External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G51680 89 / 8e-22
AtSDR2
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G29250 82 / 2e-19
AtSDR4
short-chain dehydrogenase reductase 4, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT3G26770 82 / 3e-19
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G47140 80 / 8e-19
AtSDR5
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G29260 79 / 1e-18
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G47120 77 / 2e-17
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G47130 74 / 2e-16
AtSDR3
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT4G03140 74 / 4e-16
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G26760 73 / 5e-16
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT1G52340 70 / 9e-15
SIS4, SDR1, ISI4, GIN1, ATABA2, SRE1, ABA2, ATSDR1
SHORT-CHAIN DEHYDROGENASE REDUCTASE 1, IMPAIRED SUCROSE INDUCTION 4, GLUCOSE INSENSITIVE 1, SHORT-CHAIN DEHYDROGENASE/REDUCTASE 1, ARABIDOPSIS THALIANA ABA DEFICIENT 2, ABA DEFICIENT 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10021258
225 / 5e-75
AT2G47140 177 / 2e-54
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10021257
131 / 2e-38
AT2G47140 222 / 3e-72
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10013595
125 / 5e-36
AT2G47140 224 / 3e-73
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10016178
120 / 7e-34
AT2G47140 210 / 2e-67
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10022562
117 / 5e-33
AT3G29250 223 / 5e-72
short-chain dehydrogenase reductase 4, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10029369
117 / 1e-32
AT3G51680 226 / 3e-73
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10016996
115 / 4e-32
AT3G51680 211 / 3e-67
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10016177
114 / 1e-31
AT3G26770 219 / 1e-70
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10010544
112 / 5e-31
AT3G51680 216 / 2e-69
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G206500
122 / 1e-34
AT1G52340 246 / 4e-81
SHORT-CHAIN DEHYDROGENASE REDUCTASE 1, IMPAIRED SUCROSE INDUCTION 4, GLUCOSE INSENSITIVE 1, SHORT-CHAIN DEHYDROGENASE/REDUCTASE 1, ARABIDOPSIS THALIANA ABA DEFICIENT 2, ABA DEFICIENT 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.004G199900
117 / 5e-33
AT2G47140 256 / 1e-85
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.004G200100
116 / 1e-32
AT3G26770 257 / 2e-85
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.006G206700
110 / 5e-30
AT3G51680 236 / 4e-77
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.006G206400
109 / 7e-30
AT3G51680 245 / 2e-80
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.016G073900
109 / 1e-29
AT3G51680 252 / 2e-83
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.005G106100
107 / 6e-29
AT2G47140 234 / 9e-77
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.006G207100
105 / 2e-28
AT4G03140 241 / 2e-78
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.006G206800
103 / 8e-28
AT1G52340 246 / 5e-81
SHORT-CHAIN DEHYDROGENASE REDUCTASE 1, IMPAIRED SUCROSE INDUCTION 4, GLUCOSE INSENSITIVE 1, SHORT-CHAIN DEHYDROGENASE/REDUCTASE 1, ARABIDOPSIS THALIANA ABA DEFICIENT 2, ABA DEFICIENT 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.016G073800
100 / 3e-26
AT1G52340 250 / 6e-83
SHORT-CHAIN DEHYDROGENASE REDUCTASE 1, IMPAIRED SUCROSE INDUCTION 4, GLUCOSE INSENSITIVE 1, SHORT-CHAIN DEHYDROGENASE/REDUCTASE 1, ARABIDOPSIS THALIANA ABA DEFICIENT 2, ABA DEFICIENT 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF00106
adh_short
short chain dehydrogenase
Representative CDS sequence
>Lus10013594 pacid=23168751 polypeptide=Lus10013594 locus=Lus10013594.g ID=Lus10013594.BGIv1.0 annot-version=v1.0
ATGAAAGGATCTGCTCTGCTGCTCAGTCCCATTGCTCAGAGGTTAAAAGACAAGGTCGCCGTGATAACCGGCGGAGCCAGTGGAATCGGAGCCTCGACCG
CCAGGCATTTTGCCAAGCACGGCGCCAAGGTTGTCATCGCCGACATCCAAACCGACCGGGGACACGCCTTGGCCGAACAGATCAACTCCGAATTCGGAGC
GGCCACCACTACCTACGTCCACTGCGATGTCTCTAGCGAGCCGGACGTCGAGAACGCGGTGGCGACGGCGGTTACGGCCCACGGGAAGCTGGACATCATG
TACAGCAACGCTGGGATAGGCGGCCAGAACAACATTCTCGCCAAGATAACCGACGTCGACCGTTCCGACCTGGAACGGGTCTTCGCGGTCAACGTCTACG
GCGGGGGCTTTGCCGCGAAATACGGCCCGCGAAGTACGCGGCGAGGGCGATGA
AA sequence
>Lus10013594 pacid=23168751 polypeptide=Lus10013594 locus=Lus10013594.g ID=Lus10013594.BGIv1.0 annot-version=v1.0
MKGSALLLSPIAQRLKDKVAVITGGASGIGASTARHFAKHGAKVVIADIQTDRGHALAEQINSEFGAATTTYVHCDVSSEPDVENAVATAVTAHGKLDIM
YSNAGIGGQNNILAKITDVDRSDLERVFAVNVYGGGFAAKYGPRSTRRGR
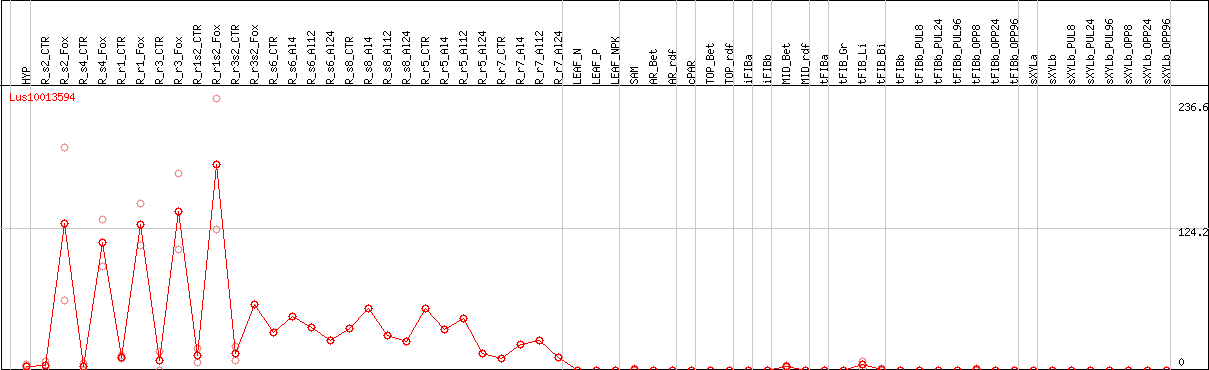
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10013594 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.