGGASSSAEKKPAGRGRGG
SSSAGAKRKR
|
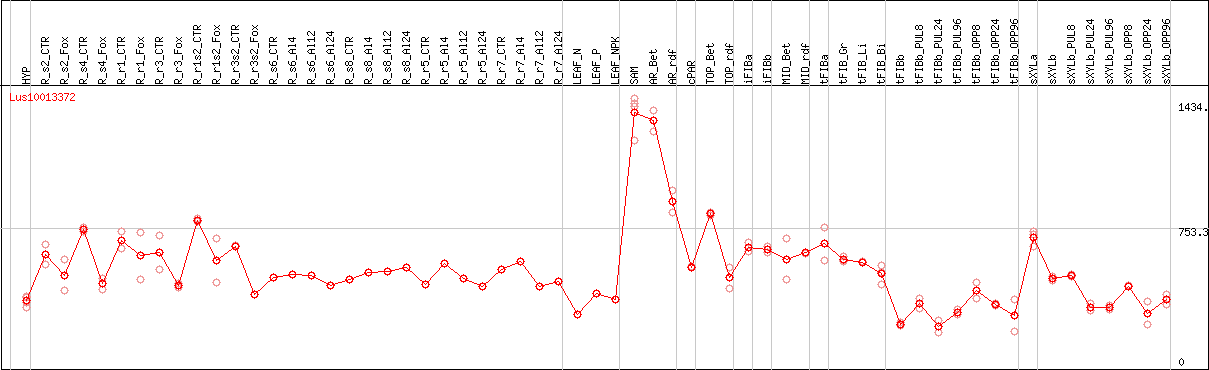
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10013372 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
