Lus10014829 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10014829 pacid=23150207 polypeptide=Lus10014829 locus=Lus10014829.g ID=Lus10014829.BGIv1.0 annot-version=v1.0
ATGCATGATCAGGACTCCCGTCCACAACGGCCGCCTATTCGCGGCCGTCTTCTTGTGAATCCTTTGACCAGGACCCTCGACCCACCACCTCGCGGCCGCC
TCCTAGTTCGTCATGGCGATCCTCCTCCTCTTCGTGGCTGCCTTCTAACTACTCCTACAATTCAAAATCCTTCTCCTTCCGACGACCAGCCTACAACCAC
AATACCACCTTTGTCGAAATACGACGCATTCATAAGCTTCCGAGGGGAGGACATCCGTAACAAATTCCTGAGTCACCTATACACTCACATGCACGACAGG
AGTATAGAAACTTACAAAGACGACGTCAGCCTCAGGCGAGGAGATCAAATCTCCCCTGCCCTCGCCCGAGGCATCGAGATGTCATCAACGTACGTCGTGG
TCTTCTCCCCGAACTACGCCACATCGAAGTGGTGCTTGGACGAGCTTGTGAAGATTTTGGAGTGCAGTGTAAAGTACGGGAGGAAGTTGATACCCATATT
CTACGACGGAGTGGACCCTTCAATTGTTAAGAACAGAACAGGGAGCTACGGCGATGCGTTTAAGGAACACGAGAAACTTCCCCCGGTGATGCAGCCTAAA
GTCCAAAGATGGAGAGATGCTCTGCGGGAAGCTGCTCACTTTGCCGGGTTTGATTCCCTGGTCGTTACGCCTGAAAGTCGCCTCATCAGAGAAGTGGCAA
AAAGTGTATTGGACGCAGTAATACCACATCAGAAGGACCCATCTCCCAGCATGAACAATTTGGTGGGCATAGACCATGAGATTGAAAAAATCAATGCATT
ACTAGACATTAACGATGAAGAAAAGAACCGGATCATAGGTTTATGGGGGATGGGTGGCATTGGCAAGACCACCCTAGCCAAAGAAATATTCTACAACATC
GATCCAGAACGATTTGAAAGTTGTCATTACATCCAAAACTTTGCAGCCCAATTGAAGAGAACACAAGTGACTGATCTACAAAATGAACTCTTCTCCATTT
TGTTACAAGATGAAAGTGCACACACCATGCCCATGAATCTGAAACTCAACCGCCTTAGCCGAAAAAGATGTTTCGTAATCATCGATGACGTATCGAGCAC
AATACAATTAGAAGCTCTACTTCTTGGAAGATACCGTGATCTATTCGGACCAGGGAGTCGAATCATCTTGACAAGCACGGATCGACAAGTTCTGAGGCAG
GTTTGTGAAGTTAAAGAAATCCACAAAGTTAAAGGGCTAGATGCTGATAAAGCCCTGCAACTGCTAAGCATGTATGCTTTGAAGAAGGATAGCCCACCAA
GTGAATACGCAGAGAGGTGTAAGAAGGTTGTTTCTTATGCGAGTGGGAACCCATTGGCATTAAAAGTATTGGGTTCAGCTTTGTACCAGCAAAAGATTGA
GTATTGGGATGGTACATTGGAGAAGCTGAGGAGCCTTCCTGCTACTAAAGAGATTGAGGATATACTGATGATTAGTTTTGATGAGTTGAGTGATGAGGAA
CAGAATGCGTTCCTTGATATTGCTTGCTTCTTCAACCATGAGGATCTTGACTTTGTGAGGAGGGTTTTGGATGACGTCATGGTTACAAATTTGGTGGACA
AGTCCCTTGCAGCGGAGGGGAGCACCGATGACAATAAAGGATGGATAATAGAAATGCATGCCTTGCTTCAACATTTGGGCGAGAGGATTGTAAACAAGGA
AAAACTCATTGGAAATCGGTCTAGGTTGTGGAATCCCAACGATGTGCATCCTTTGATCAACCAAAATATGGGAACAGAAAACACTCAGGGCATCTTTCTG
GACTTGAGCAAAACAGAGGAGATGTCTAGTCCCAAATCCGACAGTTTTGAGAGGATGAGGAACCTGAGATTGTTGTCCATTCAGGTATCCATGAATCGAA
AGCTTAACTCAGCTCATGATCTTCCTCTTGAAGTGAAGAAGAAGAAAATGAGTGTCGTTGGGGGGATTAAGTCGCTTCCTCGTGAGCTAAGGTATCTGCA
ATGGGACTTGTTTACTCAAACATCTCTACCATCTGGACTTTGGGCCCAAAATCTTCTCGTGCTCAGGCTGAAACATAGCAATATCTTACATCTATGGGAA
GGACCTAACCTGGACCTTGGGAATTTGAAAGAGCTCAATCTGTTTGCATCTAAATTCCTAACCAACTTACCTGACTTATCAACCTCTAAAAAGCTGGAGG
TGATGGACCTCACAAGTTGTACAAGCCTAACAGAGCTTCATGACTCCATCATAAATCTTCCCAGCCTCAAAATTCTCATCGTAAATTCCTGCGGAGCCCT
CAACCTTGACAATCTGAACAACAAGGAGGATGGCAAAAATCCAGCCTTCCCAAGTTTGGAAGAAGCAAACTTAGACAAGACACCAATAGACATGGTTCCC
CACTTGATCATATATGCTCCCAATCTCTCTAAACTCAGTTGCTCATCCTGCCTCCAACTGTCAGAGTTTCCAAATACCTCGGAGCCTCAGGAAACATTAC
TGCAACTTAATGTGAGCAATTGTGGCAGGATATCGGCCGTTCCTGCTAGGGTAACAAATTTTCTGAACTTGAAGGTGTTTGATCTATCGGGGACAGGGAT
CCTGGTGCTACCATATTCTGTCTTAGAGTTGGATAAATTGGAGGAGTTGAAGTTTGCCCAGTGTAAGAAGCTTGTTATTATACCACGTAGCTTCAAGAGG
CTGACTCATTTGGAAAAGCTAGAGCTCCAAGGTTGTGATCTGCTGATTAATCAGAAGCCAGAGTTGCCCGACGGCTTGGAGTATACAGACGACGGAGAAC
ACAACAACACCAAGAGCAAAGTTCGAAAGTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10014829 pacid=23150207 polypeptide=Lus10014829 locus=Lus10014829.g ID=Lus10014829.BGIv1.0 annot-version=v1.0
MHDQDSRPQRPPIRGRLLVNPLTRTLDPPPRGRLLVRHGDPPPLRGCLLTTPTIQNPSPSDDQPTTTIPPLSKYDAFISFRGEDIRNKFLSHLYTHMHDR
SIETYKDDVSLRRGDQISPALARGIEMSSTYVVVFSPNYATSKWCLDELVKILECSVKYGRKLIPIFYDGVDPSIVKNRTGSYGDAFKEHEKLPPVMQPK
VQRWRDALREAAHFAGFDSLVVTPESRLIREVAKSVLDAVIPHQKDPSPSMNNLVGIDHEIEKINALLDINDEEKNRIIGLWGMGGIGKTTLAKEIFYNI
DPERFESCHYIQNFAAQLKRTQVTDLQNELFSILLQDESAHTMPMNLKLNRLSRKRCFVIIDDVSSTIQLEALLLGRYRDLFGPGSRIILTSTDRQVLRQ
VCEVKEIHKVKGLDADKALQLLSMYALKKDSPPSEYAERCKKVVSYASGNPLALKVLGSALYQQKIEYWDGTLEKLRSLPATKEIEDILMISFDELSDEE
QNAFLDIACFFNHEDLDFVRRVLDDVMVTNLVDKSLAAEGSTDDNKGWIIEMHALLQHLGERIVNKEKLIGNRSRLWNPNDVHPLINQNMGTENTQGIFL
DLSKTEEMSSPKSDSFERMRNLRLLSIQVSMNRKLNSAHDLPLEVKKKKMSVVGGIKSLPRELRYLQWDLFTQTSLPSGLWAQNLLVLRLKHSNILHLWE
GPNLDLGNLKELNLFASKFLTNLPDLSTSKKLEVMDLTSCTSLTELHDSIINLPSLKILIVNSCGALNLDNLNNKEDGKNPAFPSLEEANLDKTPIDMVP
HLIIYAPNLSKLSCSSCLQLSEFPNTSEPQETLLQLNVSNCGRISAVPARVTNFLNLKVFDLSGTGILVLPYSVLELDKLEELKFAQCKKLVIIPRSFKR
LTHLEKLELQGCDLLINQKPELPDGLEYTDDGEHNNTKSKVRK
|
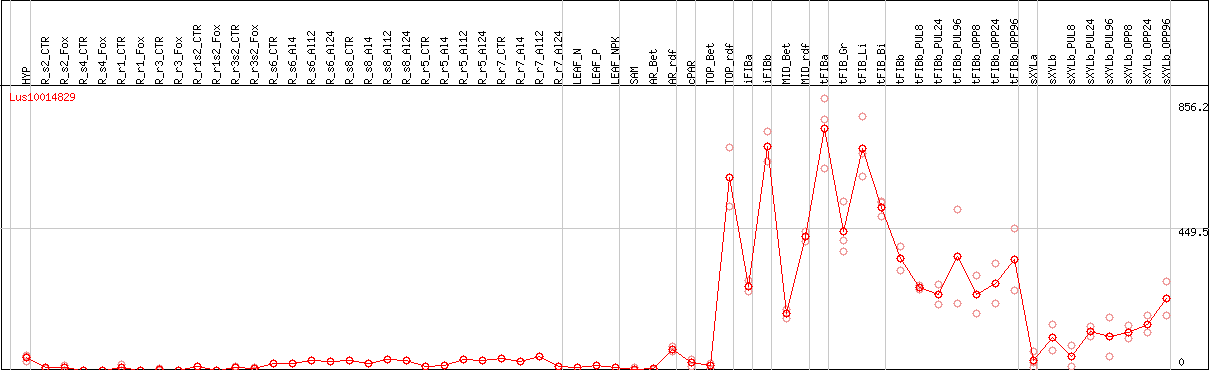
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10014829 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
