External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G80580 81 / 5e-18
AP2_ERF
Integrase-type DNA-binding superfamily protein (.1)
AT1G28370 66 / 2e-13
AP2_ERF
AtERF11
ERF domain protein 11 (.1)
AT1G53170 66 / 4e-13
AP2_ERF
ATERF8, ATERF-8
ETHYLENE RESPONSE ELEMENT BINDING FACTOR 4, ethylene response factor 8 (.1)
AT3G20310 66 / 5e-13
AP2_ERF
ATERF7, ATERF-7
ethylene response factor 7 (.1)
AT3G15210 66 / 6e-13
AP2_ERF
ATERF4, RAP2.5, ATERF-4
RELATED TO AP2 5, ethylene responsive element binding factor 4 (.1)
AT1G12980 66 / 1e-12
AP2_ERF
DRN, ESR1
ENHANCER OF SHOOT REGENERATION 1, DORNROSCHEN, Integrase-type DNA-binding superfamily protein (.1)
AT1G28360 62 / 6e-12
AP2_ERF
AtERF12
ERF domain protein 12 (.1)
AT1G24590 64 / 7e-12
AP2_ERF
ESR2, DRNL, SOB2, DRN-LIKE
FOR SUPPRESSOR OF PHYTOCHROMEB-4 2, ENHANCER OF SHOOT REGENERATION 2, DORNROSCHEN-like (.1)
AT5G44210 62 / 7e-12
AP2_ERF
AtERF9, ATERF-9
ERF DOMAIN PROTEIN- 9, erf domain protein 9 (.1)
AT1G50640 62 / 2e-11
AP2_ERF
ATERF3
ethylene responsive element binding factor 3 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026053
173 / 6e-54
AT5G18560 97 / 7e-24
Integrase-type DNA-binding superfamily protein (.1)
Lus10033664
64 / 2e-12
AT3G15210 119 / 2e-33
RELATED TO AP2 5, ethylene responsive element binding factor 4 (.1)
Lus10017907
64 / 4e-12
AT1G24590 134 / 1e-37
FOR SUPPRESSOR OF PHYTOCHROMEB-4 2, ENHANCER OF SHOOT REGENERATION 2, DORNROSCHEN-like (.1)
Lus10039857
63 / 5e-12
AT5G44210 138 / 8e-41
ERF DOMAIN PROTEIN- 9, erf domain protein 9 (.1)
Lus10035076
64 / 6e-12
AT1G24590 143 / 2e-40
FOR SUPPRESSOR OF PHYTOCHROMEB-4 2, ENHANCER OF SHOOT REGENERATION 2, DORNROSCHEN-like (.1)
Lus10015415
64 / 7e-12
AT5G44210 134 / 8e-39
ERF DOMAIN PROTEIN- 9, erf domain protein 9 (.1)
Lus10031526
62 / 2e-11
AT1G50640 112 / 2e-30
ethylene responsive element binding factor 3 (.1)
Lus10035129
61 / 6e-11
AT5G13910 152 / 7e-46
LEAFY PETIOLE, Integrase-type DNA-binding superfamily protein (.1)
Lus10035329
61 / 7e-11
AT3G15210 115 / 8e-31
RELATED TO AP2 5, ethylene responsive element binding factor 4 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0081
MBD-like
PF00847
AP2
AP2 domain
Representative CDS sequence
>Lus10014345 pacid=23164680 polypeptide=Lus10014345 locus=Lus10014345.g ID=Lus10014345.BGIv1.0 annot-version=v1.0
ATGTCGACGTCTTTCTTCCCGCAACCCCAAGGAACTGATCATTTGTTTCAAGAAGCAGTCGAAACGGTGCCGGTTCTGGAGGGGATAGCTGCCGTTGTTG
GGGAGGAGATCCTGTACGGTACCACCACCATCCGACGAGCGACGAGCTGTACACGTGTCAACGAGCAACAACAAGGTGGGGGGGACCACCACCACCAGGC
GACGACAGGCGGAGCGAGAAACTACAGAGGGGTGAGGAAGCGGCCGTGGGGGAGGTTTTCGGCGGAGATAAGGGACAGGGTGGGGAGGTGCCGTCACTGG
CTGGGCACGTTTGACACTGCGGAGGAGGCGGCGCTTGCTTACGACTCCGCCGCTTTGAGGCTCAGAGGCGCCAAGGCCAAGACCAACTTCCTCGTCGTCC
CTTCTACTTCCCTACTTCCCTCGCCTACCTCCCTACGCCCCCACCAAAAAAACACAACCCGCAGCCACCGTAAAAAAAGAAGAGAAACGACGGCGATTGT
GGTGGTGGTGATAGTCATGAATGGGCGGTGGAGACTAGTTCGGCGGCTCGTCTTTTCAGCGGTGGTGGTGATGGTGGTTATGGACTCGGCGGTGTTAGTA
ATAACTACACTAGTAGTCGTGGTAGTACTAGTGGTGGTGCGGTGGGGCTGGAGCTGGAGTTGGGGTTTGGTGTAG
AA sequence
>Lus10014345 pacid=23164680 polypeptide=Lus10014345 locus=Lus10014345.g ID=Lus10014345.BGIv1.0 annot-version=v1.0
MSTSFFPQPQGTDHLFQEAVETVPVLEGIAAVVGEEILYGTTTIRRATSCTRVNEQQQGGGDHHHQATTGGARNYRGVRKRPWGRFSAEIRDRVGRCRHW
LGTFDTAEEAALAYDSAALRLRGAKAKTNFLVVPSTSLLPSPTSLRPHQKNTTRSHRKKRRETTAIVVVVIVMNGRWRLVRRLVFSAVVVMVVMDSAVLV
ITTLVVVVVLVVVRWGWSWSWGLV
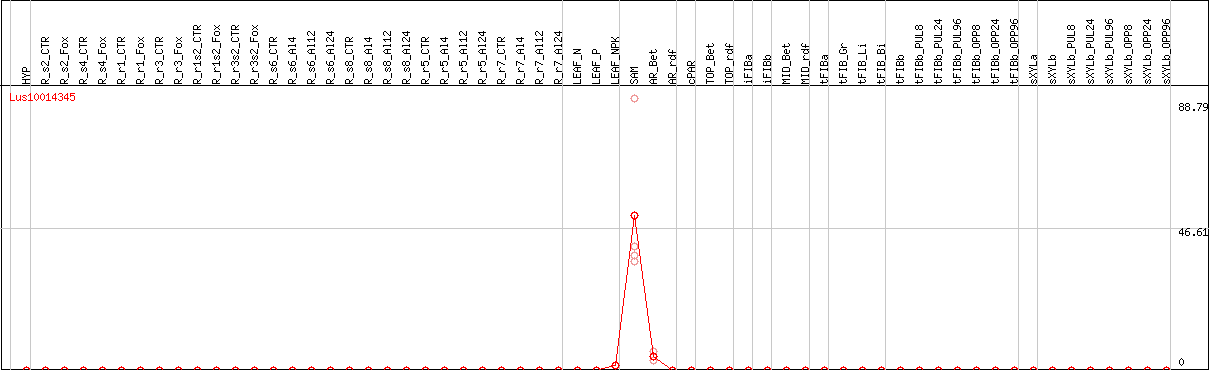
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10014345 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.