Lus10015648 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10015648 pacid=23147266 polypeptide=Lus10015648 locus=Lus10015648.g ID=Lus10015648.BGIv1.0 annot-version=v1.0
ATGGCTGCAGATATTTCTTCTTCTGCACCTCGTACTTCTCCTTATACCGGAGAATGGGAGTACGATGTTTTCTTGTGTTTTAGAGGCGACACACGCCACG
GTTTCACCAGCCACCTCCTGTCTGCTTTATCAGACAAGCAAATCAGAACCTTCATTGACCACAAGCTCGCAAAAACTGAAAGCATCGATGAGCTAATCTC
GATCCTCCAGAGATGTGCTCTTTCTGTGGTGGTTTTCTCCGAAAAGTTTGCGGATTCAGAGTGGTGCTTGGAAGAGGTGGTTACTATTGCTGAAAGGATG
AAGAAAGTTGGGCACAGAGTTCTTCCGGTTTTCTATAAAGTGGATCCGTTCGATGTGACAGATGAACCGAGAAGCTATATGGCTACCATCGATCGTGAAT
ATAAAGCTAGAAGTAGCTTTTTGGAGGATAAGAAGAGATGGATGGATGCTGTGAACGCTGTGGCTAATTGTGCTGGTCATACTTCGCAGGCCATCAAAAT
TGAATCGGAATTGATTAAGGCGGTTGTGGAAACTGTTCAGAAGCAATTAATTGACATGTCACCGAGTATCAATCGTAATAACTTGGTTGCTATGGGTTCA
CGTATTTTCGAGATTGAGCGACTATTAGCTATGGACAAACTAGATGATACTTGCATCATTGGGCTATGGGGAATGGGCGGTGTTGGGAAAACAACTCTAG
CCGAAGCGTGTTATGAAAGGGTAACTTCGTCCAACAAAGGAATCAAACATCTTTTCGTTCGAAACGTGAATGAAATATGTGAAAAGCATCATGGCGTTGA
AAAAATTGTTCACAAACTCTATTCAAAATTGTTGGACGAGAATAACATCGACCGTGAAGATTTGAATATTGCTTATCGAAGGGAACGTCTTTCGCGCTCG
AGGGTATTCGTTGTTCTAGACAACGTTGAAACTTTGGAGCAATTGGAGCAATTGGCTTTAGGATATGTGTTCAATCTCAGCAAAGTCTTTGCAGCGGGGA
GTAGGATCATCATCACCACAAGAAATAAAAAGGTGCTTCAAAATGCTATGGCCAAGATTTACAATGTCGAGTGTTTGAATAACAAGGAATCTATCCGATT
GTTTAGCTTACATGCATTCAAACAAGATCGACCTCAAGATAATTGGACGGATAAGTCACATCTTGCAATATCATATTGCAAGGGGAATCCTTTGGCACTC
AAAATCTTGGGGGGTGCATTGTTCGGTGAAGATGTACATTATTGGAGAAGTCTCTTGACCGGTTTGAGGCAATCTGGAAACCTAGGTATTGAGAGTATTC
TGAGGAGAAGTTATGATAAACTTGGGAAGGAAGAGAAGAAAATATTTATGGATGTTGCCTGTCTACTCTATGGCATGTCAAGATCTAGACTGATTGATTA
TATGGCGACCATGTATTCCTCTTCTTATGTCAGGGTGAAGGATTTAATTGATAAATCCCTATTAACATGTGTTCCTAGCGAGAATGGAGAGATGATTGAG
GTGCATGATTTGCTTAAAGAAATGGCTTGGAACATTGTCAAAGAAGAACCAAAGCTTGGCAAACGAAGCAGATTAGTAGATCCGGATGATGTCCATAAAT
TATTGTCAACTTCAGAGGTTAAAAATTGGAGTACATCAATCGTCAACCTCTTCAAAGGCATAGTGATGGTCATACCAAGAAGGAAAAGAAGAAAAGTCAC
TGACATGCATGAGAGAGGCTATGATCCATTGGAGGAACATCGAACAACTGAAGGTATATGTTTGGATCTCTCTGGAACTAAAGAGATGTATTTGAAAGCA
AATGCCTTTGAGGGTATGAATTCTCTTACATTTCTCAAATTCAAGTCGCCAGAACTTGATTATGCTCAATATCCTTTAAAAAATGTGAAGACGAAAATTC
ATTTGCCTTACGATGGACTTAACTCCCTTCCGGAGGGGTTAAGATGGTTGCAGTGGGATGGATATCCTTCAAAATCTCTACCGGCAAAGTTTTATCCCCA
ACACCTTGTTCATCTCATTATACGTGGAAGTCCAATTCGAAGATGTTGGGAAGGATATGATCAACCCCAACTTGTGAACTTGATTGTGCTTGATCTCCGC
TACTGTACAAATCTAATCGCTATTCCGGATATATCAAGTTCTTTGAATCTCGAAGAACTCCTTCTTTTTGGATGCAGAAGCCTAGTTGAGGTACCATTTC
ATGTTCAATATTTGACCAAGCTGGTAACGCTAGACATCAATGTTTGCAAAAATCTCAAGCGTCTTCCACCTAAACTTGACTCAAAGTTACTTAAGCACGT
TCGGATGCAAGGGCTCGGGATCACTCGCTGTCCAGAGATTGATTCAAGAGAGTTGGAGATATTTGATTTACGTTTTACATCTCTTGGAGAGTTACCCAGT
GCAATTTACAATGTCAAGCAAAATGGAGTCCTACGTCTCCATGGCAAAAACATTACCAAATTTCCTGGAATTACAACAATATTGAAGCTTTTTACATTAA
GCCGTACTTCAATAAGGGAGATTGATTTGGCCGATTATCATCAACAGCATCAGACTTCTGATGGATTACTATTGCCAAGATTTCAGAATTTGTGGTTGAC
AGGCAACCGGCAACTTGAGGTCCTGCCAAACAGTATATGGAACATGATTTCTGAAGAACTTTATATTGGTCGGAGCCCGCTGATTGAAAGTTTGCCGGAA
ATCTCAGAGCCTATGAGTACCTTAACTTCTCTTCATGTATTCTGTTGTAGAAGCCTCACAAGTATTCCAACTAGCATCAGCAATTTAAGATCTCTGAGGT
CGCTTCGTTTAGTAGAGACGGGTATTAAGTCACTGCCTTCTAGCATCCAAGAACTAAGGCAGCTTTTTTCGATTGATCTTAGAGATTGCAAAAGCCTCGA
GTCAATTCCAAACAGTATCCATAAGCTTTCCAAGTTGGTAACTTTGTCAATGTCTGGCTGTGAGATCATTATATCTTTGCCTGAACTCCCACCAAATCTG
AAGACCTTGAACGTAAGTGGTTGCAAGTCTTTACAAGCTCTACCAAGCAACACATGTAAGCTGTTGTACCTGAATAGGATCTATTTTGAGGAATGTCCAC
AAGTGGATCAGACTATTCCTGCTGAGTTCATGGCAAATTTCCTCGTTCATGCCAGCTTGTCTCCGTCATATGAGCGGCAAGTTAGATGTTCAGGAAGCGA
GCTTCCGAAATGGTTTTCGTACAGAAGCATGGAGGACGAGGATTGCAGTACTGTGAAAGTGGAGTTGCCTCTTGCTAATGATAGTCCTGACCATCCAATG
ATTAAGGGAATTGCATTCGGCTGTGTGAATTCTTCCGACCCTTACTACTCTTGGATGCGTATGGGATGTCGTTGCGAGGTCGGTAACACCACTGTTGCAT
CTTGGGTTTCCAACGAAAAAGTAATGGGCCCTGAAGAAAAGTCATCAGAAAAGGTATGGTTGGTGTTCAACAAGAATTTATCAAGTACGGGGTCTATGGG
AAGCGAAGAAGATGAAGCGTGGTATGTGAAGTATGGCGGGTTCGATGTCTCATTCAATTTCTATTTTCTAGATTACGACGATGAAATCATAAAGAAGGTG
AAAATAAAAAGATGTGGAGTCTCTCTGATGTATTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10015648 pacid=23147266 polypeptide=Lus10015648 locus=Lus10015648.g ID=Lus10015648.BGIv1.0 annot-version=v1.0
MAADISSSAPRTSPYTGEWEYDVFLCFRGDTRHGFTSHLLSALSDKQIRTFIDHKLAKTESIDELISILQRCALSVVVFSEKFADSEWCLEEVVTIAERM
KKVGHRVLPVFYKVDPFDVTDEPRSYMATIDREYKARSSFLEDKKRWMDAVNAVANCAGHTSQAIKIESELIKAVVETVQKQLIDMSPSINRNNLVAMGS
RIFEIERLLAMDKLDDTCIIGLWGMGGVGKTTLAEACYERVTSSNKGIKHLFVRNVNEICEKHHGVEKIVHKLYSKLLDENNIDREDLNIAYRRERLSRS
RVFVVLDNVETLEQLEQLALGYVFNLSKVFAAGSRIIITTRNKKVLQNAMAKIYNVECLNNKESIRLFSLHAFKQDRPQDNWTDKSHLAISYCKGNPLAL
KILGGALFGEDVHYWRSLLTGLRQSGNLGIESILRRSYDKLGKEEKKIFMDVACLLYGMSRSRLIDYMATMYSSSYVRVKDLIDKSLLTCVPSENGEMIE
VHDLLKEMAWNIVKEEPKLGKRSRLVDPDDVHKLLSTSEVKNWSTSIVNLFKGIVMVIPRRKRRKVTDMHERGYDPLEEHRTTEGICLDLSGTKEMYLKA
NAFEGMNSLTFLKFKSPELDYAQYPLKNVKTKIHLPYDGLNSLPEGLRWLQWDGYPSKSLPAKFYPQHLVHLIIRGSPIRRCWEGYDQPQLVNLIVLDLR
YCTNLIAIPDISSSLNLEELLLFGCRSLVEVPFHVQYLTKLVTLDINVCKNLKRLPPKLDSKLLKHVRMQGLGITRCPEIDSRELEIFDLRFTSLGELPS
AIYNVKQNGVLRLHGKNITKFPGITTILKLFTLSRTSIREIDLADYHQQHQTSDGLLLPRFQNLWLTGNRQLEVLPNSIWNMISEELYIGRSPLIESLPE
ISEPMSTLTSLHVFCCRSLTSIPTSISNLRSLRSLRLVETGIKSLPSSIQELRQLFSIDLRDCKSLESIPNSIHKLSKLVTLSMSGCEIIISLPELPPNL
KTLNVSGCKSLQALPSNTCKLLYLNRIYFEECPQVDQTIPAEFMANFLVHASLSPSYERQVRCSGSELPKWFSYRSMEDEDCSTVKVELPLANDSPDHPM
IKGIAFGCVNSSDPYYSWMRMGCRCEVGNTTVASWVSNEKVMGPEEKSSEKVWLVFNKNLSSTGSMGSEEDEAWYVKYGGFDVSFNFYFLDYDDEIIKKV
KIKRCGVSLMY
|
DESeq2's median of ratios [FLAX]
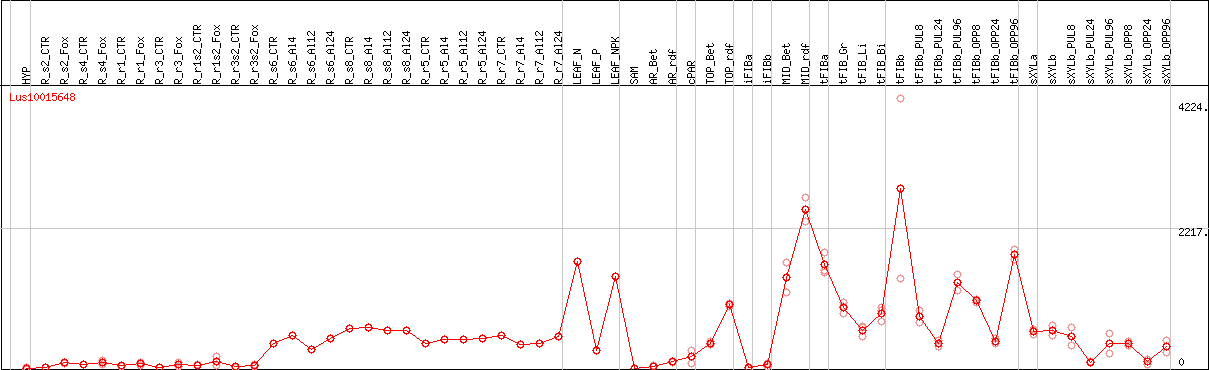
Coexpressed genes
Lus10015648 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
